13.3
Impact Factor
Theranostics 2016; 6(3):428-434. doi:10.7150/thno.13986 This issue Cite
Research Paper
68Ga-Pentixafor-PET/CT for Imaging of Chemokine Receptor 4 Expression in Glioblastoma
1. Department of Nuclear Medicine, University Hospital Würzburg, Würzburg, Germany
2. Comprehensive Cancer Center Mainfranken, Würzburg, Germany
3. Institute of Pathology, University of Würzburg, Würzburg, Germany
4. Department of Neurosurgery, University Hospital Würzburg, Würzburg, Germany
5. Scintomics GmbH, Fürstenfeldbruck, Germany
6. Pharmaceutical Radiochemistry, Technische Universität München, Munich, Germany
7. Department of Molecular and Medical Pharmacology, David Geffen School of Medicine at UCLA, Los Angeles, USA
* Equal contribution
Received 2015-9-28; Accepted 2015-11-20; Published 2016-1-25
Abstract
Chemokine receptor-4 (CXCR4) has been reported to be overexpressed in glioblastoma (GBM) and to be associated with poor survival. This study investigated the feasibility of non-invasive CXCR4-directed imaging with positron emission tomography/computed tomography (PET/CT) using the radiolabelled chemokine receptor ligand 68Ga-Pentixafor.
15 patients with clinical suspicion on primary or recurrent glioblastoma (13 primary, 2 recurrent tumors) underwent 68Ga-Pentixafor-PET/CT for assessment of CXCR4 expression prior to surgery. O-(2-18F-fluoroethyl)-L-tyrosine (18F-FET) PET/CT images were available in 11/15 cases and were compared visually and semi-quantitatively (SUVmax, SUVmean). Tumor-to-background ratios (TBR) were calculated for both PET probes. 68Ga-Pentixafor-PET/CT results were also compared to histological CXCR4 expression on neuronavigated surgical samples.
68Ga-Pentixafor-PET/CT was visually positive in 13/15 cases with SUVmean and SUVmax of 3.0±1.5 and 3.9±2.0 respectively. Respective values for 18F-FET were 4.4±2.0 (SUVmean) and 5.3±2.3 (SUVmax). TBR for SUVmean and SUVmax were higher for 68Ga-Pentixafor than for 18F-FET (SUVmean 154.0±90.7 vs. 4.1±1.3; SUVmax 70.3±44.0 and 3.8±1.2, p<0.01), respectively. Histological analysis confirmed CXCR4 expression in tumor areas with high 68Ga-Pentixafor uptake; regions of the same tumor without apparent 68Ga-Pentixafor uptake showed no or low receptor expression.
In this pilot study, 68Ga-Pentixafor retention has been observed in the vast majority of glioblastoma lesions and served as readout for non-invasive determination of CXCR4 expression. Given the paramount importance of the CXCR4/SDF-1 axis in tumor biology, 68Ga-Pentixafor-PET/CT might prove a useful tool for sensitive, non-invasive in-vivo quantification of CXCR4 as well as selection of patients who might benefit from CXCR4-directed therapy.
Keywords: Glioblastoma, neuro-oncology, chemokine receptor, CXCR4, PET.
Introduction
Glioblastoma (GBM) is the most common malignant primary tumor of the central nervous system, accounting for 45%-50% of all gliomas with an incidence of about 3-4 per 100.000 inhabitants per year [1]. Despite multimodality treatment approaches including surgery and (concomitant) radiotherapy as well as temozolomide, 5 year survival is abysmal at 5% [1, 2]. Thus, there is still an urgent clinical need for new therapeutic targets.
Chemokine receptor-4 (CXCR4) is a member of the G-protein-coupled chemokine receptor family [3]. The receptor plays an important role in a variety of physiologic processes that rely on the recruitment and homing of stem cells, progenitor cells, and immune cells [3, 4]. It is thus important in embryogenesis, neoangiogenesis, hematopoiesis, and inflammation. CXCR4 is overexpressed in more than 20 human tumor types including ovarian, prostate, esophageal, and renal cell carcinoma [5], promoting tumor growth and progression, tumor invasiveness, and metastasis [6]. In GBM, CXCR4 has also been reported to be overexpressed and associated with tumor angiogenesis [7] as well as poor patient survival [8, 9]. Moreover, in animal xenograft experiments, treatment with a CXCR4 antagonist significantly inhibited the tumorigenicity and the growth of tumors [10], suggesting that CXCR4 may play a critical role in promoting the progression of human gliomas. A very recent study reported on activation of the ERK and PI3K/AKT signalling pathway by CXCL12/CXCR4 and its potential implication for targeted therapy of GBM [11]. Hence, the CXCR4/CXCL12 axis represents a highly relevant molecular target of cancer biology and offers promising new approaches and techniques for targeted cancer therapy [12, 13].
Recently, 68Ga-Pentixafor, a radiolabelled cyclic pentapeptide with high affinity to CXCR4, excellent PET imaging characteristics and favorable human dosimetry has been developed [14-16]. Proof-of-concept for visualization of CXCR4-expression has recently been demonstrated in patients with hematologic malignancies [17, 18]. The present study aimed at the evaluation of 68Ga-Pentixafor-PET in patients with glioblastoma.
Patients and Methods
68Ga-Pentixafor was administered on a compassionate use base in compliance with §37 of the Declaration of Helsinki and The German Medicinal Products Act, AMG §13 2b and in accordance with the responsible regulatory body (Regierung von Oberfranken). The study adhered to the standards established in the declaration of Helsinki. All patients gave written informed consent to receive CXCR4-PET/CT imaging on a compassionate use basis.
Patients
From April 2014 to April 2015, a total of 15 patients (7 males, 8 females, mean age, 61±12 years; range, 42-77 y) scheduled for surgical resection or biopsy of suspected primary (13/15 pt.) or recurrent (2/15 pt.) GBM were enrolled (for details Table 1). Both patients with suspected recurrence had previously undergone surgical resection followed by irradiation and concomitant and adjuvant chemotherapy with temozolomide. The time from completion of radiation to the PET scan was > 12 weeks in these 2 patients.
68Ga-Pentixafor-PET was done within 1-23 days prior (mean, 6±6 days) to surgery/ tumor biopsy. O-(2-18F-fluoroethyl)-L-tyrosine (18F-FET) PET scans for comparison were available in 11/15 subjects. Scans were performed at a mean interval of 12±6 days (range, 2-36 days) before surgery. The median time interval between both PET scans was 1 day (range, 0-35 days). Standard magnetic resonance imaging (MRI) was available in all subjects.
With surgery/biopsy scheduled in all cases, one patient rapidly deteriorated after imaging and was therefore unable to undergo tumor resection. In the remaining 14 subjects, histopathology served as gold standard to verify diagnosis as well as the presence of viable tumor tissue.
Patients' characteristics and imaging results.
| CXCR4 | FET | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Nr | Sex | Age (y) | Disease | SUVmean | SUVmax | TBR SUVmean | TBR SUVmax | SUVmean | SUVmax | TBR SUVmean | TBR SUVmax |
| 1 | F | 60 | GBM | 6.14 | 7.34 | 68.2 | 35.0 | 4.55 | 5.40 | 3.1 | 3.1 |
| 2 | M | 77 | GBM | 5.07 | 7.3 | 253.5 | 73.0 | n/a | n/a | n/a | n/a |
| 3 | F | 75 | GBM | 2.23 | 2.87 | 111.5 | 57.4 | 4.12 | 5.07 | 3.6 | 3.7 |
| 4 | F | 64 | GBM | 3.68 | 4.48 | 184.0 | 89.6 | n/a | n/a | n/a | n/a |
| 5 | M | 73 | GBM | 2.41 | 3.04 | 241.0 | 152.0 | n/a | n/a | n/a | n/a |
| 6 | M | 59 | PCNSL | 4.02 | 6.01 | 201.0 | 75.1 | n/a | n/a | n/a | n/a |
| 7 | M | 42 | GBM | 2.48 | 2.96 | 248.0 | 148.0 | 5.04 | 6.48 | 6.1 | 5.7 |
| 8 | M | 64 | GBM | neg | neg | n/a | n/a | 3.12 | 3.96 | 3.3 | 3.1 |
| 9 | F | 57 | Oligo III | 0.75 | 0.89 | 25.0 | 12.7 | 6.15 | 7.19 | 4.7 | 4.3 |
| 10 | F | 48 | Ischemia | 2.59 | 3.3 | 86.3 | 27.5 | 2.64 | 3.10 | 1.9 | 1.7 |
| 11 | F | 69 | GBM | 0.95 | 1.29 | 95.0 | 64.5 | 2.81 | 3.23 | 5.5 | 4.4 |
| 12 | F | 76 | GBM | 3.29 | 4.32 | 65.8 | 43.2 | 9.38 | 11.62 | 5.3 | 5.1 |
| 13 | M | 42 | Astro III | neg | neg | n/a | n/a | 4.23 | 5.14 | 4.8 | 4.1 |
| 14 | F | 60 | GBM | 3.14 | 4.21 | 314.0 | 105.3 | 2.88 | 3.36 | 2.3 | 2.0 |
| 15 | M | 50 | GBM | 2.18 | 3.02 | 109.0 | 30.2 | 3.12 | 3.71 | 4.2 | 4.1 |
In addition to clinical factors, results of both 68Ga-Pentixafor- as well as 18F-FET-PET are given. n/a = not applicable; SUV = standardized uptake value; TBR = tumor-to-background ratio.
Preparation of 68Ga-Pentixafor and 18F-FET
Synthesis of 68Ga-Pentixafor was performed by means of a fully GMP compliant automated synthesizer (GRP, Scintomics, Germany) [15, 19]. 18F-FET was prepared according to the corresponding monography in the European Pharmacopoeia as previously described [20, 21].
PET Imaging
All PET scans were performed on a dedicated (PET/CT) scanner (Siemens Biograph mCT 64; Siemens Medical Solutions). For 68Ga-Pentixafor, injected activity ranged from 83 to 158 MBq (mean: 130±28 MBq). Imaging started 60 min after tracer injection. 18F-FET scans were started 20 min after injection of 180-229 MBq (mean: 207±14 MBq).
Low dose CT scans of the brain for attenuation correction were acquired (35 mAs, 120 keV, a 512 × 512 matrix, 5 mm slice thickness, increment of 30 mm/s, rotation time of 0.5 s, and pitch of 0.8). All PET images were reconstructed using corrections for attenuation, dead-time, random events and scatter. The PET scanner is periodically checked for calibration accuracy as part of quality control according to published guidelines [22].
Image Analysis
Images were analyzed as recently described [23]. In brief, images were first inspected visually (CL, KH). Then the axial PET image slice displaying the maximum tumor uptake was selected by drawing a 3D-volume of interest (VOI) around the whole tumor area. Tumor regions of interest (ROIs) were defined in 2 ways. First, a standardized 10-mm circular region was placed over the area with the peak activity. This first ROI was used to derive maximum (SUVmax) and mean standardized uptake values (SUVmean). A normal reference brain region was defined by drawing a ROI (diameter of 25 mm) involving the entire contralateral hemisphere at the level of the centrum semiovale to derive tumor-to-background ratios. Additionally, another ROI (3D isocontour) was placed in the superior sagittal sinus (at the tumor level) to derive an estimate of blood pool activity. The radiotracer concentration in the ROIs was normalized to the injected dose per kilogram of patient's body weight to derive the SUVs.
Neurosurgery
68Ga-Pentixafor-PET/CT and T1-weighted rapid three-dimensional gradient-echo technique (MP-RAGE) MR images were transferred to a neuro-navigation system (Stealth Station S7, Medtronic Navigation, Louisville, USA) and combined in image fusion planning (StealthMerge Image Registration, Medtronic Navigation, Louisville, USA). During surgery of the gliomas, neuro-navigated biopsy specimens were obtained from areas with 68Ga-Pentixafor uptake by three experienced neurosurgeons (M.L.; T.L.; A.F.K). In 5 cases, separate tumor samples with high and low/no tracer accumulation were biopsied, respectively.
Histological characterization of tumors
All tumors were histologically assessed and graded on formalin fixed and paraffin embedded tissue sections by an experienced neuropathologist (CMM) and pathologist (IK), respectively, according to the 2007 criteria of the World Health Organization [24] and as previously described [20]. Glial origin of the tumor cells was confirmed by the positive reaction with gliofilament antiserum GFAP (1:200, Clone 6F2, Dako, Hamburg, Germany). To determine the proliferative activity of tumor cells, Ki-67 labeling index was calculated after immunostaining for MIB-1 (monoclonal, clone Ki-67, 1:50, Dako, Hamburg, Germany) by determining the number of positive nuclei among 100 tumor cells per high power field (HPF) (x400) in a total of 10 HPF per sample.
CXCR4-immunohistochemistry was performed and scored as described previously [17] using 3µm thick paraffin-slices and an anti-CXCR4 rabbit polyclonal antibody (ab2074; Abcam, Cambridge, UK) followed by detection with the DAKO en vision system according to the manufacturer′s protocol. In brief, a so-called immune-reactive score (IRS) was calculated by multiplying the percentage of CXCR4+ cells x the intensity of staining. Score points range from 0-12.
All immunostained sections were counterstained for 2 minutes with hematoxylin in order to identify the specific kind of cell (tumoral, parenchymal) exhibiting positive CXCR4 immunostaining.
Statistical Analysis
Statistical analyses were performed using PASW Statistics software (version 22.0; SPSS, Inc. Chicago, IL). Quantitative values were expressed as mean ± standard deviation or median and range as appropriate. Comparisons of related metric measurements were performed using Wilcoxon-signed rank test. The Chi square- or Fisher exact test was conducted for comparison of frequency data between independent subgroups. For bivariant correlation analyses Spearman or Pearson correlation coefficients were calculated. All statistical tests were performed two-sided and a p-value < 0.05 was considered to indicate statistical significance. No correction for p-values was applied to adjust for multiple tests.
Results
Tumor characteristics
In all patients the brain lesions were located in the cerebral hemispheres. 7/15 subjects presented with temporal (temporo-occipital) lesions, 6/15 with frontal lesions, 1 had a parietal (parieto-occipital) tumor and 1 patient had occipital disease.
Final diagnosis was established by histopathology of surgical or biopsy samples in 14/15 patients. Both patients with recurrent disease had GBM with secondary GBM evolving from grade III astrocytoma in 1 patient (patient #6). The other patient suffered from recurrence of previously diagnosed primary GBM (patient #8).
In 8 patients with suspected primary GBM the clinical diagnosis was confirmed by histopathology. One patient presented with oligodendroglioma III°, another with anaplastic astrocytoma III°. In one patient primary central nervous system lymphoma (PCNSL) could be diagnosed, inflammatory changes after stroke mimicked GBM in the remaining patient.
Visual PET Image Analysis
All patients with suspected primary and recurrent high-grade gliomas (with contrast-enhancement in T1-weighted MRI) showed enhanced 18F-FET uptake (11/11), whereas one patient with GBM (patients #8) and the single patient with anaplastic astrocytoma III° (patient #13) did not display 68Ga-Pentixafor uptake (detection rate, 13/15). All lesions were clearly delineated from normal brain tissue. Of note, highly 68Ga-pentixafor positive lesions were mainly contrast-enhancing. Overall, the two tracers showed a matching image pattern upon visual assessment in 4/11 cases with inter-individual differences in uptake intensity without a tracer preference; however, these were not of relevance for image interpretation. In 7 subjects, tracer distribution varied distinctly. One example is given in Figure 1.
Semiquantitative Image Analysis
SUVmean and SUVmax for 18F-FET were higher than those for 68Ga-Pentixafor (4.4±2.0 and 5.3±2.3; 3.0±1.5 and 3.9±2.0 respectively (p=0.05 and p=0.1)). 18F-FET and 68Ga-Pentixafor SUVmean and SUVmax were not correlated (r= 0.12 and r=0.05; p=ns).
Background SUVmean and SUVmax were much higher for 18F-FET as compared to 68Ga-Pentixafor (1.1±0.4 and 1.4±0.5 versus 0.03±0.02 and 0.08±0.05; p<0.01). Consistently, TBR for SUVmean and SUVmax were higher for 68Ga-Pentixafor than for 18F-FET (SUVmean 154.0±90.7 vs. 4.1±1.3; SUVmax 70.3±44.0 and 3.8±1.2, p<0.01), respectively (Table 1). The patient with PCNSL (patient #6) presented with 68Ga-Pentixafor SUVmean and SUVmax of 4.02 and 6.01, respectively (no 18F-FET available). In the subject with ischemic infarction (patient #10), both PET scans were visually positive. Corresponding 68Ga-Pentixafor SUVmean and SUVmax were 2.6 and 3.3 (18F-FET: 2.6 [SUVmean] and 3.1 [SUVmax], respectively) (Table 1).
In 68Ga-Pentixafor-PET, blood pool activity ranged between 1.2 and 3.2 (median, 1.6; mean, 1.7±0.6) and was significantly lower than glioma SUV (p<0.01).
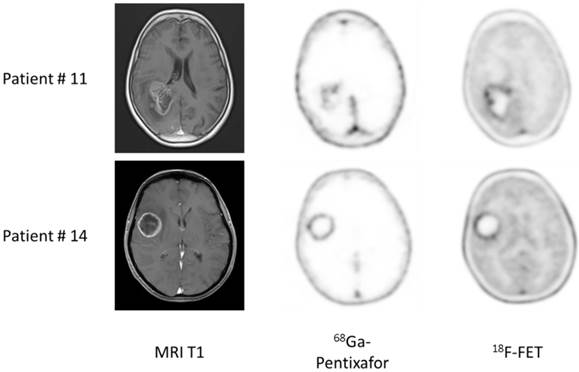
Example of dis- and concordant distribution of 68Ga-Pentixafor and 18F-FET in glioblastoma patients. Given are transaxial slices of contrast-enhanced, T1-weighted MRI, 68Ga-Pentixafor- and 18F-FET-PET in two patients with glioblastoma. Whereas tracer distribution is concordant in patient #14, marked differences can be observed in patient #11.
Immunohistochemical work-up
In 14/15 patients, imaging results could be compared to immunohistological staining for CXCR4. 2/14 samples were rated “weakly” (both IRS score 2), 6/14 “moderately” (IRS scores 4-8) and 5/14 “strongly” (IRS scores 9-12) positive (Table 2). The sample of the patient diagnosed with an ischemic stroke (patient 10) was rated “negative” (IRS score 0) due to the absence of CXCR4+ tumor cells. A high degree of CXCR4-expressing macrophages and glial cells in this specimen was found to account for the positive CXCR4-PET. The two cases with weak CXCR4-expression presented with a negative 68Ga-Pentixafor-PET scan. This apparent discrepancy is explained by the low percentage (<2% and 10-20%, respectively) of CXCR4-positive tumor cells. Considering the moderately and strongly positive sections, CXCR4 score did not correspond to the intensity of 68Ga-Pentixafor uptake (SUVmean, SUVmax, TBRmean, TBRmax).
Apart from the tissue samples from CXCR4+ regions, additional neuro-navigated biopsy specimens were available from areas with low/no 68Ga-Pentixafor uptake in another 5 patients (#1, 9, 10, 11, 12). Interestingly, a negative or low 68Ga-Pentixafor signal was associated with a negative or rather weak CXCR4 expression on tumor cells in all cases (Table 2 and Figure 2).
Immunohistochemical assessment of Ki-67 was available in 10/15 patients (Table 2), but was neither correlated with PET-derived parameters nor with CXCR4 expression (p=ns; Supplementary Figure 1). Application of the Spearman's rho correlation coefficients resulted for 68Ga-Pentixafor in R=0.354 (R2=0.125; p=0.316) for both SUVmax and SUVmean. Corresponding values for IRS were R=0.332 (R2=0.11; p=0.348). In 2 cases, the level of Ki-67 expression could be compared in 68Ga-Pentixafor positive vs. negative sections: while the proliferative index of the CXCR4- section was lower than that of the CXCR4+ one in patient #9 (5% vs. 15%), it did not differ in the other patient (#11; 5% for both).
Immunohistochemical assessment of surgical samples for CXCR4 expression and Ki-67 compared to 68Ga-Pentixafor-PET.
| Nr | CXCR4 SUVmean | Biopsy site | CXCR4 IRS score | CXCR4 IRS classification | Ki-67 |
|---|---|---|---|---|---|
| 1 | 6.14 | CXCR4+ | 6 | 2 | 5% |
| CXCR4- | 0 | 0 | n/a | ||
| 2 | 5.07 | CXCR4+ | 9 | 3 | 20% |
| 3 | 2.23 | n/a | n/a | n/a | n/a |
| 4 | 3.68 | CXCR4+ | 6 | 2 | 20% |
| 5 | 2.41 | CXCR4+ | 6 | 2 | 10% |
| 6 | 4.02 | CXCR4+ | 12 | 3 | >80% |
| 7 | 2.48 | CXCR4+ | 4 | 2 | 10% |
| 8 | CXCR4- | 2 | 1 | 2% | |
| 9 | 0.75 | CXCR4+ | 9 | 3 | 15% |
| CXCR4- | 4 | 2 | 5% | ||
| 10 | 2.59 | CXCR4+ | 0 | 0 | 25% |
| CXCR4- | 0 | 0 | n/a | ||
| 11 | 0.95 | CXCR4+ | 9 | 3 | 5% |
| CXCR4- | 2 | 1 | 5% | ||
| 12 | 3.29 | CXCR4+ | 12 | 3 | 40% |
| CXCR4- | 2 | 1 | n/a | ||
| 13 | neg | CXCR4- | 2 | 1 | 15% |
| 14 | 3.14 | CXCR4+ | 6 | 2 | 30% |
| 15 | 2.18 | CXCR4+ | 6 | 2 | n/a |
Immunohistochemical assessment of CXCR4+ and CXCR4- tumor samples (as assessed by 68Ga-Pentixafor-PET) in comparison to 68Ga-Pentixafor-PET results (CXCR4-SUVmean) and Ki-67 labelling index. For histological graduation, a so-called immunoreactive score was calculated by multiplying the percentage of CXCR4+ cells x the intensity of staining. Score points range from 0-12. For classification, the following system was used: 0-1 = 0 (negative); 2-3 = 1 (weak expression); 4-8 = 2 (mild expression); 9-12 = 3 (strong expression).
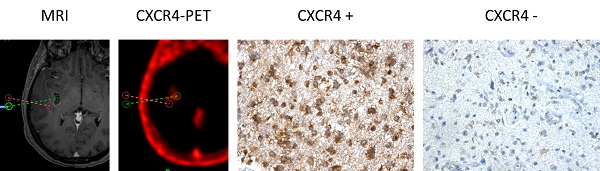
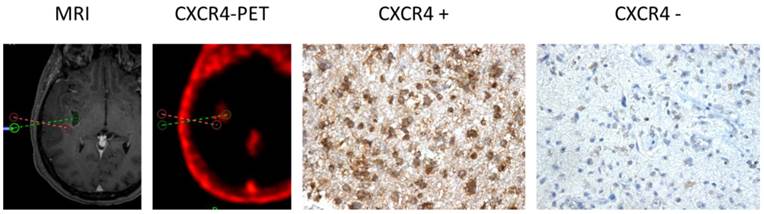
CXCR4 Expression in CXCR4+ and CXCR4- tumor samples (as assessed by 68Ga-Pentixafor-PET). Exemplary depiction of immunohistochemical staining for CXCR4 in biopsies from both 68Ga-Pentixafor negative (red dotted line; positive in 18F-FET-PET) and positive (green) tumor areas, respectively (patient #1). 68Ga-Pentixafor-PET/CT and T1-weighted rapid three-dimensional gradient-echo technique (MP-RAGE) MR images were transferred to a neuronavigation system for specific sampling. Nuclei are stained with H&E. Magnification: 200x.
Discussion
The CXCR4/CXCL12 axis plays a fundamental role in the biology of various tumor entities. In gliomas, CXCR4 mRNA and protein expression has been shown to be connected to higher tumor grading [25], earlier recurrence [7] and shorter survival [25]. In our study, we could demonstrate for the first time that non-invasive imaging of CXCR4 in human malignant glioma using 68Ga-Pentixafor is feasible. In 11/13 patients with high-grade glioma, 68Ga-Pentixafor-PET yielded positive results and receptor expression of malignant cells could be confirmed by immunhistochemical work-up. Whereas 68Ga-Pentixafor uptake was not higher or even lower than amino acid transport, -due to the extremely low background activity in 68Ga-Pentixafor-PET- tumor-to-background ratios were very favorable and significantly higher than for 18F-FET. Of note, tumor samples from areas without apparent tracer accumulation demonstrated absent or only low CXCR4-expression, thus confirming specificity of the radiopharmaceutical.
Due to its critical role for proliferation and spread of malignant gliomas, CXCR4 is an attractive target for new treatment approaches. Recently, a therapeutic vector for CXCR4 has been developed by Wester and co-workers [14, 15, 26] and successfully applied in a proof-of-concept study in advanced stage myeloma patients [27]. Given that CXCR4-directed therapy alone [28] or in combination with temozolomide [29] has already yielded promising results, endoradiotherapy with radiolabelled CXCR4-ligands seems a promising new approach for glioblastoma therapy. In this scenario, the main application of 68Ga-Pentixafor would not primarily be tumor delineation but identification of potential candidates for targeted therapy. CXCR4-directed PET/CT should be performed as a first non-invasive tool to select imaging-positive patients as potential candidates. As reports have hinted at a particularly high receptor expression in cancer stem cells [30], anti-CXCR4 treatment strategies might be even more effective than conventional approaches.
Interestingly, in our cohort both inflammatory changes after brain ischemia as well as primary CNS lymphoma presented with 68Ga-Pentixafor uptake. This was accounted for by CXCR4+ macrophages and lymphoma cells, respectively, which both have been demonstrated to overexpress CXCR4 on the cell surface as well [18, 31]. Though a certain influence of non-tumoral cells on the Pentixafor-signal in GBM cannot be excluded, physiologically abundant microglial cells as well as lymphocytes do not generate a relevant 68Ga-Pentixafor signal as can be deducted from the extremely low background values in our cohort. This finding further underlines the versatility of this new tracer in a broad spectrum of pathologies ranging from inflammatory diseases such as autoimmune diseases, cardiac regeneration after acute myocardial infarction and recovery after cerebral stroke to tumorigenesis, cancer cell proliferation and metastasis in more than 30 different tumor entities.
Our results are in line with a recently published study on the feasibility of non-invasive visualization of CXCR4 expression using PET with another 68Ga-labelled imaging agent [32]. In 8 patients with suspected high-grade glioma, the authors could also demonstrate the specific binding of their tracer to the chemokine receptor on the cell membrane. Interestingly, in our cohort the patients with gliomas WHO III° showed rather low (patient #9, anaplastic oligodendroglioma III°) or no (patient #13, anaplastic astrocytoma III°) 68Ga-Pentixafor-uptake. Whereas immunohistochemistry corroborated negative imaging findings in one patient (#13), the remaining Pentixafor-negative patient presented with a high histological receptor expression. The reason for this discordant observation is not fully understood yet, but might be influenced by receptor kinetics and internalization, given that CXCR4 expression at the cell surface is dynamically regulated and receptor internalization is induced by ligand binding. The biological implications of this finding need to be further investigated on in future larger series. In contrast to a published study investigating neuroendocrine neoplasms [33], we could not demonstrate a correlation between Ki-67 and CXCR4 expression in our cohort, though some cases with low Ki-67 expression also demonstrated low Pentixafor uptake (e.g. patients #7, #9, and #11). This observation should also be confirmed in larger series.
Our study has several limitations. First, only a limited number of patients could be included, though it is larger than the series reported with 68Ga-NOTA-NFB before [32]. Staining for stem cell markers such as CD133 to confirm the theory with particularly high receptor expression on the cell surface of glioblastoma stem cells was not performed, thereby heterogeneity of CXCR4 in GBM cannot be addressed. Future studies comparing radiotracer binding with histologic CXCR4 expression need to be conducted. 68Ga-Pentixafor uptake did not correlate with histological receptor expression, maybe due to receptor kinetics and internalization. However, since imaging-negative patients proved to be CXCR4-negative in immunostaining, 68Ga-Pentixafor might serve as readout to identify or exclude potential candidates for CXCR4-directed therapy.
This pilot study is the first report to demonstrate the feasibility of non-invasive imaging of CXCR4 with 68Ga-Pentixafor, a tracer that has already been demonstrated to exhibit excellent imaging characteristics in lymphoproliferative diseases. Given the need for new effective treatment alternatives in GBM, endoradiotherapy with 90Y- or 177Lu-labelled, 68Ga-Pentixafor-derived vectors might be a new option for glioblastoma patients. Further research investigating the underlying mechanistic and biologic implications is therefore highly warranted.
Conclusion
In this pilot study, 68Ga-Pentixafor retention has been observed in the vast majority of glioblastoma lesions and served as readout for non-invasive visualization of intracerebral CXCR4 expression. Given the paramount importance of the CXCR4/SDF-1 axis in tumor biology, 68Ga-Pentixafor-PET/CT might prove a useful (theranostic) tool for sensitive, non-invasive in-vivo quantification of CXCR4 (tumor phenotyping), and, potentially, prognostication and selection of patients who might benefit from CXCR4-directed therapy.
Supplementary Material
Supplementary Figure 1.
Conflicts of interest
SKr is CEO of Scintomics. HJW is founder and shareholder of Scintomics. All other authors declare no conflicts of interest.
References
1. Ostrom QT, Gittleman H, Farah P, Ondracek A, Chen Y, Wolinsky Y. et al. CBTRUS statistical report: Primary brain and central nervous system tumors diagnosed in the United States in 2006-2010. Neuro-oncology. 2013;15(Suppl 2):ii1-56
2. Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ. et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. The New England journal of medicine. 2005;352:987-96
3. Zlotnik A, Burkhardt AM, Homey B. Homeostatic chemokine receptors and organ-specific metastasis. Nature reviews Immunology. 2011;11:597-606
4. Jacobson O, Weiss ID. CXCR4 chemokine receptor overview: biology, pathology and applications in imaging and therapy. Theranostics. 2013;3:1-2
5. Zhao H, Guo L, Zhao H, Zhao J, Weng H, Zhao B. CXCR4 over-expression and survival in cancer: a system review and meta-analysis. Oncotarget. 2015;6:5022-40
6. Domanska UM, Kruizinga RC, Nagengast WB, Timmer-Bosscha H, Huls G, de Vries EG. et al. A review on CXCR4/CXCL12 axis in oncology: no place to hide. European journal of cancer. 2013;49:219-30
7. Tabouret E, Tchoghandjian A, Denicolai E, Delfino C, Metellus P, Graillon T. et al. Recurrence of glioblastoma after radio-chemotherapy is associated with an angiogenic switch to the CXCL12-CXCR4 pathway. Oncotarget. 2015;6:11664-75
8. Bian XW, Yang SX, Chen JH, Ping YF, Zhou XD, Wang QL. et al. Preferential expression of chemokine receptor CXCR4 by highly malignant human gliomas and its association with poor patient survival. Neurosurgery. 2007;61:570-8 discussion 8-9
9. Lv B, Yang X, Lv S, Wang L, Fan K, Shi R. et al. CXCR4 Signaling Induced Epithelial-Mesenchymal Transition by PI3K/AKT and ERK Pathways in Glioblastoma. Molecular neurobiology. 2014
10. Rubin JB, Kung AL, Klein RS, Chan JA, Sun Y, Schmidt K. et al. A small-molecule antagonist of CXCR4 inhibits intracranial growth of primary brain tumors. Proceedings of the National Academy of Sciences of the United States of America. 2003;100:13513-8
11. Yao C, Li P, Song H, Song F, Qu Y, Ma X. et al. CXCL12/CXCR4 Axis Upregulates Twist to Induce EMT in Human Glioblastoma. Molecular neurobiology. 2015
12. Uy GL, Rettig MP, Motabi IH, McFarland K, Trinkaus KM, Hladnik LM. et al. A phase 1/2 study of chemosensitization with the CXCR4 antagonist plerixafor in relapsed or refractory acute myeloid leukemia. Blood. 2012;119:3917-24
13. Kuhne MR, Mulvey T, Belanger B, Chen S, Pan C, Chong C. et al. BMS-936564/MDX-1338: a fully human anti-CXCR4 antibody induces apoptosis in vitro and shows antitumor activity in vivo in hematologic malignancies. Clinical cancer research: an official journal of the American Association for Cancer Research. 2013;19:357-66
14. Demmer O, Gourni E, Schumacher U, Kessler H, Wester HJ. PET imaging of CXCR4 receptors in cancer by a new optimized ligand. ChemMedChem. 2011;6:1789-91
15. Herrmann K, Lapa C, Wester HJ, Schottelius M, Schiepers C, Eberlein U. et al. Biodistribution and radiation dosimetry for the chemokine receptor CXCR4-targeting probe 68Ga-pentixafor. Journal of nuclear medicine: official publication, Society of Nuclear Medicine. 2015;56:410-6
16. Gourni E, Demmer O, Schottelius M, D'Alessandria C, Schulz S, Dijkgraaf I. et al. PET of CXCR4 expression by a (68)Ga-labeled highly specific targeted contrast agent. Journal of nuclear medicine: official publication, Society of Nuclear Medicine. 2011;52:1803-10
17. Philipp-Abbrederis K, Herrmann K, Knop S, Schottelius M, Eiber M, Luckerath K. et al. In vivo molecular imaging of chemokine receptor CXCR4 expression in patients with advanced multiple myeloma. EMBO molecular medicine. 2015;7:477-87
18. Wester HJ, Keller U, Schottelius M, Beer A, Philipp-Abbrederis K, Hoffmann F. et al. Disclosing the CXCR4 expression in lymphoproliferative diseases by targeted molecular imaging. Theranostics. 2015;5:618-30
19. Martin R, Juttler S, Muller M, Wester HJ. Cationic eluate pretreatment for automated synthesis of [(6)(8)Ga]CPCR4.2. Nuclear medicine and biology. 2014;41:84-9
20. Lapa C, Linsenmann T, Luckerath K, Samnick S, Herrmann K, Stoffer C. et al. Tumor-associated macrophages in glioblastoma multiforme-a suitable target for somatostatin receptor-based imaging and therapy? PloS one. 2015;10:e0122269
21. Wester HJ, Herz M, Weber W, Heiss P, Senekowitsch-Schmidtke R, Schwaiger M. et al. Synthesis and radiopharmacology of O-(2-[18F]fluoroethyl)-L-tyrosine for tumor imaging. Journal of nuclear medicine: official publication, Society of Nuclear Medicine. 1999;40:205-12
22. Boellaard R. Need for standardization of 18F-FDG PET/CT for treatment response assessments. Journal of nuclear medicine: official publication, Society of Nuclear Medicine. 2011;52(Suppl 2):93S-100S
23. Lapa C, Linsenmann T, Monoranu CM, Samnick S, Buck AK, Bluemel C. et al. Comparison of the amino acid tracers 18F-FET and 18F-DOPA in high-grade glioma patients. Journal of nuclear medicine: official publication, Society of Nuclear Medicine. 2014;55:1611-6
24. Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A. et al. The 2007 WHO classification of tumours of the central nervous system. Acta neuropathologica. 2007;114:97-109
25. Lv S, Sun B, Zhong X, Dai C, Wang W, Ma X. et al. The Clinical Implications of Chemokine Receptor CXCR4 in Grade and Prognosis of Glioma Patients: A Meta-Analysis. Molecular neurobiology. 2015;52:555-61
26. Demmer O, Dijkgraaf I, Schumacher U, Marinelli L, Cosconati S, Gourni E. et al. Design, synthesis, and functionalization of dimeric peptides targeting chemokine receptor CXCR4. Journal of medicinal chemistry. 2011;54:7648-62
27. Lapa C, Knop S, Schirbel A, Osl T, Poschenrieder A, Haenscheid H. et al. First in man experience of CXCR4-directed endoradiotherapy with 177Lu- and 90Y-labelled Pentixather in multiple myeloma patients. Journal of Nuclear Medicine. 2015;56:14
28. Liu SC, Alomran R, Chernikova SB, Lartey F, Stafford J, Jang T. et al. Blockade of SDF-1 after irradiation inhibits tumor recurrences of autochthonous brain tumors in rats. Neuro-oncology. 2014;16:21-8
29. Redjal N, Chan JA, Segal RA, Kung AL. CXCR4 inhibition synergizes with cytotoxic chemotherapy in gliomas. Clinical cancer research: an official journal of the American Association for Cancer Research. 2006;12:6765-71
30. Ehtesham M, Mapara KY, Stevenson CB, Thompson RC. CXCR4 mediates the proliferation of glioblastoma progenitor cells. Cancer letters. 2009;274:305-12
31. Gupta SK, Pillarisetti K, Lysko PG. Modulation of CXCR4 expression and SDF-1alpha functional activity during differentiation of human monocytes and macrophages. Journal of leukocyte biology. 1999;66:135-43
32. Wang Z, Zhang M, Wang L, Wang S, Kang F, Li G. et al. Prospective Study of (68)Ga-NOTA-NFB: Radiation Dosimetry in Healthy Volunteers and First Application in Glioma Patients. Theranostics. 2015;5:882-9
33. Kaemmerer D, Trager T, Hoffmeister M, Sipos B, Hommann M, Sanger J. et al. Inverse expression of somatostatin and CXCR4 chemokine receptors in gastroenteropancreatic neuroendocrine neoplasms of different malignancy. Oncotarget. 2015;6:27566-79
Author contact
![]() Corresponding author: h.j.westerde
Corresponding author: h.j.westerde
 Global reach, higher impact
Global reach, higher impact