13.3
Impact Factor
Theranostics 2017; 7(11):2878-2887. doi:10.7150/thno.19358 This issue Cite
Research Paper
Blood Droplet-Based Cancer Diagnosis via Proteolytic Activity Measurement in Cancer Progression
1. Laboratory of Pharmacogenetics, College of Pharmacy, Kyung Hee University, Seoul 02447, Republic of Korea;
2. Department of Biomedical Engineering, Yonsei University, Wonju 26493, Republic of Korea;
3. School of Public Health, Harvard University, Boston, MA 02115, USA;
4. Department of Hemato-Oncology, Yuseong Sun Hospital, Daejeon 34078, Republic of Korea;
5. Section of Hematology-Oncology, Samsung Medical Center, and Sungkyunkwan University (SKKU) School of Medicine, Seoul 06351, Republic of Korea;
6. Department of Urology, Kangbuk Samsung Hospital, Sungkyunkwan University (SKKU) School of Medicine, Seoul 03181, Republic of Korea;
7. Biomechanics Laboratory, College of Sport Science, Sungkyunkwan University (SKKU), Suwon 16419, Republic of Korea;
8. SKKU Advanced Institute of Nano Technology (SAINT), Sungkyunkwan University (SKKU), Suwon 16419, Republic of Korea.
* These authors (J.W.C. and H.L.) made equal contribution to this work.
Received 2017-1-26; Accepted 2017-3-31; Published 2017-7-8
Abstract
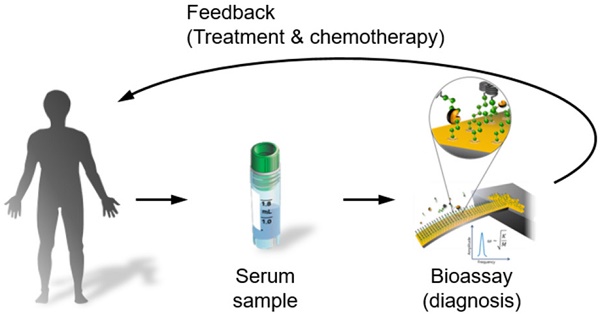
Matrix metalloproteinase (MMP) is a key marker and target molecule for cancer diagnosis, as MMP is able to cleave peptide chains resulting in degradation of extracellular matrix (ECM), a necessary step for cancer development. In particular, MMP2 has recently been recognized as an important biomarker for lung cancer. Despite the important role of detecting MMP molecules in cancer diagnosis, it is a daunting task to quantitatively understand a correlation between the status of cancer development and the secretion level of MMP in a blood droplet. Here, we demonstrate a nanoscale cancer diagnosis by nanomechanical quantitation of MMP2 molecules under cancer progression with using a blood droplet of lung cancer patients. Specifically, we measured the frequency dynamics of nanomechanical biosensor functionalized with peptide chains mimicking ECM in response to MMP2 secreted from tumors in lung with different metastasis level. It is shown that the frequency shift of the biosensor, which exhibits the detection sensitivity below 1 nM, enables the quantitation of the secretion level of MMP2 molecules during the progression of cancer cells or tumor growth. More importantly, using a blood droplet of lung cancer patients, nanomechanical biosensor is shown to be capable of depicting the correlation between the secretion level of MMP2 molecules and the level of cancer metastasis, which highlights the cantilever-based MMP2 detection for diagnosis of lung cancer. Our finding will broaden the understanding of cancer development activated by MMP and allow for a fast and point-of-care cancer diagnostics.
Keywords: proteolysis, matrix metalloproteinase, cancer diagnosis, nanomechanical detection.
Introduction
As matrix metalloproteinase (MMP) has been appreciated as a marker and target molecule for cancer diagnosis [1-3], it is of great importance to characterize the expression level of MMPs and MMP-driven proteolysis (of peptides), which is necessary for enabling the development of prognostic model for cancer patient [3]. In particular, for cancer development (e.g. metastasis), the key process is MMP-driven degradation of extracellular matrix (ECM) circumventing tumors [4, 5], e.g. see Figure 1a, and this ECM degradation is a priori requisite for angiogenesis during cancer progression [6]. Among the members of MMP family, MMP2 has been found to play a role in degrading type IV collagen fibrils [7], which are major component of ECM, indicating that MMP2 is a good marker molecule for cancer diagnosis. Specifically, MMP2 has been recognized as an important marker for lung cancer [8-10], particularly non-small cell lung cancer (NSCLC), which occupies ~85% of lung cancer patients in the United States of America [11].
Though MMP molecules can be detected by conventional bioassays such as immunohistochemistry (IHC) [12], western blot [13], and enzyme-linked immunosorbent assay (ELISA) [14], these bioassays are quite restricted for quantitative characterization of MMP-driven proteolysis during cancer metastasis, which is attributed to their limited detection sensitivity, as well as the inability of antibody used for these bioassays to distinguish MMP from its precursor (i.e. proMMP) [15]. Here, we note that proMMP is unable to cleave peptide chains, which implies that proMMP cannot be a marker for cancer diagnosis due to its incapability of ECM degradation. To overcome the limitation of these bioassays based on antibody, the bioassays using zymography [9, 16, 17] or fluorogenic nanosensor [18-20] have been employed to sense only MMP by detecting the cleavage (i.e. proteolysis) of peptide chains due to MMP. By contrast, they are incapable of real-time quantitation of the MMP-driven proteolysis (i.e. peptide cleavage) during cancer development, which restricts the understanding of prognostic model for cancer patients.
Nanomechanical biosensors such as cantilever sensors have recently been highlighted as a bioassay toolkit that is useful for cancer diagnosis at molecular level [21-24], since they allow for not only the highly sensitive label-free detection of biomolecules such as DNA [25-28], RNA [29], proteins [30-34], and enzymes [35, 36], but also the real-time monitoring of biomolecular interactions [28, 33]. In addition, nanomechanical biosensors have recently been employed for quantitative understanding of the viability of living organisms [37] such as cancer cell [38] and bacteria [39] in response to drug molecules. The basic principle of cantilever-based detection is a direct transduction of biomolecular recognition (occurring on the surface of cantilever biosensor) into the dynamic motion of the biosensor due to biomolecular interaction. Specifically, the biomolecular recognition on the surface of a nanomechanical biosensor leads to the shift of its resonant frequency [21] or its deflection fluctuations [37]. For a recent decade, the frequency shift due to biomolecular interaction has shown to be a useful measure for depicting the kinetics of biomolecular interactions [28] such as proteolysis [36]. In recent years, we have shown that nanomechanical biosensor is able to sensitively detect membrane type-1 MMP [40], which is a marker molecule expressed on the surface of cancer cell. However, to the best of our knowledge, it is a daunting task to quantitatively identify the status of cancer development based on a label-free detection of MMP molecules secreted in a blood droplet of cancer patient. Specifically, a relationship between the secretion level of MMP and cancer metastasis has remained elusive, while an insight into such a relationship is of great importance to develop the prognostic model for cancer patients. Moreover, nanomechanical biosensor-based bioassay has not been implemented using a blood droplet of cancer patient, albeit a blood droplet-based bioassay is necessary for cancer diagnosis at clinical level.
In this paper, we first demonstrate a nanomechanical biosensor-based diagnosis of lung cancer by detecting the peptide cleavage driven by MMP2 secreted in a blood droplet of lung cancer patients with different metastasis level. In particular, we measured the frequency shift of nanomechanical biosensor due to MMP2-driven proteolysis of peptide chains (functionalized on the biosensor's surface) in order to gain insight into a relationship between the secretion level of MMP2 (or equivalently, the amount of cleaved peptide chains due to MMP2) and the level of cancer metastasis. Here, with using the blood droplet of animal model or lung cancer patient, we show that the nanomechanical biosensor allows for quantitative identification of cancer progression by evaluating the amount of peptide chains cleaved by MMP2 under different level of cancer progression. Our study sheds light on the nanomechanical bioassay, which paves the way for developing a fast and point-of-care cancer diagnostics.
Materials and Methods
Theory
We employed continuum elastic model such as Euler-Bernoulli beam model to gain insight into a relationship between the frequency shift of nanomechanical biosensor and the mass of biomolecules that involve in biomolecular interaction (e.g. proteolysis). The relationship between the frequency shift and the change of overall mass for the biosensor due to biomolecular interaction is given by [21]
(1)
where Δω and Δm represent the frequency shift of the biosensor and the change of overall mass for the biosensor, respectively, while ω0 and m0 indicate the resonant frequency of the biosensor and its mass before biomolecular interaction occurs.
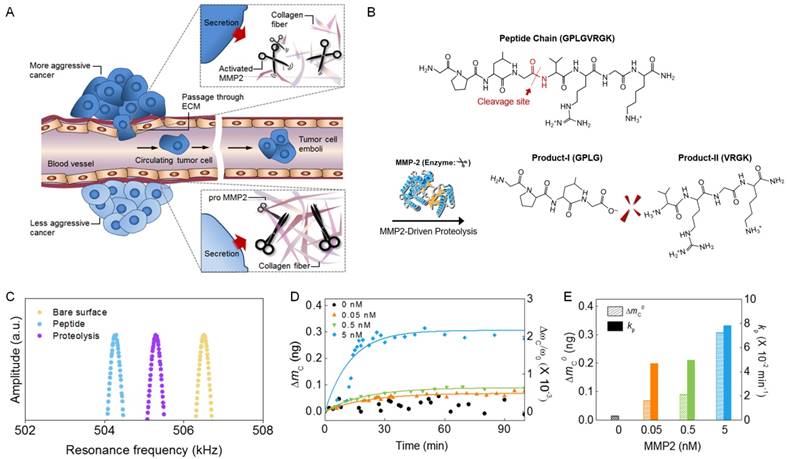
Nanomechanical detection of MMP2 molecules. (A) Schematic illustration of MMP2-driven degradation of ECM due to the ability of MMP2 to cleave peptide chains that are major component of ECM. (B) Chemical structure of peptide chain, which can be cleaved by MMP2. The cleavage site of a peptide chain is indicated by a red arrow. MMP2 cleaves the chemical bond between glycine and valine in the peptide chain. (C) Resonant frequency curves of a bare biosensor (yellow), peptide-functionalized biosensor (blue), and the functionalized biosensor exposed to MMP2 (purple), respectively. (D) In situ measurement of the mass of peptide chains cleaved by MMP2. (E) Dependence of the total mass of cleaved peptide chains and the kinetic rate of proteolysis on MMP2 concentrations.
To quantitate the kinetic rate of proteolysis driven by MMP, we consider the Langmuir kinetic model, which provides a rate equation for proteolysis as follows [36].
(2)
where N(t) is the number of peptide chains that are functionalized on the surface of the nanomechanical biosensor as a function of time t, and kP is the kinetic rate of proteolysis. We note that the time constant of proteolysis is given by τ = 1/kP. The rate equation given by Eq. (2) provides the number of peptide chains that are cleaved by MMP in the form of
(3)
Since the frequency shift of nanomechanical biosensor due to MMP is proportional to the number of cleaved peptide chains, the frequency shift due to MMP-driven proteolysis is given by [36,40]
(4)
This elucidates that Langmuir kinetic model given by Eq. (4) enables the extraction of the kinetic rate of proteolysis (kP) from the in situ measurement of the frequency shift due to MMP as a function of time.
Nanomechanical Bioassay
For label-free detection of MMP2-driven proteolysis, the surface of nanomechanical biosensor was chemically functionalized with peptide chains (whose sequence is shown in Figure 1b) in such a way that PEGylated peptide chains were immobilized on the amine-modified surface of the biosensor. Specifically, the surface of a cantilever biosensor was chemically modified by 3-aminopropyltrimethoxysilane (200 μL) in deionized water (40 mL) at 85 ºC for 24 hr. Subsequently, the amine-modified surface of the biosensor was purified by excessive deionized water and ethanol. Then, in order to immobilize peptide chains on the surface of the biosensor, the amine-functionalized biosensor was immersed in the PEGylated peptide solution for 24 hr at room temperature after adding EDC (3 μmol). The details of bio-functionalization of the biosensor's surface are well described in our previous works [36,40]. Here, we used a commercially available cantilever (Bruker AXS, Madison) with its dimension of 40 × 6.25 × 125 μm3 (width × thickness × length) and its force constant of 200 N/m.
To measure the frequency response of a nanomechanical biosensor, we utilized Biocatalyst (Bruker AXS, Madison), which enabled us to evaluate the resonant frequency of the biosensor in a real-time. For in situ real-time monitoring of MMP2-driven proteolysis, the biosensor (functionalized with peptide chains) was mounted in a liquid cell, that is, a fluid cantilever holder with O-ring (whose volume is ~80 μL). Subsequently, at room temperature, we injected a biological sample (e.g. buffer solution containing MMP2, cell culture medium, or blood droplet) into the liquid cell, and then the resonant frequency of the biosensor was monitored for every 1 min after injecting the biological sample.
Cell Culture
Cancer cell line H460 (human large cell lung carcinoma) and A549 (human alveolar carcinoma) were purchased from American Type Culture Collection, while WI-26 (human fetal lung cell) and H322 (human bronchioaveolar carcinoma) were acquired from Korea Cell Line Bank. For cell cultivation, we used RPMI (for A549 cell line) or Dulbecco's modified eagle's medium (for other cell lines), which contains 10% fetal bovine serum (FBS) and 1% antibiotic. Cells were seeded at ~5 × 105, 106, and 2 × 106 cells/mL, respectively, in 60 mm-sized dishes.
Zymography Assay
Cells were incubated for 24 and 48 h, respectively, and seeded (at 105 cells/mL) in RPMI containing 10% FBS. After starving the cells in serum-free RPMI for 2 h, cell culture medium (20 mL) was mixed with 5× FOD buffer (0.125 Tris-HCl, pH 6.8, containing 5% SDS, 20% glycerol, and 0.01% bromophenol blue) and subjected to 10% SDS-PAGE containing 1 mg/mL gelatin. The gel was washed for each 20 min, followed by incubation with reaction buffer (50 mM Tris-HCl, pH 7.5, containing 10 mM CaCl2, 150 mM NaCl, 1 M ZnCl2, 1% Triton X-100, and 0.002% sodium azide) for 24 h at 37 ºC. The gel was washed with distilled water, and then stained with Coomasie blue R250, followed by destaining with 35% methanol.
Animal Model
All animal tests were conducted under approval of Institutional Research Board (IRB) under IRB No. SMC-2013-02-058 for clinical research at Kangbuk Samsung Hospital. We consider tumor transplantation to mouse model by using two types of cancer cell lines such as H460 and LLC (Lewis lung carcinoma).
H460-transplanted mouse was developed in such a way that 2 × 107 H460 cells were subcutaneously inoculated into 6 week old female nude mice (Orient Bio, Korea). In 10 days after transplantation, the diameter of tumor reaches 3 to 5 mm. This state is referred to as stage I. Then the blood of the mice was collected from 3 mice by cardiac puncture and tumors were isolated. We refer to stage II as 20 days after transplantation into the mice, and the blood sample at this stage was collected.
As LLC tumor originates from a carcinoma of the lung of C57BL/6 mouse, we also considered LLC transplantation into the mice. The procedure of transplantation is similar to the case of H460 transplantation (as described above). In case of LLC transplantation, we define the stage I as 3 weeks after transplantation, while stage II is referred to as 6 weeks after transplantation.
Immunohistochemistry (IHC) Staining
Tumor mass for each tested mouse was isolated, flushed with phosphate buffered saline (PBS), and then mixed with 10% formalin for 24 h. The tissues were then processed with paraffin block. After de-paraffinization and hydration, the tissue slides were stained by hematoxylin and eosin (i.e. H&E staining). IHC staining was accomplished as follows. For antigen retrieval, the slides were incubated with 0.3% H2O2, and then boiled in 0.01 M citrate buffer. After blocking with 4% bovine serum albumin (BSA), antibodies H-76 (purchased from Santacruz biotechnology) against MMP2 were added for 2 h. Then polymer kit (Dako, Denmark) and diaminobenzidine (Dako, Denmark) were added for IHC tissue staining.
Human Translational Research
The blood sera of normal people and lung cancer patients were collected under the approval of IRB under IRB No. SMC-2005-10-024 for clinical research at Samsung Medical Center. Here, 15 patients suffering from lung cancer at stage IV (but different metastasis level) were randomly selected for obtaining the blood serum of cancer patient.
Results
Nanomechanical Detection of MMP2-Driven Proteolysis
To sense MMP2-driven proteolysis, we prepared a nanomechanical biosensor by functionalizing its surface with peptide chains, which can be cleaved by MMP2. The peptide-functionalization of a bare biosensor (with its resonant frequency of 506.3 kHz) decreases its frequency with the amount of ΔωF = 2.23 kHz, which corresponds to the total mass of functionalized peptide chains being ΔmF = 640 pg (Figure 1c). When the functionalized biosensor was exposed to MMP2 molecules, the resonant frequency of the biosensor (vibrating in normal air) is increasing with the amount of ΔωC = 1.01 kHz, which is attributed to the mass of cleaved peptide chains being measured as ΔmC = 293 pg (Figure 1c). For quantitative analysis of the MMP2-driven proteolysis, we consider an in situ real-time monitoring of the frequency shift of a nanomechanical biosensor immersed in a buffer solution due to MMP2 molecules with their concentrations ranging from 0.05 to 5 nM. Figure 1d suggests that the in situ frequency shift of the biosensor due to MMP2 is well dictated by Langmuir kinetic model (see Materials and Methods, and also ref. [36,40]). Figure 1e shows the dependence of the total mass of cleaved peptide chains and the kinetic rate of proteolysis on the concentration of MMP2 molecules. Our nanomechanical detection of MMP2-driven proteolysis was validated by atomic force microscopy (AFM) imaging (see Figure S1), which is useful in visualizing the surface where biomolecular interactions occur [41,42]. Here, we note that after MMP2-driven proteolysis of peptide chain, the N-terminal amino acids (VRGK) remains on the surface of the biosensor, while C-terminal amino acids (GPLG) are cleaved (Figure 1b). This is consistent with AFM images (Figure S1), which show that the peptide-functionalization of a bare biosensor leads to the increase of AFM height into 20 nm (average value) due to the length of peptide chains, while MMP2-driven proteolysis reduces the value of AFM height into 10 nm (average value) owing to the length of cleaved peptide fragments. Moreover, we consider negative control experiments with using the buffer solution containing other MMP family (i.e. MMP3, MMP9, and MMP14) in order to verify that our biosensor is only responsive to MMP2 rather than other MMP family, since MMP2 is an important biomarker for lung cancer [8-10]. The results of negative control experiments show that the biosensor is able to selectively detect only MMP2 rather than other MMP family (Figure S2).
The MMP2-Driven Proteolysis under Cancer Cell Progression
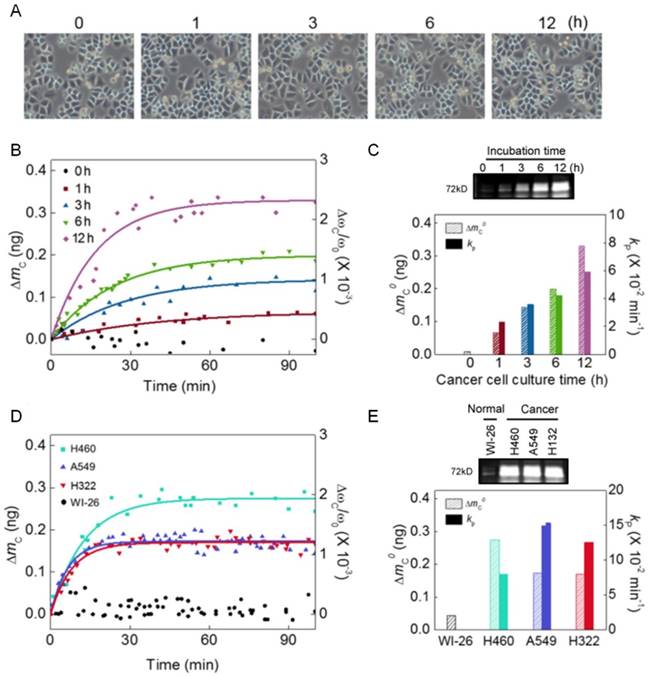
To understand the role of MMP2 in cancer development, we consider the nanomechanical biosensor exposed to cell culture medium, where cancer cells were incubated. Here, we take into account H460 cells with different incubation time from 0 to 12 h. We note that since cellular doubling time is 23 h, our measurement based on incubation time up to 12 h is aimed towards understanding the secretion level of MMP2 molecules released from cancer cells under their progression before cell division occurs. While optical microscope images show that cellular confluency (corresponding to the number of cancer cells) is not increased (Figure 2a), the frequency shift of the nanomechanical biosensor due to MMP2 is increasing with respect to incubation time (Figure 2b). The MMP2 secretion was confirmed by zymography bioassay (see the upper panel of Figure 2c), though this bioassay is unable to quantitate the kinetic rate of MMP2-driven proteolysis. As shown in Figure 2c, the dependence of frequency shift due to MMP2 on the incubation time suggests that during cancer progression, though the number of cancer cells is not increased, the secretion level of MMP2 is likely to increase. It is shown that the total mass of peptide chains (ΔmC0) cleaved by MMP2, which was released from H460 cells, is linearly proportional to the incubation time (T) such as d(ΔmC0)/dT = 25 pg/h.
Moreover, we investigate whether MMP2 secretion is a generic process for cancer progression by considering a nanomechanical biosensor that was exposed to cell culture media, where different types of cancer cells such as H460, A549, and H322, respectively, were incubated for 12 h. We observe the frequency shifts of nanomechanical biosensors when they were exposed to aforementioned cell culture media, respectively, which confirms the MMP2 secretion from these cancer cell lines. It is shown that nanomechanical biosensor does not respond to the cell culture medium, in which normal cells such as WI-26 cells were incubated. This provides an evidence that MMP2 secretion does not occur for normal cells. We found that the total mass of peptide chains cleaved by MMP2 secreted from H460 cells is measured as ~300 pg, which is larger than that (i.e. ~200 pg) from other cancer cell lines (Figure 2d and e). This observation suggests that MMP2 secretion is a generic process for cancer progression [43], while the secretion level of MMP2 molecules is more remarkable for lung cancer.
Correlation between MMP2 Activity and Tumor Growth State
To explore the potential of nanomechanical biosensor for cancer diagnosis, we focus on the frequency dynamics of the biosensor in response to MMP2 that is likely to be secreted from tumors appearing in animal model. In particular, we used the blood droplet of animal model, where tumor cells such as H460 or LLC cells were injected. The details of animal model are described in Materials and Methods, and the results of nanomechanical bioassay using LLC-transplanted animal model are presented in Figure S3. Here, we concentrate on the results of nanomechanical bioassay using H460-transplanted animal model.
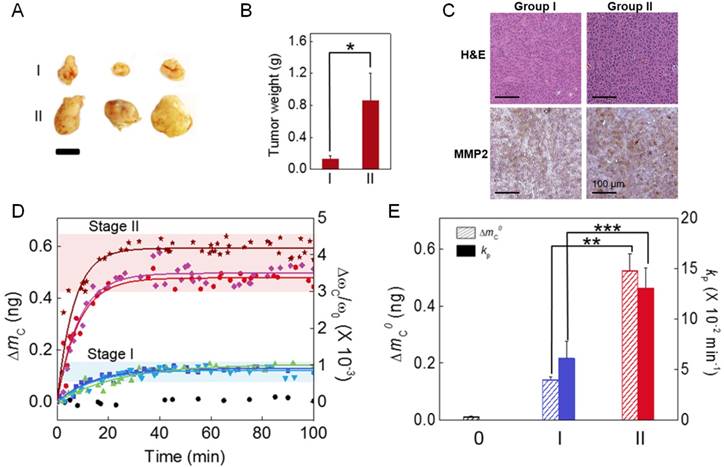
We categorize the H460-transplanted animal models into two groups (i.e. stage I and II) according to tumor size. The weight of tumors at stage I and II is measured as ~0.2 and ~0.8 g, respectively (Figure 3a and b). H&E staining shows that cellular density and necrotic region are similar between stage I and II (see the upper panel of Figure 3c), whereas IHC experiment qualitatively suggests that MMP2 seems to be vigorously secreted for stage II when compared with stage I (see the lower panel of Figure 3c). To characterize the peptide cleavage driven by MMP2 secreted from tumors at two different tumor growth states, we consider the frequency dynamics of nanomechanical biosensor in response to injection of blood droplet (obtained from animal model) into a liquid cell, in which the biosensor was mounted. Figure 3d depicts that the frequency shift of a nanomechanical biosensor depends on the tumor growth state, which underlies that MMP2 is a useful marker to identify the tumor growth state. It is shown that the total mass of cleaved peptide chains and the kinetic rate of proteolysis are measured as ΔmC0 = 148 pg and kP = 6.12 × 10-2 min-1 for stage I, respectively, while they are evaluated as ΔmC0 = 548 pg and kP = 13.07 × 10-2 min-1 for stage II. Our results validate the potential of nanomechanical biosensor for quantitative identification of tumor growth state, which implies that nanomechanical bioassay allows for quantitative understanding of cancer development.
Nanomechanical detection of MMP2 molecules secreted from cancer cells during their progression. (A) Optical microscope images of cancer cells incubated at 1 to 12 h. (B) The mass of cleaved peptide chains (measured from the frequency shift) due to MMP2 secreted from H460 cells incubated at 1 to 12 h. (C) MMP2 secretion was confirmed by zymography bioassay (upper panel). The total mass of cleaved peptide chains and the kinetic rate of proteolysis are shown to depend on the cell incubation time (lower panel). (D) The mass of cleaved peptide chains due to MMP2 molecules that are likely to be secreted from different types of cancer cells such as H460, A549, and H322, respectively, incubated at 12 h. (E) The MMP2 secretion from different types of cancer cell lines was verified by zymography bioassay (upper panel). The total mass of cleaved peptide chains and the kinetic rate of proteolysis are found to depend on the type of cancer cell lines (lower panel).
Relationship between the secretion level of MMP2 and tumor growth state. (A) Photographic images of tumors at stage I and II, respectively. Scale bar indicates 10 mm. (B) The weight of tumors at these two different stages. (C) H&E images (upper panel) and IHC images (lower panel) for tissues at stages I and II, respectively. (D) In situ measurement of the mass of cleaved peptide chains due to MMP2 for two different tumor growth states. (E) Dependence of the total mass of cleaved peptide chains and the kinetic rate of proteolysis on the tumor growth state.
Blood Droplet-Based Cancer Diagnosis: Proteolytic Activity in Cancer Metastasis
To evaluate the potential of nanomechanical bioassay for blood droplet-based cancer diagnosis, we monitored the frequency dynamics of a nanomechanical biosensor in response to injection of a blood droplet obtained from patients suffering from lung cancer, i.e. NSCLC (see Figure 4a). Here, the blood samples were acquired from 15 patients suffering from NSCLC at the same stage IV but different metastasis level (Figure S4). The negative control experiments were taken into account using a blood droplet obtained from 6 normal people (Figure S5), which confirms that MMP2 secretion does not occur for normal people. By contrast, we observe the frequency shift of the nanomechanical biosensor in response to injection of a blood droplet acquired from 15 lung cancer patients. The in situ frequency shifts of the biosensor due to MMP2 molecules secreted from tumors for all 15 lung cancer patients are presented in Figure S6. The total mass of peptide chains cleaved by MMP2 is measured in a range of 150 to 600 pg, and the kinetic rate of proteolysis is estimated in a range between 4 × 10-2 and 11 × 10-2 min-1 (Figure 4b). The variation of these values is attributed to the different level of cancer metastasis (see below). The average time constant of MMP2-driven proteolysis is evaluated as τ = 14.59 min, which alludes that nanomechanical cancer diagnosis can be implemented within few hours leading to acute and fast cancer diagnosis.
The total mass of cleaved peptide chains is a useful measure in understanding the secretion level of MMP2 molecules released from tumors, which leads to the quantitation of tumor growth state. The total mass of cleaved peptide chains is measured as 600 pg for lung cancer patients #11 and #12 (Figure S6), which is comparable to the mass of peptide chains cleaved by MMP2 secreted from tumors at stage II in animal model (e.g. Figure 3c). This implies that the expression level of MMP2 molecules in animal model at stage II may be similar to that for the blood sample of lung cancer patients #11 and #12. However, for lung cancer patients #08 and #09, the total mass of cleaved peptide chains is estimated as 150 pg, which is comparable to that for tumor growth stage I in animal model. This observation suggests that even though 15 patients suffer from lung cancer at stage IV, the expression level of MMP2 molecules may be different with dependence on the size of tumor. That is, the variation of the values of the total mass of cleaved peptide chains may be attributed to the different size of tumors under different level of metastasis (see below).
Blood droplet-based nanomechanical cancer diagnosis. (A) Schematic illustration of nanomechanical cancer diagnosis using a blood droplet of cancer patient. The lung cancer was indicated by red arrows in the computed tomography images. When the blood droplet of cancer patient was injected into a liquid cell, where a nanomechanical biosensor was mounted, its frequency dynamics was monitored in order to quantitate MMP2-driven proteolysis. (B) The total mass of cleaved peptide chains (indicated by green open-circular dots) and the kinetic rate of proteolysis (shown as red open-triangular dots) for cancer patients. (C) The total mass of cleaved peptide chains with respect to the level of distant metastasis. (D) The kinetic rate of proteolysis as a function of the level of distant metastasis. (E) The total mass of cleaved peptide chains with respect to the degree of spread to regional lymph node. (F) The kinetic rate of proteolysis as a function of the degree of spread to regional lymph node. Here, it should be noted that red-colored abscissa indicates the average value of the quantity such as the total mass of cleaved peptide chains or the kinetic rate of proteolysis.
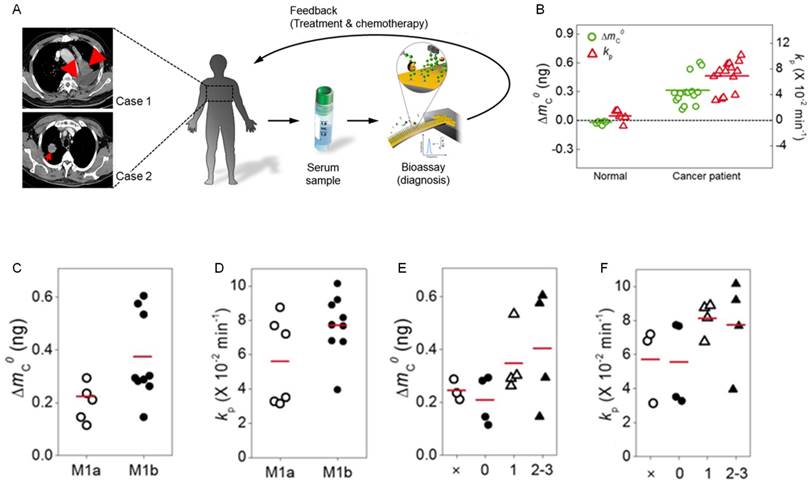
We further explore the ability of nanomechanical bioassay to identify the level of cancer metastasis. In particular, we investigate whether the frequency response of nanomechanical biosensor to MMP2 is correlated with the level of cancer metastasis. Here, in order to describe the level of cancer metastasis, we consider two types of classifications - (i) the level of distant metastasis (abbreviated as M), and (ii) the degree of spread to regional lymph node (denoted as N). We found that the average values of the total mass of cleaved peptide chains and the kinetic rate of proteolysis are measured as ΔmC0 = 226 pg and kP = 5.58 × 10-2 min-1 for stage M1a, respectively, while they are evaluated as ΔmC0 = 348 pg and kP = 7.7 × 10-2 min-1 for stage M1b (Figure 4c and d). This indicates that the proteolytic activity of MMP2 is highly correlated with the level of distant metastasis. In addition, the variation of the total mass of cleaved peptide chains is found to be larger for stage M1b than that for stage M1a, which implies that the variation of tumor sizes may be larger for stage M1b than stage M1a. It should be noted that the largest value of the total mass of cleaved peptide chains at stage M1a is less than 348 pg (that is the average value of the total mass of cleaved peptide chains at stage M1b). The value of 600 pg for the total mass of cleaved peptide chains corresponds only to 2 patients suffering from stage M1b (Figure 4c). This observation suggests that the total mass of cleaved peptide chains is a useful measure for quantitating the secretion level of MMP2 with a relation to the level of distant metastasis. In a similar manner, the total mass of cleaved peptide chains and the kinetic rate of proteolysis are shown to be critically dependent on the degree of spread to regional lymph node (Figure 4e and f). In particular, we found that the value of 600 pg for the total mass of cleaved peptide chains corresponds only to 2 patients suffering from N2-3 stage. It is shown that the largest value of the total mass of cleaved peptide chains for Nx stage is still less than the average value of the total mass of cleaved peptide chains at N1 and N2-3 stages. This suggests that the average value of the total mass of cleaved peptide chains is a useful quantity in identifying the degree of spread to regional lymph node. These results highlight the important role of measuring the level of MMP2-driven proteolysis in lung cancer diagnosis.
Discussion
In this work, we first demonstrate the nanomechanical biosensor-based cancer diagnosis by a label-free detection of MMP2 molecules secreted from tumors appearing in lung cancer patients under different level of cancer metastasis. We show that the frequency dynamics of nanomechanical biosensor in response to MMP2 is a useful measure in quantitating the proteolytic activity of MMP2 in a relation to the level of cancer metastasis. Specifically, the mass of cleaved peptide chains, which can be calculated from the frequency shift of the biosensor due to MMP2, is found to be correlated with the level of cancer metastasis. Here, we note that our nanomechanical biosensor exhibits the high detection sensitivity for sensing the secretion level of MMP molecules. The detection sensitivity of the nanomechanical biosensor is 0.05 nM (e.g. see Figure 1d), which is better than the detection sensitivity of other methods such as fluorogenic nanosensor (2 nM) [19], ELISA (0.5 nM) [14], and zymography (100 nM) [9].
Moreover, our work can be further extended for in vivo evaluation of drug efficacy by measuring the frequency response of nanomechanical biosensor to MMP2, whose activity can be regulated by drug injected into animal model or cancer patient, as some drugs have recently been designed to directly target MMP molecules [44, 45]. In particular, as shown in our previous study [40], the treatment of tumors with drug molecules directly targeting MMP will reduce the expression level of MMP, which implies that the total mass of cleaved peptide chains will decrease with drug treatment. The amount of decrease in the total mass of cleaved peptide chains due to drug will provide a quantitative insight into how drug molecules effectively regulate the secretion level of MMP molecules released from tumors. In addition, as various types of MMP molecules are important biomarkers for various types of cancers, the multiplexed detection of MMP molecules is of great importance for cancer diagnosis [46]. For our future study, we will consider the cantilever biosensor array [25,34] for the multiplexed detection of various types of MMP molecules released from tumors. Furthermore, MMP is also a key marker for inflammation such as pulmonary emphysema [47]. Under inflammation condition, the over-expression of MMP molecules is likely to happen. This suggests that our work can be also applicable for studying the expression level of MMP molecules released from macrophage under inflammation condition by measuring the mass of peptide chains cleaved by MMP under different inflammation conditions.
In conclusion, we first suggest the nanomechanial MMP detection-based diagnosis of lung cancer under different metastasis level. Our study implies that nanomechanical bioassay, which is able to quantitatively characterize the status of cancer development with using a blood droplet of cancer patients, may pave the way for developing a prognostic model for cancer patients. Our work sheds light on nanomechanical bioassay that can help to understand the cancer development activated by MMP as well as to develop a fast and point-of-care cancer diagnostics.
Acknowledgements
T.K. appreciates the financial support from the National Research Foundation of Korea (NRF) under Grant No. NRF-2015R1A2A1A15052758 and NRF-2016R1D1A1B03933719. K.E. gratefully acknowledges the financial support from NRF under Grant No. NRF-2015R1A2A2A04002453.
Supplementary Material
Figure S1 - S6.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Egebald M, Werb Z. New functions for the matrix metalloproteinases in cancer progression. Nat Rev Cancer. 2002;2:161-74
2. Chiang AC, Massague J. Molecular basis of metastasis. N Eng J Med. 2008;359:2814-23
3. Coussens LM, Fingleton B, Matrisian LM. Matrix metalloproteinase inhibitors and cancer: trials and tribulatioins. Science. 2002;295:2387-92
4. Woessner JF, Nagase H. Matrix Metalloproteinases and TIMPs. New York: Oxford University Press. 2000
5. McCawley LJ, Matrisian LM. Matrix metalloproteinases; multifunctional contributors to tumor progression. Mol Med Today. 2000;6:149-56
6. McCawley LJ, Matrisian LM. Matrix metalloproteinases; they're not just for matrix anymore!. Curr Opin Cell Biol. 2001;13:534-40
7. Mahajan N, Dhawan V, Mahmood S, Malik S, Jain S. Extracellular matrix remodeling in Takayasu's arteritis; role of matrix metalloproteinases and adventitial inflammation. Arch Med Res. 2012;43:406-10
8. Qian Q, Wang Q, Zhan P, Peng L, Wei S-Z, Shi Y, Song Y. The role of matrix metalloproteinase 2 on the survival of patients with non-small cell lung cancer: a systematic review with meta-analysis. Cancer Investig. 2010;28:661-9
9. Garbisa S, Scagliotti G, Masiero L, Di Francesco C, Caenazzo C, Onisto M, Micela M, Stetler-Stevenson WG, Liotta LA. Correlation of serum metalloproteinase levels with lung cancer metastasis and response to therapy. Cancer Res. 1992;52:4548-9
10. Passlick B, Sienel W, Seen-Hibler R, Wockel W, Thetter O, Mutschler W, Pantel K. Overexpression of matrix metalloproteinase 2 predicts unfavorable outcome in early-stage non-small lung cancer. Clin Cancer Res. 2000;6:3944-8
11. Molina JR, Yang P, Cassivi SD, Schild SE, Adjei AA. Non-small cell lung cancer; epidemiology, risk factors, treatment, and survivorship. Mayo Clinc Proc. 2008;83:584-94
12. Brehmer B, Biesterfeld S, Jakse G. Expression of matrix metalloproteinases (MMP-2 and -9) and their inhibitors (TIMP-1 and -2) in prostate cancer tissue. Prostate Cancer P D. 2003;6:217-22
13. Daja MM, Niu X, Zhao Z, Brown JM, Russell PJ. Characterization of matrix metalloproteinases and tissue inhibitors of metalloproteinases in prostate cancer cell line. Prostate Cancer P D. 2003;6:15-26
14. Ylisirnio S, Hoyhtya M, Turpeenniemi-Hujanen T. Serum matrix metalloproteinases -2, -9 and tissue inhibitors of metalloproteinases -1, -2 in lung cancer; TIMP-1 as prognostic marker. Anticancer Res. 2000;20:1311-6
15. Brown PD, Bloxidge RE, Stuart NSA, Galler KC, Carmichael J. Association between expression of activated 72-kilodalton gelatinase and tumor spread in non-small-cell lung carcinoma. J Natl Cancer Inst. 1993;85:574-8
16. Kleiner DE, Stetlerstevenson WG. Quantitative zymography; detection of picogram quantities of gelatinases. Anal Biochem. 1994;218:325-9
17. Wilson MJ, Casey C, Woodson M, Sinha AA. Reverse zymography studies of protease inhibitors in the secretions of different lobes of rat prostate. Arch Andrology. 1999;42:109-18
18. Ouyang M, Lu S, Li X-Y, Xu J, Seong J, Giepmans BNG, John Y-JS, Weiss SJ, Wang Y. Visualization of polarized membrane type 1 matrix metalloproteinase activity in live cells by fluorescence resonance energy transfer imaging. J Biol Chem. 2008;283:17740-8
19. Lee S, Ryu JH, Park K, Lee A, Lee S-Y, Youn I-C, Ahn C-H, Yoon SM, Myung S-J, Moon DH, Chen X, Choi K, Kwon IC, Kim K. Polymeric nanoparticle-based activatable near-infrared nanosensor for protease determination in vivo. Nano Lett. 2009;9:4412-6
20. Lee S, Cha E-J, Park K, Lee S-Y, Hong J-K, Sun I-C, Kim SY, Choi K, Kwon IC, Kim K, Ahn C-H. A near-infrared-fluorescence-quenched gold-nanoparticle imaging probe for in vivo drug screening and protease activity determination. Angew Chem Int Ed. 2008;47:2804-7
21. Eom K, Park HS, Yoon DS, Kwon T. Nanomechanical resonators and their applications in biological/chemical detection; nanomechanics principles. Phys Rep. 2011;503:115-63
22. Arlett JL, Myers EB, Roukes ML. Comparative advantages of mechanical biosensors. Nat Nanotech. 2011;6:203-15
23. Boisen A, Dohn S, Keller SS, Schmid S, Tenje M. Cantilever-like micromechanical sensors. Rep Prog Phys. 2011;74:036101
24. Waggoner PS, Craighead HG. Micro- and nanomechanical sensors for environmental, chemical, and biological detection. Lab Chip. 2007;7:1238-55
25. Fritz J, Baller MK, Lang HP, Rothuizen H, Vettiger P, Meyer E, Guntherodt HJ, Gerber C, Gimzewski JK. Translating biomolecular recognition into nanomechanics. Science. 2000;288:316-8
26. McKendry R, Zhang JY, Arntz Y, Strunz T, Hegner M, Lang HP, Baller MK, Certa U, Meyer E, Guntherodt HJ, Gerber C. Multiple label-free biodetection and quantitative DNA-binding assays on a nanomechanical cantilever array. Proc Natl Acad Sci USA. 2002;99:9783-8
27. Wu GH, Ji HF, Hansen K, Thundat T, Datar R, Cote R, Hagan MF, Chakraborty AK, Majumdar A. Origin of nanomechanical cantilever motion generated from biomolecular interactions. Proc Natl Acad Sci USA. 2001;98:1560-4
28. Kwon T, Eom K, Park J, Yoon DS, Lee HL, Kim TS. Micromechanical observation of the kinetics of biomolecular interactions. Appl Phys Lett. 2008;93:173901
29. Zhang J, Lang HP, Huber F, Bietsch A, Grange W, Certa U, McKendry R, Guntherodt HJ, Hegner M, Gerber C. Rapid and label-free nanomechanical detection of biomarker transcripts in human RNA. Nat Nanotech. 2006;1:214-20
30. Wu GH, Datar RH, Hansen KM, Thundat T, Cote RJ, Majumdar A. Bioassay of prostate-specific antigen (PSA) using microcantilevers. Nat Biotech. 2001;19:856-60
31. Mukhopadhyay R, Sumbayev VV, Lorentzen M, Kjems J, Andreasen PA, Besenbacher F. Cantilever sensor for nanomechanical detection of specific protein conformations. Nano Lett. 2005;5:2385-8
32. Gruber K, Horlacher T, Castelli R, Mader A, Seeberger PH, Hermann BA. Cantilever array sensors detect specific carbohydrate-protein interactions with picomolar sensitivity. ACS Nano. 2011;5:3670-8
33. Kwon TY, Eom K, Park JH, Yoon DS, Kim TS, Lee HL. In situ real-time monitoring of biomolecular interactions based on resonating microcantilevers immersed in a viscous fluid. Appl Phys Lett. 2007;90:223903
34. Yue M, Stachowiak JC, Lin H, Datar R, Cote R, Majumdar A. Label-free protein recognition two-dimensional array using nanomechanical sensors. Nano Lett. 2008;8:520-4
35. Weizmann Y, Elnathan R, Lioubashevski O, Willner I. Endonuclease-based logic gates and sensors using magnetic force-amplified readout of DNA scission on cantilevers. J Am Chem Soc. 2005;127:12666-72
36. Kwon T, Park J, Yang J, Yoon DS, Na S, Kim C-W, Suh JS, Huh YM, Haam S, Eom K. Nanomechanical in situ monitoring of proteolysis of peptide by Cathepsin B. PLoS ONE. 2009;4:e6248
37. Kasas S, Ruggeri FS, Banadiba C, Maillard C, Stupar P, Tournu H, Dietler G, Longo G. Detecting nanoscale vibrations as signature of life. Proc Natl Acad Sci USA. 2015;112:378-81
38. Wu S, Liu X, Zhou X, Liang XM, Gao D, Liu H, Zhao G, Zhang Q, Wu X. Quantification of cell viability and rapid screening anti-cancer drug utilizing nanomechanical fluctuation. Biosens Bioelectron. 2016;77:164-73
39. Longo G, Alonso-Sarduy L, Rio LM, Bizzini A, Trampuz A, Notz J, Dietler G, Kasas S. Rapid detection of bacterial resistance to antibiotics using AFM cantilevers as nanomechanical sensors. Nat Nanotech. 2013;8:522-6
40. Lee G, Eom K, Park J, Yang J, Haam S, Huh Y-M, Ryu JK, Kim NH, Yook JI, Lee SW, Yoon DS, Kwon T. Real-time quantitative monitoring of specific peptide cleavage by a proteinase for cancer diagnosis. Angew Chem Int Ed. 2012;51:5837-41
41. Park J, Yang J, Lee G, Lee CY, Na S, Lee SW, Haam S, Huh Y-M, Yoon DS, Eom K, Kwon T. Single-molecule recognition of biomolecular interaction via Kelvin probe force microscopy. ACS Nano. 2011;5:6981-90
42. Sinensky AK, Belcher AM. Label-free and high-resolution protein/DNA nanoarray analysis using Kelvin probe force microscopy. Nat Nanotech. 2007;2:653-9
43. Vihinen P, Kähäri V-M. Matrix metalloproteinases in cancer; Prognostic markers and therapeutic targets. Int J Cancer. 2002;99:157-66
44. Fisher JF, Mobashery S. Recent advances in MMP inhibitor design. Cancer Metastasis Rev. 2006;25:115-36
45. Kang S-g, Zhou G, Yang P, Liu Y, Sun B, Huynh T, Meng H, Zhao L, Xing G, Chen C, Zhao Y, Zhou R. Molecular mechanism of pancreatic tumor metastasis inhibition by Gd@C82(OH)22 and its implication for de novo design of nanomedicine. Proc Natl Acad Sci USA. 2012;109:15431-6
46. Kwong GA, von Maltzahn G, Murugappan G, Abudayyeh O, Mo S, Papayannopoulos IA, Sverdlov DY, Liu SB, Warren AD, Popov Y, Schuppan D, Bhatia SN. Mass-encoded synthetic biomarkers for multiplexed urinary monitoring of disease. Nat Biotech. 2013;31:63-70
47. Cobos-Correa A, Trojanek JB, Diemer S, Mall MA, Schultz C. Membrane-bound FRET probe visualizes MMP12 activity in pulmonary inflammation. Nat Chem Biol. 2009;5:628-30
Author contact
![]() Corresponding authors: taeyunkwonedu (T.K.) and kilhoeomedu (K.E.)
Corresponding authors: taeyunkwonedu (T.K.) and kilhoeomedu (K.E.)
 Global reach, higher impact
Global reach, higher impact