13.3
Impact Factor
Theranostics 2017; 7(11):2888-2899. doi:10.7150/thno.19425 This issue Cite
Research Paper
A Normalization-Free and Nonparametric Method Sharpens Large-Scale Transcriptome Analysis and Reveals Common Gene Alteration Patterns in Cancers
1. State Key Laboratory of Genetic Resources and Evolution, Kunming Institute of Zoology, Chinese Academy of Sciences, Kunming 650223, China;
2. Key Laboratory of Animal Models and Human Disease Mechanisms, Kunming Institute of Zoology, Chinese Academy of Sciences, Kunming 650223, China;
3. KIZ/CUHK Joint Laboratory of Bioresources and Molecular Research in Common Diseases, Kunming 650223, China;
4. Department of Pathology, the Second Xiangya Hospital, Central South University, Changsha 410013, China;
5. Farm Animal Genetic Resources Exploration and Innovation Key Laboratory of Sichuan Province, Sichuan Agricultural University, Chengdu 611130, China;
6. Laboratory for Conservation and Utilization of Bio-Resources, Yunnan University, Kunming 650091, China;
7. School of Life Sciences, University of Science and Technology of China, Hefei 230027, China;
8. Key Laboratory of Computational Biology, CAS-MPG Partner Institute for Computational Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai 200031, China;
9. Kunming College of Life Science, University of Chinese Academy of Sciences, Beijing 100049, China.
* These authors contributed equally to this work.
Abstract
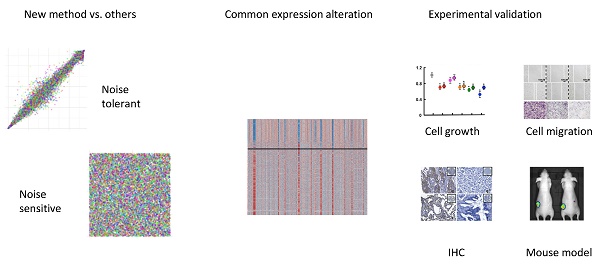
Heterogeneity in transcriptional data hampers the identification of differentially expressed genes (DEGs) and understanding of cancer, essentially because current methods rely on cross-sample normalization and/or distribution assumption—both sensitive to heterogeneous values. Here, we developed a new method, Cross-Value Association Analysis (CVAA), which overcomes the limitation and is more robust to heterogeneous data than the other methods. Applying CVAA to a more complex pan-cancer dataset containing 5,540 transcriptomes discovered numerous new DEGs and many previously rarely explored pathways/processes; some of them were validated, both in vitro and in vivo, to be crucial in tumorigenesis, e.g., alcohol metabolism (ADH1B), chromosome remodeling (NCAPH) and complement system (Adipsin). Together, we present a sharper tool to navigate large-scale expression data and gain new mechanistic insights into tumorigenesis.
Keywords: Cross-Value Association Analysis, normalization-free, pan-cancer, transcriptome, heterogeneity.
 Global reach, higher impact
Global reach, higher impact