Impact Factor
Theranostics 2017; 7(12):3106-3117. doi:10.7150/thno.19016 This issue Cite
Review
Circular RNAs: Regulators of Cancer-Related Signaling Pathways and Potential Diagnostic Biomarkers for Human Cancers
1. Bone and Soft Tissue Tumors Research Center of Yunnan Province, Department of Orthopaedics, the Third Affiliated Hospital of Kunming Medical University (Tumor Hospital of Yunnan Province), Kunming, Yunnan 650118, China;
2. Department of Medical Oncology, The Third Affiliated Hospital of Kunming Medical University (Tumor Hospital of Yunnan Province), Kunming, Yunnan 650118, China;
3. State Key Laboratory of Protein and Plant Gene Research, College of Life Sciences, Peking University, Beijing, China.
* These authors contributed equally to this work and should be considered as co-first authors.
Received 2017-1-2; Accepted 2017-5-7; Published 2017-7-22
Abstract
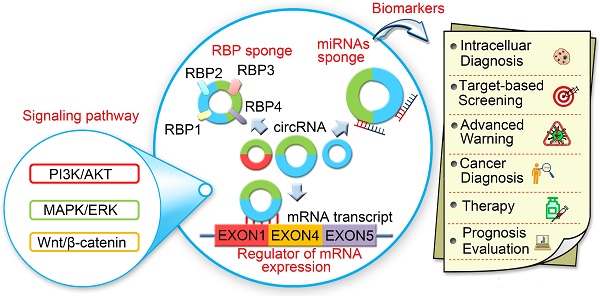
Circular RNAs (circRNAs) are newly discovered endogenous non-coding RNAs featuring structural stability, high abundance, and tissue-specific expression. CircRNAs are prevalent and conserved in mammalian cells. They are involved in cellular processes and regulate gene expression at the transcriptional or post-transcriptional level by interacting with microRNAs (miRNAs) and other molecules. Recent studies have shown that circRNAs play an important role in the progression of various human diseases including atherosclerosis, nervous system disorders, diabetes, and cancer. In this review, we summarize the advances on endogenous circRNAs in eukaryotic cells and elucidate their diagnostic and prognostic significance in human cancers. Especially, we highlight the involvement of circRNAs in signal transduction pathways as well as their clinical potential to serve as biomarkers.
Keywords: circular RNA, cancer, diagnosis, biomarkers.
Introduction
Circular RNAs (circRNAs) are newly discovered endogenous non-coding RNAs hundreds or even thousands of bases in length with a covalently closed structure [1, 2] and are transcribed by RNA polymerase II with the same efficiency as linear RNAs [3]. However, unlike in linear RNA splicing processes, circRNAs are produced when the splice donor site is joined to a splice acceptor site upstream in the primary transcript [4, 5]. Previous studies have demonstrated that circRNAs are tissue and cell-type specific, developmentally regulated, and exist during the eukaryotic tree of life suggesting they have important functions in various biological activities [6, 7]. Some circRNAs can be produced by the same gene with different isoforms showing different expression profiles, such as circSTAU2a and circSTAU2b [8].
CircRNAs are produced by circularization of a single exon [9, 10], several exons [11], exons and introns named exon-intron circRNA (ElcirRNA) [12], or intronic sequences alone (circularized intron RNA; ciRNA) [13, 14]. CircRNAs were first found in some plant viroids as covalently closed circular RNA molecules with the base pairing between inverted complementary sequences at the 3' and 5' ends of linear molecules [15]. This was confirmed by an electron microscopic study which also indicated that circRNAs are mainly located in the cytoplasm of eukaryotic cells [16]. In 1986, Kos et al. discovered a single-stranded circular RNA molecule in the Hepatitis delta virus (HDV) by electron microscopy which was the first animal virus identified with a circular RNA genome [17].
A decade ago, Cocquerelle et al. identified novel human ets-1 transcripts which corresponded to circular RNA molecules containing only exons in genomic order. These novel transcripts were structurally stable and localized in the cytoplasmic compartment of the cells. This new type of circRNAs was thought to represent a novel aspect of gene expression and hold some interesting clues about the splicing mechanism [18]. However, these new discoveries did not receive much attention and circRNAs were merely considered to be a class of low-abundance RNA molecules formed by mis-splicing of exon transcripts.
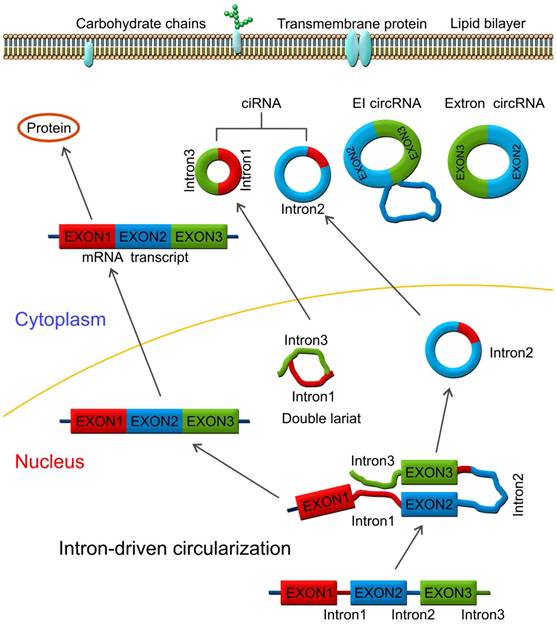
With the progress of deep sequencing of long RNAs and computational algorithms, a large number of novel circRNAs were identified in plants, invertebrates, and vertebrates regardless of whether they were replicated and/or induced in vivo accumulation of small RNAs [19, 20]. High-throughput sequencing analyses have shown that circRNAs are abundant and stable in eukaryotic transcriptosomes suggesting that they may be the key players in the RNA world [7, 21-23]. CircRNAs exhibit tissue- and developmental-specific expression and play crucial roles in multiple cellular processes, such as functioning as miRNA sponges by sequestering miRNA thereby affecting the stability of target mRNAs and dynamically regulating mRNA translation (Figure 1) [24-26]. Recently, Memczak et al. detected 1950, 1903, and 724 circRNAs in human, mice, and C. elegans, respectively [25]. Zheng et al. generated ribosomal-depleted RNA sequencing data from six normal tissues and seven cancers and detected at least 27,000 circRNA candidates [27]. They claimed that many of these circRNAs were differentially expressed between the normal and cancerous tissues. Recently, a new role for circRNAs as regulators of gene expression in the nucleus has been described. CircRNAs were reported to enhance the expression of their parental genes in cis and regulate the transcription activity of target genes via specific RNA-RNA interactions between U1 snRNA and EIcirRNAs [12]. Thousands of well-expressed, stable circRNAs were identified in various human diseases and life processes, such as aging [28-32], insulin secretion [33, 34], tissue development [35, 36], atherosclerotic vascular disease risk [37-39], cardiac hypertrophy [40-44], and cancer [26, 45-49].
Lariat-driven circularization model. CircRNAs are produced by a lariat-driven circularization model containing exon 2 and exon 3 produced from exon skipping. The lariat subsequently undergoes internal splicing and circRNAs are generated with or without removing introns 1 and 3. The important roles of circRNAs include functioning as miRNA sponges and sequestering miRNAs by interacting with miRNAs via miRNA response elements (MRE), regulating mRNA translation and targeting mRNA stability.
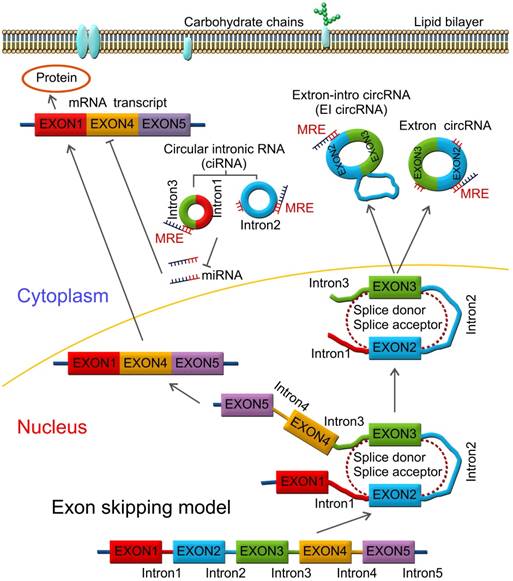
Formation of circRNAs and key features
Pre-mRNAs are spliced into linear molecules that alternately join the exons and retain the exon order [50, 51]. The alternative splicing generates mRNAs encoding proteins [52-54]. Recently, it was shown that exon transcripts in pre-mRNAs might also be non-linearly reverse-spliced into a circular RNA [55-59]. There are two kinds of circRNAs: exonic circRNAs [60-62] and intronic circRNAs [63-65]. The biogenesis and formation of exonic circRNA mainly include three types of circularization driven by lariats [66, 67], intron-pairing [68, 69], and RNA-binding proteins (RBPs) [70, 71], which are described below.
Mostly, circRNAs are formed through lariat-driven circularization model (Figure 1). The mechanism involves the creation of a lariat containing an exon produced from exon skipping. For example, the splice donor of 3'-end of exon 1 and splice acceptor of 5'-end of exon 4 are covalently joined together. The lariat is subsequently processed by internal splicing. Finally, the intron is released and the circRNA consisting of exon2-exon3 is formed.
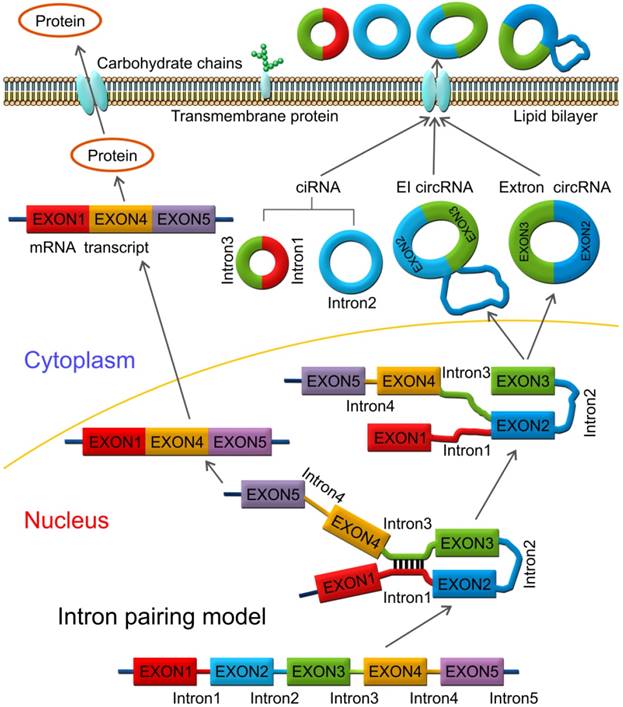
In some other cases, circRNAs are formed via direct back splicing (Figure 2). Two unspliced introns interact by complementary base pairing, thereby juxtaposing the branch point of the 5'-intron and the 3-intron'-exon junction (3-splice donor) for a nucleophilic attack and cleavage. Then, the 3'-splice donor attacks the 5'-intron-exon junction (5-splice acceptor) joining the two introns and releasing the circularized exon [72].
Direct back splicing model. The unspliced introns 1 and 3 interact through complementary base pairing. CircRNAs are formed and released by circularization of exon 2 - exon 3, or exon 2 - intron 2 - exon 3. CircRNAs are present in blood, bodily fluids, and tissues and may serve as biomarkers for human cancers, such as the has_circ_0029 in gastric cancer; Has_circ_100855 in laryngeal squamous cell cancer; circ_CDYL and has_circ_001569 in colorectal cancer; has_circ_0005075 in hepatocellular carcinoma etc.
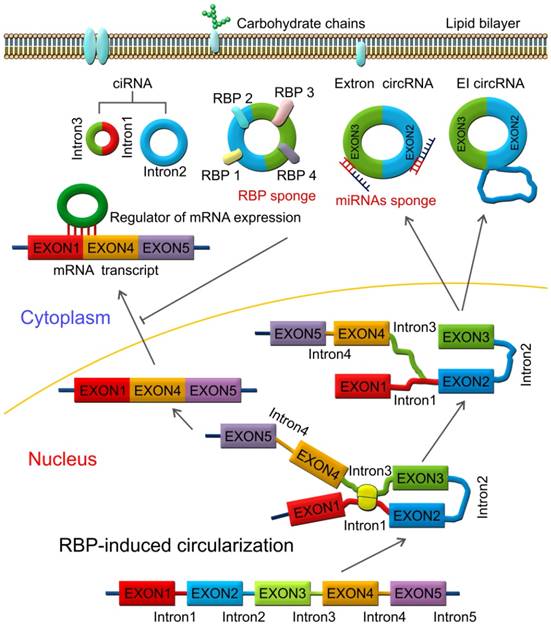
RBPs are key components in the circularization of RNAs. As shown in Figure 3, base pairing is mediated by RBP between intron 1 and intron 3. Thus, a circular structure is formed and the introns are removed to form circRNAs [26, 70]. The formation of circRNAs has also been shown to be affected by RBPs, which bind to the target sites in the flanking introns and induce circularization and biogenesis of circRNA (Figure 3). Two such examples are of RBPs, such as muscleblind (MBNL1) and adenosine deaminase 1 acting on RNA (ADAR1), which have been reported to enhance or suppress circRNA formation and biogenesis by different modes of action. MBNL1 binds to its own pre-mRNA and bridges two flanking introns close together to induce back-splicing [59]. ADAR1 regulation is involved in Adenosine-to Inosine (A-to-I) editing and regulates circRNA biogenesis directly through its dsRNA binding activity [71]. Intronic circRNAs are formed by intron-driven circularization model [73-75].
RNA-binding protein (RBP)-induced circularization model. Base-pairing is mediated by RBP between intron 1 and intron 3. Circular structure is formed and the introns are then removed to form circRNAs, which can function as miRNA or RBP sponges and regulators of mRNA expression.
During the intron-driven circularization, lariat introns are processed from primary transcripts in a two-step mechanism. First, the 2'-OH group of the branch point adenosine (bpA) attacks the 5'-splice site generating a 3'-OH group at the 5'-exon. Next, the newly generated 3'-OH group carries out a nucleophilic attack on the 3'-splice site forming an excised lariat intron and a linear RNA transcript composed of the alternative combined exons (Figure 4) [76].
Intron-driven circularization model. Intronic circRNAs are generated via a two-step mechanism. The 2'-OH group of a defined adenosine within intron 2 attacks the 5'-exon and the lariat intermediate is formed as well as a free 3'-OH group at the 5'-exon. Next, the generated 3'-OH group attacks the 3'-splice site to produce an excised lariat intron and a linear RNA product.
CircRNAs in diagnosis and prognosis of human cancers
CircRNAs have been reported to play important roles in various diseases, such as atherosclerosis [77], nervous system disorders [78], diabetes [79], and cancer [80, 81]. The unique features and diverse functions of circRNAs make them promising candidates with clinical potentials in life sciences and medicine. Here, we summarize the role of circRNAs in cancer with specific reference to cellular development, proliferation, and apoptosis.
CircRNAs are differentially expressed in radioresistant cancer cells
Complex multidisciplinary methods incorporating surgery, chemotherapy, radiotherapy, and immunotherapy are applied for the treatment of human cancers and the molecular alterations that ensue following these treatments have been amply studied. However, the expression profiles of circRNAs in various therapeutic models have not been widely studied. Differential expression of cirRNAs in radioresistant cancer cells has been reported [82]. Radiotherapy is one of the most important modalities for cancer treatment [82, 83]. Acquired radio-resistance during the course of the treatment is considered to be the main reason for the failure of radiation therapy and tumor recurrence [84-86]. CircRNAs, functioning as miRNA sponges, regulate the expression of numerous cancer-related miRNAs and are believed to be involved in the progression of human cancers [87]. The circRNA-miRNA-mRNA axis has been reported to play a role in several cancer-associated pathways with both agonist or antagonist effects on carcinogenesis and metastasis [88]. In this study, a comprehensive expression and functional profile of differentially expressed circRNAs in radioresistant esophageal cancer cells was determined which indicated the possible involvement of the dysregulated circRNAs in the development of radiation resistance. Among the detected 3752 circRNA candidates, 57 circRNAs were significantly upregulated and 17 circRNAs were downregulated in human radioresistant esophageal cancer cell line KYSE-150R compared to the parental cell line KYSE-150 (fold change >/=2.0 and P < 0.05). Furthermore, GO bioinformatics analysis showed the enrichment of 400 target genes including most miRNAs in the Wnt signaling pathway. CircRNAs_001059 and _000167 were the two largest nodes in circRNA/miRNA co-expression network [82].
CircRNAs are involved in signaling pathways in human cancers
CircRNAs are relatively stable in animal cells and can regulate the target gene expression in several ways. First, circRNAs, have a covalently closed loop with no 5'-3' polarity or poly A tail; the circular loop consists of various miRNA binding sites known as miRNA response elements (MREs) enabling circRNAs to act as miRNA sponges. Second, circRNAs can also regulate mRNA expression by functioning as RBP sponges. And third, circRNAs can regulate mRNA expression by directly binding to targeted mRNAs. By virtue of their interactions with miRNA, circRNAs play key roles in regulating cancer progression and may be involved in a variety of signaling pathways in cancers [89-93] (Figure 5) such as mitogen-activated protein kinase (MAPK)/extracellular signal-regulated kinase (ERK1/2), phosphatidylinositol 3-kinase (PI3K)/protein kinase B (AKT) intracellular signaling pathway, and Wnt/beta-catenin pathway [94-99] (Table 2). Here, we present a brief overview of numerous examples of specific and mainly cancer-related circRNAs that have the potential of directly or indirectly impacting key signaling pathways and cellular processes.
CircRNAs and the related signaling pathways in human cancers. Wnt/β-catenin signaling pathway plays an important role in the progression of human cancers. Cir-ITH and circ-cdr1as are two key circRNAs that regulate the proteins or miRNAs in Wnt/β-catenin signaling pathway. CircTCF25, circ-Foxo3 and circZNF292 are three key circRNAs that regulate cell cycle progression in various cancers. MAPK/ERK and PI3K/AKT are key signaling pathways involved in human cancers. Several circRNAs such as ciRS-7, circ001042, circ-TTBK2 and circTCF25 have been identified as regulators of these signaling networks.
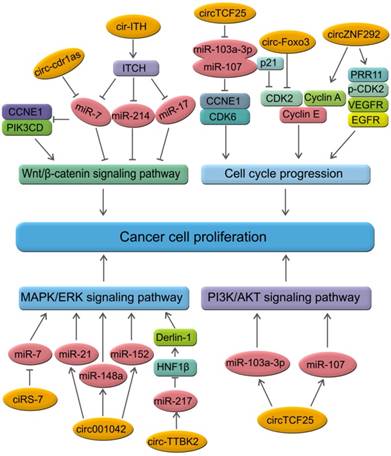
CircRNAs and the related signaling pathways in human cancers
| circRNAs | MiRNA target | m RNA-miRNA target | Cancers | Signaling pathway | reference |
|---|---|---|---|---|---|
| circRNA_001059 and circRNA_000167 | / | / | Radioresistant esophageal cancer | Wnt signaling pathway | [82] |
| Cir-ITCH | miR-7, miR-17 and miR-214 | ITCH | Lung cancer | Cir-ITCH inhibits lung cancer progression by enhancing its parental gene, ITCH, expression and suppresses the activation of Wnt/beta-catenin signaling. | [106-108] |
| Cdr1as | miR-7 | CCNE1 and PIK3CD | hepatocellular carcinoma | Knockdown of Cdr1as inhibited HCC cell proliferation and invasion by targeting miR-7 and Cdr1asand functioned as an oncogene partly through targeting miR-7 in HCC. | [116] |
| cZNF292 | / | PRR11, Cyclin A, p-CDK2, VEGFR-1/2, p-VEGFR-1/2 and EGFR | human glioma U87MG and U251 cells | cZNF292 is an important oncogenic circRNA. Knockdown of cZNF292 significantly inhibited proliferation of glioma cells and cell cycle progression by suppressing tube formationand could be a therapeutic target and biomarker in glioma. | [118] |
| CircTCF25 | MiR-103a-3p/miR-107 | CDK6 | Bladder cancer | Over-expression of circTCF25 down-regulated miR-103a-3p and miR-107, increased CDK6 expression, and promoted proliferation and migration in vitro and in vivo | [83] |
| Foxo3 circRNA | Eight miRNAs, including miR-103a-3p/miR-107 | FOXO3, CDK6, CDK2, p21,CCNE1 | Mouse cancer cells lines (NIH3T3, 4T1 and B16) | Ectopic expression of Foxo3 circRNA inhibited tumor growth, cancer cell proliferation, and survival by suppressing angiogenesis and cell cycle progression. | [119, 120] |
| circHIPK3 | miR-124 | Human cells | circHIPK3 binds to miR-124 and inhibits its activity. Knowckdown of circHIPK3 significantly suppressed human cancer | [27] |
Several circRNAs have been reported to impact the EGFR/RAF1/MAPK signaling. Activation of the EGFR/RAF1/MAPK pathway has been reported via suppression of miR-7 activity. The circRNA CiRS-7 was proposed to be a potential miR-7 sponge thus impacting the EGFR/RAF1/MAPK [100]. MRPS35_hsa_circ_001042 was predicted to interact simultaneously with miR-21, miR-148a, miR-148b, and miR-152 in the MAPK signaling pathway [101]. In osteoarthritis (OA), circRNA-CER was reported to function as a sponge by competitively binding miR-136 and could target and regulate MMP13 through TGFβ, JNK, and ERK pathways [102]. Circ-TTBK2 was upregulated in glioma tissues and cell lines and exerted an oncogenic role in glioma cells. Inhibiting circ-TTBK2 restrained malignant progression of glioma cells by upregulating miR-217, which exerted tumor-suppressive function through downregulating HNF1β, which, in turn, activated Derlin-1 via binding to its promoter and activated PI3K/AKT and ERK signaling pathways [103]. Thus, inhibition of circ-TTBK2/miR-217/HNF1β/Derlin-1 axis may be a potential therapeutic target for human gliomas. Another example is of miR-21/ERK signaling which was reported to be a potential therapeutic pathway for the prevention of myocardial fibrosis [104]. Upregulation of miR-21 and miR-200 blocked hsa-circ-000595 by regulating cell apoptosis [105]. Another example is of circTCF25-miR-103a-3p/miR-107 axis which participates in many significant pathways in cancer. Significantly, the PI3K-AKt signaling pathway overlaps with the miR-103a-3p and miR-107 axis [83].
Circular RNA-ITCH (cir-ITCH) was significantly decreased in lung cancer tissues compared to the paired adjacent noncancer tissues obtained from 78 patients. Ectopic expression of cir-ITCH markedly enhanced the level of its parental cancer-suppressive gene, ITCH, and inhibited the cell viability of lung cancer cells by targeting oncogenic miR-7 and miR-214. Furthermore, cir-ITCH acted as a sponge of oncogenic miRNAs and suppressed Wnt/beta-catenin signaling [106]. This was also confirmed in esophageal squamous cell carcinoma (ESCC) in which cir-ITCH acted as a sponge for miR-7, miR-17, and miR-214. Increased expression of ITCH by cir-ITCH, in turn, enhanced the ubiquitination and degradation of phosphorylated Dvl2 (Segment polarity protein disheveled homolog DVL-2), thereby inhibiting the Wnt/beta-catenin pathway. These data revealed that cir-ITCH has an inhibitory effect on ESCC via its effect on the Wnt pathway [107]. In another study of 45 pairs of specimens from colorectal cancer (CRC) patients, cir-ITCH expression was significantly down-regulated in CRC tissues compared to the peritumoral tissue further confirming that cir-ITCH was involved in the inhibition of the Wnt/beta-catenin pathway [108].
Chen et al. compared the differentially expressed circRNAs in gastric cancer (GC) and paired normal tissues and identified at least 5500 distinct circRNA candidates. CircPVT1, derived from the PVT1 gene, was upregulated in GC tissues and promoted tumor proliferation by acting as a sponge for members of the miR-125 family. CircPVT1 could serve as an independent prognostic marker for overall survival and disease-free survival in patients with GC [109]. CiRS-7, a potential miR-7 sponge, was significantly up-regulated in colorectal cancer tissues compared with matched normal mucosae and emerged as an independent risk factor for overall survival. Thus, CiRS-7 was a promising prognostic biomarker and served as a therapeutic target for reducing EGFR-RAF1 activity in colorectal cancer patients [100].
CircRNA sponges withstand enzymatic degradation and are completely resistant to miRNA-mediated degradation. Studies showed that circRNAs against miR-21 or miR-22 are more effective than the typical linear miRNA sponges and the miRNA inhibitors in repressing miRNA targets. Some circRNAs also display superior anti-cancer activities compared to the linear sponges as shown in malignant cell lines [110, 111]. Especially, the expression levels of circRNAs and linear RNAs showed an inverse trend for many cancer-related signaling pathways including NFkB, PI3k/AKT, and TGF-beta [112-115].
Circular RNA Cdr1as expression was upregulated in hepatocellular carcinoma tissues compared to the adjacent non-tumor tissues. Knockdown of Cdr1as or overexpression of miR-7 suppressed HCC proliferation and invasion by inhibiting the CCNE1 and PIK3CD expression. Importantly, knockdown of Cdr1as suppressed the HCC cell proliferation and invasion through targeting miR-7. Thus, Cdr1as plays an important role to promote HCC progression as an oncogene partly through targeting miR-7 [116]. This was contrary to the myocardial infarction mouse model in which Cdr1as and miR-7a were both upregulated and Cdr1as overexpression in myocardial cardiomyocytes promoted cell apoptosis but was reversed by miR-7a overexpression [117]. This study implied the roles of circRNAs were tissue specific and the circRNA-miRNA association was bidirectional.
It has been reported that cZNF292 is a new circular oncogenic RNA involved in the progression of tube formation. Knockdown of cZNF292 significantly inhibited the proliferation and cell cycle progression in glioma cells by suppressing tube formation. Cell cycle progression in human glioma U87MG and U251 cells was halted at S/G2/M phase via the Wnt/beta-catenin signaling pathway and the levels of related genes, such as PRR11, Cyclin A, p-CDK2, VEGFR-1/2, p-VEGFR-1/2, and EGFR were altered suggesting that cZNF292 silencing plays an important role in the tube formation process. Cznf292 could, therefore, be used as a therapeutic target and biomarker in gliomas [118].
In bladder carcinoma tissues, circRNA expression profiles were screened by microarray analysis. The results demonstrated that in a total of 469 dysregulated circular transcripts, 285 circRNAs were up-regulated and 184 were down-regulated in bladder cancer tissues compared to normal tissues. Over-expression of circTCF25 could sequester and down-regulate miR-103a-3p/miR-107 potentially leading to the up-regulation of thirteen targets including CDK6 which promoted cell proliferation, migration, and invasion in vitro and in vivo [83].
Another example is of the forkhead family of transcription factor 3 (Foxo3) circRNA, which binds to eight miRNAs to regulate Foxo3. Ectopic expression of Foxo3 circRNA inhibited tumor growth, cancer cell proliferation, and survival by suppressing angiogenesis [119]. It has been reported that circ-Foxo3 was highly expressed in normal cells and was associated with cell cycle progression. Knockdown of endogenous circ-Foxo3 promoted cell growth, whereas ectopic expression of circ-Foxo3 could bind to the cell cycle proteins cyclin-dependent kinase 2 (also known as cell division protein kinase 2 or CDK2) and cyclin-dependent kinase inhibitor 1 (or p21) to form a circ-Foxo3-p21-CDK2 ternary complex blocking cell cycle progression [120].
The expression patterns of circRNAs in the mammary glands of lactating rats were analyzed. Numerous circRNAs were differentially and specifically expressed at different lactation stages. At two different lactation stages in rat mammary glands, a total of 6,824 and 4,523 circRNAs were identified. Also, 1,314 circRNAs were detected at both lactation stages. An interesting observation was that four protein-coding genes (Rev3l, IGSF11, MAML2, and LPP) transcribed high levels of circRNAs involved in breast development and breast cancer [121]. Expression profiling and the comparative analysis of 46 gliomas and normal brain samples detected thousands of circRNAs, 476 of which were differentially expressed [122].
A variety of circRNAs were identified which were potentially involved in the tumorigenesis of basal cell carcinoma (BCC). Twenty-three upregulated and 48 downregulated circRNAs with 354 MREs were reported which were capable of sequestering miRNA target sequences of the BCC miRNome [123]. Additionally, 322 circRNAs were identified as differentially expressed in cutaneous squamous cell carcinoma (cSCC). Among these, 143 circRNAs were upregulated and 179 were down-regulated. Furthermore, a total of 1603 MREs were found to be differentially expressed in cSCC participating in the epigenetic control [124].
An abundant circRNA, circHIPK3, derived from exon2 of the HIPK3 gene, was observed to sponge nine miRNAs with 18 potential binding sites. It was reported that circHIPK3 directly bound to miR-124 and inhibited miR-124 activity. Knockdown of circHIPK3 but not HIPK3 mRNA could significantly inhibit human cancer cellular growth by sponging multiple miRNAs [27].
CircRNAs are potential novel biomarkers for diagnosis and prognosis of human cancers
CircRNAs have been shown to regulate gene expression through various mechanisms and can be useful biomarkers in a variety of diseases due to their stability and specific expression. Also, circRNAs can serve as oncogenic stimuli or tumor suppressors in cancer and are therefore of potential significance in cancer prognosis and treatment (Table 1) [22, 125, 126]. Here, we summarize several circRNAs, which were used as novel biomarkers in the prognosis and clinical therapy of various human cancers.
CircRNAs as diagnostic and prognostic markers for human cancers
| circRNAs | Cancer | Function | Reference |
|---|---|---|---|
| Hsa_circ_0001649 | hepatocellular carcinoma | Regulates tumorigenesis and metastasis and might serve as a novel potential biomarker for HCC. | [129] |
| ciRS-7 (Cdr1as) | hepatocellular carcinoma | May be a promising biomarker of hepatic microvascular invasion (MVI) and a novel therapeutic target for restraining MVI in hepatocellular carcinoma. | [116, 130, 135] |
| hsa_circRNA_100855 and hsa_circRNA_104912 | laryngeal squamous cell cancer | hsa_circRNA_100855 was the most upregulated circRNA and hsa_circRNA_104912 was the most downregulated circRNA in LSCC. Patients with T3-4 stage, neck nodal metastasis, poor differentiation or advanced clinical stage had a higher level of hsa_circRNA_100855 and lower hsa_circRNA_104912 expression. | [134] |
| Hsa_circ_002059 | Gastric cancer | Significantly downregulated in gastric cancer tissues compared with paired adjacent nontumorous tissue and had a significant correlation with distal metastasis (P=0.036), TNM stage (P=0.042), gender (P=0.002), and age (P=0.022). | [136] |
| Hsa_circ_001988 | Colorectal cancer | Decreased expression was observed in colorectal cancer. Can serve as a novel potential biomarker for diagnosis and a potential novel therapeutic target for colorectal cancer | [142] |
| circTCF25 | Human bladder cancer | Could sequester miR-103a-3p/miR-107 potentially leading to the up-regulation of 13 targets related to cell proliferation, migration, and invasion and thus serve as a new promising marker for bladder cancer. | [83] |
Serum exosomal circRNAs have been used to distinguish patients with colon cancer from healthy controls [127]. More than 1,800 circular RNAs were analyzed. The ratio of circular to linear RNA isoforms was significantly lower in tumors compared to normal colon samples and much lower in colorectal cancer cell lines and correlated negatively with the proliferation index [128]. Using 89 paired specimens of HCC and adjacent liver tissues, hsa_circ_0001649 was found to be significantly downregulated and its level of expression was closely correlated with tumor size (p = 0.045) and the occurrence of tumor embolus (p = 0.017). The study concluded that hsa_circ_0001649 was involved in the regulation of tumorigenesis and metastasis of HCC and might serve as a novel potential biomarker for human HCC [129]. Another example is of CiRS-7 (also termed as Cdr1as), which is the inhibitor and sponge of miR-7 in the embryonic zebrafish midbrain and islet cells. Compared with the matched non-tumor tissues, the ciRS-7 level was significantly decreased in 60.2 % specimens of HCC tissues (65 out of 108, 60.2 %). However, high expression of ciRS-7 in HCC tissues was tightly associated with hepatic microvascular invasion (MVI), AFP level, and young age and thus partly related to the progression of HCC. Furthermore, the level of ciRS-7 in HCC tissues with concurrent MVI was inversely associated with that of miR-7. Also, ciRS-7 level was positively associated with the expression of two miR-7-targeted genes, PIK3CD (r = 0.55, p < 0.001) and p70S6K (r = 0.34, p = 0.021). Thus, ciRS-7 was one of the independent factors of hepatic MVI [130]. In addition, the levels of hsa_circ_0000520, hsa_circ_0005075, and hsa_circ_0066444 were shown to be differentially expressed in HCC tissues (n = 3) and adjacent normal liver (n = 3) tissues. The expression level of hsa_circ_0005075 was significantly higher in 60 HCC tissues than the matched nontumorous tissues, was correlated with HCC tumor size, and showed good diagnostic potential. Furthermore, hsa_circ_0005075 targeted the miRNA genes, including miR-23b-5p, miR-93-3p, miR-581, miR-23a-5p, and their corresponding mRNAs, suggesting hsa_circ_0005075 could participate in cell adhesion during HCC development [131]. The expression of miR-23b and miR-23a was associated with cancer metastasis and invasion. For example, miR-23b regulated cell migration and invasion of glioblastoma cells via targeting of Pyk2 [132]; MiR-23a modulated the migration and invasion of non-small cell lung cancer cells by targeting insulin receptor substrate-1 (IRS-1) [133].
CircRNAs were studied in 4 paired laryngeal squamous cell cancer tissues (LSCC) and adjacent non-tumor tissues by microarray analysis. The results showed that 302 circRNAs were significantly increased and 396 circRNAs were decreased in LSCC tissues; hsa_circRNA_100855 was the most upregulated circRNA and hsa_circRNA_104912 was the most downregulated circRNA. Altered levels of these two circRNAs were associated with the T3-4 stage, neck nodal metastasis or advanced clinical stage, implying their important roles in the tumorigenesis and the potential to serve as novel and stable biomarkers for the diagnosis/prognosis of LSCC [134]. The ciRS-7, the miRNA sponge for miR-7 and sex-determining region Y (SRY), regulated the biological behavior of various cancers by targeting miRNA-7 (miR-7) and miRNA-138 (miR-138) [135]. In gastric cancer patients, hsa_circ_002059 levels were significantly decreased in cancer tissues compared with paired adjacent nontumorous tissues, as well as in the plasma between preoperative and postoperative gastric cancer patients. Furthermore, lower levels of hsa_circ_002059 were significantly correlated with distal metastasis, TNM stage, gender, and age, indicating has_circ_002059 could be used as a potential novel and stable biomarker to screen gastric carcinoma [136].
In the clinical setting, circRNAs, which are found in blood, bodily fluids or tissues as a sign of human cancers, are used as biomarkers to detect human cancers (Figure 2). The stability, sensitivity, and specificity of cricRNAs must be suitable and appropriate to be used a biomarker, such as the has_circ_0029 in gastric cancer, has_circ_100855 in laryngeal squamous cell cancer, circ_CDYL and has_circ_001569 in colorectal cancer, and has_circ_0005075 in hepatocellular carcinoma. Other new circRNAs that were recently discovered might also serve as possible biomarkers.
Perspectives and future directions
With the rapid advances in RNA sequencing and analysis, thousands of endogenous circRNAs have been detected in mammalian cells which are highly conserved [137, 138]. So far, the identification and function of circRNAs in human cancers remained elusive. In this review, we described the wide-spread expression of circRNAs as well as their functions. We also summarized our current understanding of their molecular regulatory mechanisms including involvement in crucial signaling pathways. However, several outstanding issues that need to be clarified and/or areas of circRNAs research to focus on include the following:
1. Although circRNA transcripts were discovered in the early 1980s, the information on various aspects of circRNAs is still limited [16]. Recently, circRNAs have emerged as key regulators in the development and progression of multiple human cancers. Furthermore, circRNAs show a more robust expression pattern across patients than mRNAs and are believed to be involved in tumor growth and metastasis, suggesting that circRNAs have a great potential to be employed as biomarkers in clinical diagnosis for a variety of cancers. There is a need to expand the screening in a variety of human cancers and identifying new candidate circRNAs that can serve not only as reliable diagnostic and prognostic biomarkers but also as therapeutic targets [139] [140].
2. The presence of circRNAs for the detection of not only cancers but also a variety of other diseases needs to be explored in easily accessible specimens of blood, saliva, urine, and feces to realize their full clinical potential.
3. CircRNAs are novel non-coding RNAs which appear to function as oncogenes or tumor suppressor genes in human cancers. One of the molecular mechanisms through which some circRNAs regulate cancer progression is by targeting specific miRNAs [116, 141]. The role of circRNAs in cellular transcriptional control needs to be further explored with respect to cancer development.
4. CircRNAs with multiple MREs are highly stable and effective inhibitors [123, 124]. It would be important to explore whether the sequence and number of MREs can be manipulated to enhance the regulatory effects of circRNAs for the therapy of human cancers.
Acknowledgements
The work was supported by National Natural Science Foundation of China ( No. 31300624,81372322,81460440 and 81560471); the Joint Special Funds for the Department of Science and Technology of Yunnan Province-Kunming Medical University (No. 2014FB059), the Scientific Research Projects from Internal Research Institutions of Medical and Health Units in Yunnan Province (Nos. 2014NS013, 2014NS014, 2014NS015 and 2014NS016), Foundation of the Yunnan Provincial Innovative Team of Bone and Soft Tissue Tumor (No. 2015HC026), Foundation of the Young and Middle-aged Academic and Technical Leaders of Yunnan Province (No. 2014HB034), Doctor Scientific Research Startup Funds of the Third Affiliated Hospital of Kunming Medical University (No. BSJJ201406) and Doctor Scientific Research Startup Funds of the Third Affiliated Hospital of Kunming Medical University (No. BSJJ201406) and China Postdoctoral Science Foundation funded the project (Grant No. 2013M530473).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Salzman J, Chen RE, Olsen MN. et al. Cell-type specific features of circular RNA expression. PLoS Genet. 2013;9(9):e1003777
2. Afonina ZA, Myasnikov AG, Shirokov VA. et al. Formation of circular polyribosomes on eukaryotic mRNA without cap-structure and poly(A)-tail: a cryo electron tomography study. Nucleic Acids Res. 2014;42(14):9461-9
3. Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12(4):381-8
4. Lasda E. and R. Parker, Circular RNAs: diversity of form and function. RNA. 2014;20(12):1829-42
5. Dong Y, He D, Peng Z. et al. Circular RNAs in cancer: an emerging key player. J Hematol Oncol. 2017;10(1):2
6. Barrett S.P. and J. Salzman.Circular RNAs: analysis, expression and potential functions. Development. 2016;143(11):1838-47
7. Meng X, Li X, Zhang P. et al. Circular RNA: an emerging key player in RNA world. Brief Bioinform. 2016
8. Rybak-Wolf A, Stottmeister C, Glažar P. et al. Circular RNAs in the Mammalian Brain Are Highly Abundant, Conserved, and Dynamically Expressed. Mol Cell. 2015;58(5):870-85
9. Starke S, Jost I, Rossbach O. et al. Exon circularization requires canonical splice signals. Cell Rep. 2015;10(1):103-11
10. Kelly S, Greenman C, Cook PR. et al. Exon Skipping Is Correlated with Exon Circularization. J Mol Biol. 2015;427(15):2414-7
11. Wilusz JE. Circular RNAs: Unexpected outputs of many protein-coding genes. RNA Biol. 2016:1-11
12. Li Z, Huang C, Bao C. et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22(3):256-64
13. Monat C, Quiroga C, Laroche-Johnston F. et al. The Ll.LtrB intron from Lactococcus lactis excises as circles in vivo: insights into the group II intron circularization pathway. RNA. 2015;21(7):1286-93
14. Aucamp J, Bronkhorst AJ, Pretorius PJ.A Historical, Evolutionary Perspective on Circulating Nucleic Acids, Extracellular Vesicles. Circulating Nucleic Acids as Homeostatic Genetic Entities. Adv Exp Med Biol. 2016;924:91-5
15. Hewlett MJ, Pettersson RF, Baltimore D.Circular forms of Uukuniemi virion RNA. an electron microscopic study. J Virol. 1977;21(3):1085-93
16. Hsu MT, Coca-Prados M. Electron microscopic evidence for the circular form of RNA in the cytoplasm of eukaryotic cells. Nature. 1979;280(5720):339-40
17. Kos A, Dijkema R, Arnberg AC. et al. The hepatitis delta (delta) virus possesses a circular RNA. Nature. 1986;323(6088):558-60
18. Cocquerelle C, Mascrez B, Hétuin D. et al. Mis-splicing yields circular RNA molecules. FASEB J. 1993;7(1):155-60
19. Zhang Z, Qi S, Tang N. et al. Discovery of replicating circular RNAs by RNA-seq and computational algorithms. PLoS Pathog. 2014;10(12):e1004553
20. Hoffmann S, Otto C, Doose G. et al. A multi-split mapping algorithm for circular RNA, splicing, trans-splicing and fusion detection. Genome Biol. 2014;15(2):R34
21. Veneziano D, Di Bella S, Nigita G. et al. Noncoding RNA: Current Deep Sequencing Data Analysis Approaches and Challenges. Hum Mutat. 2016;37(12):1283-98
22. Lyu D, Huang S. The emerging role and clinical implication of human exonic circular RNA. RNA Biol. 2016:1-7
23. Kulcheski FR, Christoff AP, Margis R. Circular RNAs are miRNA sponges and can be used as a new class of biomarker. J Biotechnol. 2016;238:42-51
24. Ebbesen KK, Kjems J, Hansen TB. Circular RNAs: Identification, biogenesis and function. Biochim Biophys Acta. 2016;1859(1):163-8
25. Memczak S, Jens M, Elefsinioti A. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature. 2013;495(7441):333-8
26. Qu S, Yang X, Li X. et al. Circular RNA: A new star of noncoding RNAs. Cancer Lett. 2015;365(2):141-8
27. Zheng Q, Bao C, Guo W. et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat Commun. 2016;7:11215
28. Abdelmohsen K, Panda AC, De S. et al. Circular RNAs in monkey muscle: age-dependent changes. Aging (Albany NY). 2015;7(11):903-10
29. Maiese K. Disease onset and aging in the world of circular RNAs. J Transl Sci. 2016;2(6):327-29
30. Westholm JO, Miura P, Olson S. et al. Genome-wide analysis of drosophila circular RNAs reveals their structural and sequence properties and age-dependent neural accumulation. Cell Rep. 2014;9(5):1966-80
31. Xu Y, Komiyama M. Structure, function and targeting of human telomere RNA. Methods. 2012;57(1):100-5
32. Larson K, Yan SJ, Tsurumi A. et al. Heterochromatin formation promotes longevity and represses ribosomal RNA synthesis. PLoS Genet. 2012;8(1):e1002473
33. Xu H, Guo S, Li W. et al. The circular RNA Cdr1as, via miR-7 and its targets, regulates insulin transcription and secretion in islet cells. Sci Rep. 2015;5:12453
34. Schneider T, Hung LH, Schreiner S. et al. CircRNA-protein complexes: IMP3 protein component defines subfamily of circRNPs. Sci Rep. 2016;6:31313
35. Mills JD, Chen BJ, Ueberham U. et al. The Antisense Transcriptome and the Human Brain. J Mol Neurosci. 2016;58(1):1-15
36. Floris G, Zhang L, Follesa P. et al. Regulatory Role of Circular RNAs and Neurological Disorders. Mol Neurobiol. 2016
37. Burd CE, Jeck WR, Liu Y. et al. Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. PLoS Genet. 2010;6(12):e1001233
38. Holdt LM1, Stahringer A, Sass K. et al. Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat Commun. 2016;7:12429
39. Rafferty B, Dolgilevich S, Kalachikov S. et al. Cultivation of Enterobacter hormaechei from human atherosclerotic tissue. J Atheroscler Thromb. 2011;18(1):72-81
40. Xie YZ, Yang F, Tan W. et al. The anti-cancer components of Ganoderma lucidum possesses cardiovascular protective effect by regulating circular RNA expression. Oncoscience. 2016;3(7-8):203-7
41. Wang K, Long B, Liu F. et al. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur Heart J. 2016;37(33):2602-11
42. Khan MA, Reckman YJ, Aufiero S. et al. RBM20 Regulates Circular RNA Production From the Titin Gene. Circ Res. 2016;119(9):996-1003
43. Werfel S, Nothjunge S, Schwarzmayr T. et al. Characterization of circular RNAs in human, mouse and rat hearts. J Mol Cell Cardiol. 2016;98:103-7
44. Thum T. Facts and updates about cardiovascular non-coding RNAs in heart failure. ESC Heart Fail. 2015;2(3):108-111
45. Qu S, Zhong Y, Shang R. et al. The emerging landscape of circular RNA in life processes. RNA Biol. 2016:1-8
46. Guarnerio J, Bezzi M, Jeong JC. et al. Oncogenic Role of Fusion-circRNAs Derived from Cancer-Associated Chromosomal Translocations. Cell. 2016;165(2):289-302
47. Chen X, Fan S, Song E. Noncoding RNAs: New Players in Cancers. Adv Exp Med Biol. 2016;927:1-47
48. Bao C, Lyu D, Huang S. Circular RNA expands its territory. Mol Cell Oncol. 2016;3(2):e1084443
49. Athyala PK, Kanwar JR, Alameen M. et al. Probing the biophysical interaction between Neocarzinostatin toxin and EpCAM RNA aptamer. Biochem Biophys Res Commun. 2016;469(2):257-62
50. Chen I, Chen CY, Chuang TJ. Biogenesis, identification, and function of exonic circular RNAs. Wiley Interdiscip Rev RNA. 2015;6(5):563-79
51. Gilbert KB, Fahlgren N, Kasschau KD. et al. Preparation of Multiplexed Small RNA Libraries From Plants. Bio Protoc. 2014:4 (21)
52. Berman AJ, Akiyama BM, Stone MD. et al. The RNA accordion model for template positioning by telomerase RNA during telomeric DNA synthesis. Nat Struct Mol Biol. 2011;18(12):1371-5
53. Kayal E, Bentlage B, Collins AG.Insights into the transcriptional, translational mechanisms of linear organellar chromosomes in the box jellyfish Alatina alata (Cnidaria. Medusozoa: Cubozoa). RNA Biol. 2016;13(9):799-809
54. Fomina-Yadlin D, Mujacic M, Maggiora K. et al. Transcriptome analysis of a CHO cell line expressing a recombinant therapeutic protein treated with inducers of protein expression. J Biotechnol. 2015;212:106-15
55. Granados-Riveron JT, Aquino-Jarquin G. The complexity of the translation ability of circRNAs. Biochim Biophys Acta. 2016;1859(10):1245-51
56. Broadbent KM, Broadbent JC, Ribacke U. et al. Strand-specific RNA sequencing in Plasmodium falciparum malaria identifies developmentally regulated long non-coding RNA and circular RNA. BMC Genomics. 2015;16:454
57. Suzuki H, Tsukahara T. A view of pre-mRNA splicing from RNase R resistant RNAs. Int J Mol Sci. 2014;15(6):9331-42
58. Salzman J, Gawad C, Wang PL. et al. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One. 2012;7(2):e30733
59. Ashwal-Fluss R, Meyer M, Pamudurti NR. et al. circRNA biogenesis competes with pre-mRNA splicing. Mol Cell. 2014;56(1):55-66
60. Xia S, Feng J, Lei L. et al. Comprehensive characterization of tissue-specific circular RNAs in the human and mouse genomes. Brief Bioinform. 2016
61. Ye CY, Zhang X, Chu Q. et al. Full-length sequence assembly reveals circular RNAs with diverse non-GT/AG splicing signals in rice. RNA Biol. 2016:1-9
62. Yang L. Splicing noncoding RNAs from the inside out. Wiley Interdiscip Rev RNA. 2015;6(6):651-60
63. Kramer MC, Liang D, Tatomer DC. et al. Combinatorial control of Drosophila circular RNA expression by intronic repeats, hnRNPs, and SR proteins. Genes Dev. 2015;29(20):2168-82
64. Nielsen H. Group I intron ribozymes. Methods Mol Biol. 2012;848:73-89
65. Lu T, Cui L, Zhou Y. et al. Transcriptome-wide investigation of circular RNAs in rice. RNA. 2015;21(12):2076-87
66. Barrett SP, Wang PL, Salzman J. Circular RNA biogenesis can proceed through an exon-containing lariat precursor. Elife. 2015;4:e07540
67. Tang Y, Nielsen H, Masquida B. et al. Molecular characterization of a new member of the lariat capping twin-ribozyme introns. Mob DNA. 2014;5:25
68. Zhang XO, Wang HB, Zhang Y. et al. Complementary sequence-mediated exon circularization. Cell. 2014;159(1):134-47
69. Salgia SR, Singh SK, Gurha P. et al. Two reactions of Haloferax volcanii RNA splicing enzymes: joining of exons and circularization of introns. RNA. 2003;9(3):319-30
70. Souii A1, M'hadheb-Gharbi MB, Gharbi J. Cellular Proteins Act as Bridge Between 5' and 3' Ends of the Coxsackievirus B3 Mediating Genome Circularization During RNA Translation. Curr Microbiol. 2015;71(3):387-95
71. Ivanov A, Memczak S, Wyler E. et al. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep. 2015;10(2):170-7
72. Wang Y, Wang Z. Efficient backsplicing produces translatable circular mRNAs. RNA. 2015;21(2):172-9
73. Schmidt CA, Noto JJ, Filonov GS. et al. A Method for Expressing and Imaging Abundant, Stable, Circular RNAs In Vivo Using tRNA Splicing. Methods Enzymol. 2016;572:215-36
74. Suzuki H, Aoki Y, Kameyama T. et al. Endogenous Multiple Exon Skipping and Back-Splicing at the DMD Mutation Hotspot. Int J Mol Sci. 2016:17 (10)
75. Lan PH, Liu ZH, Pei YJ. et al. Landscape of RNAs in human lumbar disc degeneration. Oncotarget. 2016;7(39):63166-76
76. Liang D, Wilusz JE. Short intronic repeat sequences facilitate circular RNA production. Genes Dev. 2014;28(20):2233-47
77. Xiang J, Wang Y, Su K. et al. Ritonavir binds to and downregulates estrogen receptors: molecular mechanism of promoting early atherosclerosis. Exp Cell Res. 2014;327(2):318-30
78. van Rossum D, Verheijen BM, Pasterkamp RJ. Circular RNAs: Novel Regulators of Neuronal Development. Front Mol Neurosci. 2016;9:74
79. Wang Y, Xu XY, Tang YR. et al. Effect of endogenous insulin-like growth factor and stem cell factor on diabetic colonic dysmotility. World J Gastroenterol. 2013;19(21):3324-31
80. Marques-Rocha JL, Samblas M, Milagro FI. et al. Noncoding RNAs, cytokines, and inflammation-related diseases. FASEB J. 2015;29(9):3595-611
81. Wang Y, Mo Y, Gong Z. et al. Circular RNAs in human cancer. Mol Cancer. 2017;16(1):25
82. Su H, Lin F, Deng X. et al. Profiling and bioinformatics analyses reveal differential circular RNA expression in radioresistant esophageal cancer cells. J Transl Med. 2016;14(1):225
83. Zhong Z, Lv M, Chen J. Screening differential circular RNA expression profiles reveals the regulatory role of circTCF25-miR-103a-3p/miR-107-CDK6 pathway in bladder carcinoma. Sci Rep. 2016;6:30919
84. Xie C, Jin X, Su H. The Potential Involvement of Circular RNA in the Development of Radiation Resistance of Esophageal Cancer. Int J Radiat Oncol Biol Phys. 2016;96(2S):E149
85. Tao Z, Chen S, Mao G. et al. The PDRG1 is an oncogene in lung cancer cells, promoting radioresistance via the ATM-P53 signaling pathway. Biomed Pharmacother. 2016;83:1471-7
86. Saki M, Toulany M, Rodemann HP. Acquired resistance to cetuximab is associated with the overexpression of Ras family members and the loss of radiosensitization in head and neck cancer cells. Radiother Oncol. 2013;108(3):473-8
87. Zhang P, Zuo Z, Shang W. et al. Identification of differentially expressed circular RNAs in human colorectal cancer. Tumour Biol. 2017;39(3):1010428317694546
88. Xin Z, Ma Q, Ren S. et al. The understanding of circular RNAs as special triggers in carcinogenesis. Brief Funct Genomics. 2016;16(2):80-6
89. Li P, Xue WJ, Feng Y. et al. Long non-coding RNA CASC2 suppresses the proliferation of gastric cancer cells by regulating the MAPK signaling pathway. Am J Transl Res. 2016;8(8):3522-9
90. Yang N, Wang Y, Hui L. et al. Silencing SOX2 Expression by RNA Interference Inhibits Proliferation, Invasion and Metastasis, and Induces Apoptosis through MAP4K4/JNK Signaling Pathway in Human Laryngeal Cancer TU212 Cells. J Histochem Cytochem. 2015;63(9):721-33
91. Parra E. Inhibition of JNK-1 by small interfering RNA induces apoptotic signaling in PC-3 p;rostate cancer cells. Int J Mol Med. 2012;30(4):923-30
92. Shen W, Yuan Y, Zhao M. et al. Novel long non-coding RNA GACAT3 promotes gastric cancer cell proliferation through the IL-6/STAT3 signaling pathway. Tumour Biol. 2016;37(11):14895-902
93. Zhao JJ, Hao S, Wang LL. et al. Long non-coding RNA ANRIL promotes the invasion and metastasis of thyroid cancer cells through TGF-beta/Smad signaling pathway. Oncotarget. 2016;7(36):57903-18
94. Xiao C, Wu CH, Hu HZ. LncRNA UCA1 promotes epithelial-mesenchymal transition (EMT) of breast cancer cells via enhancing Wnt/beta-catenin signaling pathway. Eur Rev Med Pharmacol Sci. 2016;20(13):2819-24
95. Wu C, Zhuang Y, Jiang S. Interaction between Wnt/beta-catenin pathway and microRNAs regulates epithelial-mesenchymal transition in gastric cancer (Review). Int J Oncol. 2016;48(6):2236-46
96. Morris SL, Huang S. Crosstalk of the Wnt/beta-catenin pathway with other pathways in cancer cells. Genes Dis. 2016;3(1):41-47
97. McCubrey JA, Rakus D, Gizak A. et al. Effects of mutations in Wnt/beta-catenin, hedgehog, Notch and PI3K pathways on GSK-3 activity-Diverse effects on cell growth, metabolism and cancer. Biochim Biophys Acta. 2016;1863(12):2942-2976
98. Kobayashi M, Funayama R, Ohnuma S. et al. Wnt-beta-catenin signaling regulates ABCC3 (MRP3) transporter expression in colorectal cancer. Cancer Sci. 2016;107(12):1776-84
99. Guo YH, Wang LQ, Li B. et al. Wnt/beta-catenin pathway transactivates microRNA-150 that promotes EMT of colorectal cancer cells by suppressing CREB signaling. Oncotarget. 2016;7(27):42513-26
100. Weng W, Wei Q, Toden S. et al. Circular RNA ciRS-7 - A promising prognostic biomarker and a potential therapeutic target in colorectal cancer. Clin Cancer Res. 2017
101. Y Zhang, Ren XZ, Wang YZ. et al. Bioinformatics Interaction Analysis of Wnt1-Targeting miRNAs and Their Corresponding circRNAs and Target Genes. Progress in Biochemistry and Biophysics. 2016;43(11):1094-1101
102. Liu Q, Zhang X, Hu X. et al. Circular RNA Related to the Chondrocyte ECM Regulates MMP13 Expression by Functioning as a MiR-136 'Sponge' in Human Cartilage Degradation. Sci Rep. 2016;6:22572
103. Zheng J, Liu X, Xue Y. et al. TTBK2 circular RNA promotes glioma malignancy by regulating miR-217/HNF1beta/Derlin-1 pathway. J Hematol Oncol. 2017;10(1):52
104. Cheng M, Wu G, Song Y. et al. Celastrol-Induced Suppression of the MiR-21/ERK Signalling Pathway Attenuates Cardiac Fibrosis and Dysfunction. Cell Physiol Biochem. 2016;38(5):1928-38
105. Cesana M, Cacchiarelli D, Legnini I. et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell. 2011;147(2):358-69
106. Wan L, Zhang L, Fan K. et al. Circular RNA-ITCH Suppresses Lung Cancer Proliferation via Inhibiting the Wnt/beta-Catenin Pathway. Biomed Res Int. 2016;2016:1579490
107. Li F, Zhang L, Li W. et al. Circular RNA ITCH has inhibitory effect on ESCC by suppressing the Wnt/beta-catenin pathway. Oncotarget. 2015;6(8):6001-13
108. Huang G, Zhu H, Shi Y. et al. cir-ITCH plays an inhibitory role in colorectal cancer by regulating the Wnt/beta-catenin pathway. PLoS One. 2015;10(6):e0131225
109. Chen J, Li Y, Zheng Q. et al. Circular RNA profile identifies circPVT1 as a proliferative factor and prognostic marker in gastric cancer. Cancer Lett. 2016;388:208-219
110. Xie H, Ren X, Xin S. et al. Emerging roles of circRNA_001569 targeting miR-145 in the proliferation and invasion of colorectal cancer. Oncotarget. 2016;7(18):26680-91
111. Liu Y, Cui H, Wang W. et al. Construction of circular miRNA sponges targeting miR-21 or miR-221 and demonstration of their excellent anticancer effects on malignant melanoma cells. Int J Biochem Cell Biol. 2013;45(11):2643-50
112. Bitar MS, Abdel-Halim SM, Al-Mulla F. Caveolin-1/PTRF upregulation constitutes a mechanism for mediating p53-induced cellular senescence: implications for evidence-based therapy of delayed wound healing in diabetes. Am J Physiol Endocrinol Metab. 2013;305(8):E951-63
113. Ahmed I, Karedath T, Andrews SS. et al. Altered expression pattern of circular RNAs in primary and metastatic sites of epithelial ovarian carcinoma. Oncotarget. 2016;7(24):36366-36381
114. Qiao Y, Han X, Guan G. et al. TGF-beta triggers HBV cccDNA degradation through AID-dependent deamination. FEBS Lett. 2016;590(3):419-27
115. Jenkins RH, Bennagi R, Martin J. et al. A conserved stem loop motif in the 5'untranslated region regulates transforming growth factor-beta(1) translation. PLoS One. 2010;5(8):e12283
116. Yu L, Gong X, Sun L. et al. The Circular RNA Cdr1as Act as an Oncogene in Hepatocellular Carcinoma through Targeting miR-7 Expression. PLoS One. 2016;11(7):e0158347
117. Geng HH, Li R, Su YM. et al. The Circular RNA Cdr1as Promotes Myocardial Infarction by Mediating the Regulation of miR-7a on Its Target Genes Expression. PLoS One. 2016;11(3):e0151753
118. Yang P, Qiu Z, Jiang Y. et al. Silencing of cZNF292 circular RNA suppresses human glioma tube formation via the Wnt/beta-catenin signaling pathway. Oncotarget. 2016;7(39):63449-55
119. Yang W, Du WW, Li X. et al. Foxo3 activity promoted by non-coding effects of circular RNA and Foxo3 pseudogene in the inhibition of tumor growth and angiogenesis. Oncogene. 2016;35(30):3919-31
120. Du WW, Yang W, Liu E. et al. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 2016;44(6):2846-58
121. Zhang C, Wu H, Wang Y. et al. Expression Patterns of Circular RNAs from Primary Kinase Transcripts in the Mammary Glands of Lactating Rats. J Breast Cancer. 2015;18(3):235-41
122. Song X, Zhang N, Han P. et al. Circular RNA profile in gliomas revealed by identification tool UROBORUS. Nucleic Acids Res. 2016;44(9):e87
123. Sand M, Bechara FG, Sand D. et al. Circular RNA expression in basal cell carcinoma. Epigenomics. 2016;8(5):619-32
124. Sand M, Bechara FG, Gambichler T. et al. Circular RNA expression in cutaneous squamous cell carcinoma. J Dermatol Sci. 2016;83(3):210-8
125. Abu N, Jamal R. Circular RNAs as promising biomarkers: a mini-review. Frontiers in Physiology. 2016;7:355
126. Wang F, Nazarali AJ, Ji S. Circular RNAs as potential biomarkers for cancer diagnosis and therapy. Am J Cancer Res. 2016;6(6):1167-76
127. Li Y, Zheng Q, Bao C. et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res. 2015;25(8):981-4
128. Bachmayr-Heyda A, Reiner AT, Auer K. et al. Correlation of circular RNA abundance with proliferation-exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci Rep. 2015;5:8057
129. Qin M, Liu G, Huo X. et al. Hsa_circ_0001649: A circular RNA and potential novel biomarker for hepatocellular carcinoma. Cancer Biomark. 2016;16(1):161-9
130. Xu L, Zhang M, Zheng X. et al. The circular RNA ciRS-7 (Cdr1as) acts as a risk factor of hepatic microvascular invasion in hepatocellular carcinoma. J Cancer Res Clin Oncol. 2016;143(1):17-27
131. Shang X, Li G, Liu H. et al. Comprehensive Circular RNA Profiling Reveals That hsa_circ_0005075, a New Circular RNA Biomarker, Is Involved in Hepatocellular Crcinoma Development. Medicine (Baltimore). 2016;95(22):e3811
132. Loftus JC, Ross JT, Paquette KM. et al. miRNA expression profiling in migrating glioblastoma cells: regulation of cell migration and invasion by miR-23b via targeting of Pyk2. PLoS One. 2012;7(6):e39818
133. Cao M, Li Y, Lu H. et al. MiR-23a-mediated migration/invasion is rescued by its target, IRS-1, in non-small cell lung cancer cells. J Cancer Res Clin Oncol. 2014;140(10):1661-70
134. Xuan L, Qu L, Zhou H. et al. Circular RNA: a novel biomarker for progressive laryngeal cancer. Am J Transl Res. 2016;8(2):932-9
135. Zhao ZJ1, Shen J. Circular RNA participates in the carcinogenesis and the malignant behavior of cancer. RNA Biol. 2015:1-8
136. Li P, Chen S, Chen H. et al. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin Chim Acta. 2015;444:132-6
137. Yao Z, Luo J, Hu K. et al. ZKSCAN1 gene and its related circular RNA (circZKSCAN1) both inhibit hepatocellular carcinoma cell growth, migration and invasion but through different signaling pathways. Mol Oncol. 2017;11(4):422-37
138. Xu S, Xiao S, Qiu C. et al. Transcriptome-wide identification and functional investigation of circular RNA in the teleost large yellow croaker (Larimichthys crocea). Mar Genomics. 2017;32:71-8
139. Hou LD, Zhang J. Circular RNAs: An emerging type of RNA in cancer. Int J Immunopathol Pharmacol. 2017;30(1):1-6
140. Ahmed I, Karedath T, Andrews SS. et al. Altered expression pattern of circular RNAs in primary and metastatic sites of epithelial ovarian carcinoma. Oncotarget. 2016;7(24):36366-81
141. Maksimenko AV, Volkov EM, Bertrand JR. et al. Targeting of single-stranded DNA and RNA containing adjacent pyrimidine and purine tracts by triple helix formation with circular and clamp oligonucleotides. Eur J Biochem. 2000;267(12):3592-603
142. Wang X, Zhang Y, Huang L. et al. Decreased expression of hsa_circ_001988 in colorectal cancer and its clinical significances. Int J Clin Exp Pathol. 2015;8(12):16020-5
Author contact
Corresponding author: Zuozhang Yang, Ph.D, E-mail: yangzuozhangcom Tel.: +86-871-68221157 Fax: +86-871-68221157
 Global reach, higher impact
Global reach, higher impact