13.3
Impact Factor
Theranostics 2018; 8(8):2278-2288. doi:10.7150/thno.23544 This issue Cite
Research Paper
DNA Methylation Signatures Predicting Bevacizumab Efficacy in Metastatic Breast Cancer
1. III rd Medical Department with Hematology, Medical Oncology, Hemostaseology, Infectious Diseases and Rheumatology, Oncologic Center; Salzburg Cancer Research Institute (SCRI) with Laboratory of Immunological and Molecular Cancer Research (LIMCR) and Center for Clinical Cancer and Immunology Trials (CCCIT); Paracelsus Medical University Salzburg, Salzburg, Austria;
2. Division of Bioinformatics, Biocenter, Medical University of Innsbruck, Innsbruck, Austria;
3. Center for Health & Bioresources, Business Unit for Molecular Diagnostics, AIT - Austrian Institute of Technology GmbH, Vienna, Austria;
4. Department of Pathology, Paracelsus Medical University Salzburg, Salzburg, Austria;
5. Division of Cancer Genetics/Epigenetics, Department of Molecular Biology, University of Salzburg, Salzburg, Austria;
6. Cancer Cluster Salzburg, Salzburg, Austria
*These authors contributed equally to this work.
Abstract
Background: Biomarkers predicting response to bevacizumab in breast cancer are still missing. Since epigenetic modifications can contribute to an aberrant regulation of angiogenesis and treatment resistance, we investigated the influence of DNA methylation patterns on bevacizumab efficacy.
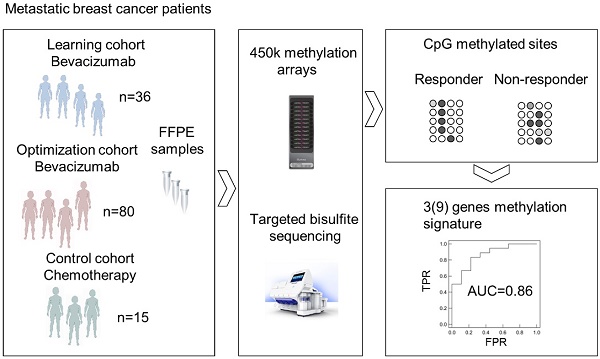
Methods: Genome-wide methylation profiling using the Illumina Infinium HumanMethylation450 BeadChip was performed in archival FFPE specimens of 36 patients with HER2-negative metastatic breast cancer treated with chemotherapy in combination with bevacizumab as first-line therapy (learning set). Based on objective response and progression-free survival (PFS) and considering ER expression, patients were divided in responders (R) and non-responders (NR). Significantly differentially methylated gene loci (CpGs) with a strong change in methylation levels (Δβ>0.15 or Δβ<-0.15) between R and NR were identified and further investigated in 80 bevacizumab-treated breast cancer patients (optimization set) and in 15 patients treated with chemotherapy alone (control set) using targeted deep amplicon bisulfite sequencing. Methylated gene loci were considered predictive if there was a significant association with outcome (PFS) in the optimization set but not in the control set using Spearman rank correlation, Cox regression, and logrank test.
Results: Differentially methylated loci in 48 genes were identified, allowing a good separation between R and NR (odds ratio (OR) 101, p<0.0001). Methylation of at least one cytosine in 26 gene-regions was significantly associated with progression-free survival (PFS) in the optimization set, but not in the control set. Using information from the optimization set, the panel was reduced to a 9-gene signature, which could divide patients from the learning set into 2 clusters, thereby predicting response with an OR of 40 (p<0.001) and an AUC of 0.91 (LOOCV). A further restricted 3-gene methylation model showed a significant association of predicted responders with longer PFS in the learning and optimization set even in multivariate analysis with an excellent and good separation of R and NR with AUC=0.94 and AUC=0.86, respectively.
Conclusion: Both a 9-gene and 3-gene methylation signature can discriminate between R and NR to a bevacizumab-based therapy in MBC and could help identify patients deriving greater benefit from bevacizumab.
Keywords: Metastatic breast cancer, bevacizumab, methylation, signature, predictive
 Global reach, higher impact
Global reach, higher impact