Impact Factor
Theranostics 2016; 6(12):2129-2140. doi:10.7150/thno.16356 This issue Cite
Research Paper
Proto-Oncogenic Src Phosphorylates EB1 to Regulate the Microtubule-Focal Adhesion Crosstalk and Stimulate Cell Migration
1. State Key Laboratory of Medicinal Chemical Biology, College of Life Sciences, Nankai University, Tianjin 300071, China.
2. Institute of Biomedical Sciences, College of Life Sciences, Key Laboratory of Animal Resistance Biology of Shandong Province, Key Laboratory of Molecular and Nano Probes of the Ministry of Education, Shandong Normal University, Jinan 250014, China.
Received 2016-6-1; Accepted 2016-8-4; Published 2016-9-12
Abstract
Cell migration, a complex process critical for tumor progression and metastasis, requires a dynamic crosstalk between microtubules (MTs) and focal adhesions (FAs). However, the molecular mechanisms underlying this event remain elusive. Herein we identify the proto-oncogenic protein Src as an important player in the regulation of the MT-FA crosstalk. Src interacts with and phosphorylates end-binding protein 1 (EB1), a member of MT plus end-tracking proteins (+TIPs), both in cells and in vitro. Systematic mutagenesis reveals that tyrosine-247 (Y247) is the primary residue of EB1 phosphorylated by Src. Interestingly, both constitutively activated Src and Y247-phosphorylated EB1 localize to the centrosome and FAs. Src-mediated EB1 phosphorylation diminishes its interactions with other +TIPs, including adenomatous polyposis coli (APC) and mitotic centromere associated kinesin (MCAK). In addition, EB1 phosphorylation at Y247 enhances the rate of MT catastrophe and significantly stimulates cell migration. These findings thus demonstrate that the Src-EB1 axis plays a crucial role in regulating the crosstalk between MTs and FAs to promote cell migration.
Keywords: EB1, Src, phosphorylation, microtubule, focal adhesion, cell migration.
Introduction
Microtubule (MT) dynamics must be tightly controlled during cell migration in order to facilitate rapid responses to signals that drive polarization1. At the leading edge of the cell, MTs are captured and stabilized by MT plus end-tracking proteins (+TIPs)2. However, despite these stabilizing interactions, MTs at focal adhesions (FAs) undergo catastrophe at a rate seven times higher than that of cytoplasmic MTs3. Increases in MT dynamics at FAs are closely associated with the activation of various signaling pathways that control cell migration4,5. A growing number of studies have demonstrated the importance of the MT-FA crosstalk in cell migration. For example, cytoplasmic linker-associated proteins (CLASPs) located around FAs have been shown to mediate MT capture and establish a transport pathway for FA turnover6. Similarly, FA kinase (FAK) and dynamin also participate in MT-driven FA disassembly7. In contrast, paxillin regulates the dynamics of FA-targeted MTs by serving as a platform for MT targeting8. However, the driving factor responsible for the changes in MT dynamics at FAs remains elusive, suggesting that additional, unknown FA-associated factors may play critical roles in the regulation of MT dynamics at FAs.
End-binding protein 1 (EB1), a member of +TIPs, binds the plus ends of growing MTs via its conserved calponin-homology domain9. EB1 regulates MT dynamics through the recruitment of other +TIPs to MT plus ends, such as SxIP motif-containing proteins and cytoskeleton-associated protein glycine-rich (CAP-Gly) domain-containing proteins10,11, or through coordination with other MT-binding proteins12,13. Interactions between EB1 and other +TIPs have been shown to be important for the regulation of MT dynamics during cell migration. For example, SxIP motif-containing +TIPs, such as adenomatous polyposis coli (APC), CLASPs, MT-actin crosslinking factor 1 (MACF1)/actin crosslinking family 7 (ACF7), and mitotic centromere associated kinesin (MCAK), regulate MT stability at bundled actin-rich sites at the cell cortex2,14,15. However, the molecular mechanisms that regulate the interactions between EB1 and other +TIPs during cell migration remain not fully understood.
A few recent studies have shown that post-translational modifications of EB1 influence its functions in MT dynamics-related activities, such as cell division. However, it is not clear whether the post-translational modification of EB1 occurs in motile cells and if so, how it is orchestrated and whether it contributes to cell migration. The proto-oncogenic protein Src is a non-receptor tyrosine kinase and phosphorylates many substrates, several of which play essential roles in cell migration and adhesion. Activated Src localizes strongly to FAs, where it phosphorylates FAK, paxillin, and vinculin16-18. In response to androgen signaling, Src has also been shown to play an important role in the regulation of centrosome activity and promotion of MT nucleation19. Together, these studies suggest that Src is well-positioned to regulate the crosstalk between MTs and FAs. In the present study, we report that Src phosphorylates EB1 primarily at tyrosine-247 (Y247) to regulate the recruitment of SxIP motif-containing +TIPs. In addition, our data demonstrate that Src-mediated EB1 phosphorylation occurs at the centrosome and FAs and promotes cell migration by regulating the MT-FA crosstalk.
Materials and Methods
Cell culture and transfection
Human embryonic kidney 293T (HEK293T) cells and human umbilical vein endothelial cells (HUVECs) were purchased from the American Type Culture Collection and grown in DMEM medium and RPMI1640 medium, respectively, supplemented with 10% fetal bovine serum at 37°C in 5% CO2. They were recently authenticated and tested for contamination. Polyethyleneimine (Sigma-Aldrich) was used for plasmid transfection into HEK293T cells, and LipoJet (SignaGen Laboratories) was used for plasmid transfection into HUVECs. Control (5'-CGUACGCGGAAUACUUCGA-3') and Src siRNAs (5'-AAACUCCCCUUGCUCAUGUAC-3') were synthesized by Ribobio and transfected with Lipofectamine RNAiMAX (Invitrogen).
Plasmids and proteins
Mammalian expression plasmids for GFP-EB1 and HA-EB1 were constructed by PCR with the pEGFPN1 and pCMV-HA vectors, respectively, as described previously12. Mammalian expression plasmids for GFP-Src, GFP-APC-C, and GFP-MCAK were constructed with the pEGFPC1 vector. Bacterial expression plasmids for GST-Src and His-EB1 were constructed with the pGEX6P3 and pRSET-A vectors, respectively, as described20. GST-Src was purified from bacteria with glutathione-sepharose beads (GE Healthcare Life Sciences), and His-EB1 was purified from bacteria with the nickel resin (Bio-Rad). Various mutants of EB1 and the Y530F mutant of Src were constructed by site-directed mutagenesis.
Antibodies and chemicals
Primary antibodies against α-tubulin (Abcam, ab7291), Src (Abcam, ab109381), vinculin (Abcam, ab18058), GFP (Roche Applied Science, 11814460001), pTyr (Millipore, 05-321), pY418-Src (Abcam, ab4816), HA (Sigma-Aldrich, H3663), GST (Sigma-Aldrich, G7781), γ-tubulin (Abcam, ab11316), His (Medical & Biological Laboratories, D291-3), and EB1 (BD Biosciences, 610534) were purchased from the indicated sources and used as 1:1000 dilution for immunoblotting and 1:100 dilution for immunofluorescence microscopy. Horseradish peroxidase-conjugated secondary antibodies were from GE Life Sciences and used as 1:2000 dilution. Fluorescein- or rhodamine-conjugated secondary antibodies were from Jackson ImmunoResearch Laboratories and used as 1:200 dilution. DAPI, SU6656, and PP2 were from Sigma-Aldrich.
Generation of the pY247-EB1 antibody
The pY247-EB1 customized antibody was obtained from PTM Biolabs. The phosphopeptide (CRIVDIL-pY247-ATDEGF) was synthesized and conjugated with the carrier protein keyhole limpet hemocyanin (KLH) for immunization, and the unmodified peptide (CRIVDIL-Y247-ATDEGFV) was synthesized for depletion. Three 8-week rabbits were immunized with the phosphopeptide for four times. Antibodies were purified with peptide affinity chromatography and verified with ELISA and dot blotting assays with good response to the phosphopeptide and no response to the unmodified peptide.
Immunoblotting, immunoprecipitation, and GST pulldown
Proteins were resolved by SDS-PAGE and transferred onto polyvinylidene difluoride membranes (Millipore). The membranes were blocked and incubated with primary antibodies and then with horseradish peroxidase-conjugated secondary antibodies. Specific proteins were visualized with enhanced chemiluminescence detection reagent (Thermo Fisher Scientific). For immunoprecipitation and GST pulldown, cell lysates or purified proteins were incubated with antibody-coated or glutathione-conjugated agarose beads at 4°C for 2 h. The beads were washed and boiled in the SDS loading buffer, and the proteins were detected by immunoblotting.
In vitro kinase assay
GFP-Src immunoprecipitated from HEK293T cells or bacterially purified GST-Src was incubated with bacterially purified His-EB1 at 30°C for 2.5 h in the kinase reaction buffer (Cell Signaling Technology) containing ATP (200 µM). The reaction samples were then subjected to immunoblotting with pTyr or pY247-EB1 antibodies.
Immunofluorescence microscopy
Cells were fixed and immunostained as described previously21,22. In brief, to visualize vinculin, cells grown on glass coverslips were fixed with 4% paraformaldehyde for 20 min and permeabilized with 0.5% Triton X-100 for 20 min. To visualize other proteins, cells were fixed with cold methanol for 4 min. Cells were incubated with primary antibodies and then rhodamine- or fluorescein-conjugated secondary antibodies followed by DAPI staining. Cells were mounted onto slides and examined under a confocal microscope (Carl Zeiss).
Analysis of GFP-EB1 comets and MT dynamics
Cells expressing GFP-EB1 or its mutants were observed under the confocal microscope, and time-lapse images of GFP-EB1 comets were acquired at 2-s intervals for 2 min with the 488-nm laser illumination as described23. To analyze MT dynamics, the time-lapse images of GFP-EB1 comets were analyzed with the PlusTipTracker software as described24. By using the quadrant-scatter-plot tool, MT growth tracks were overlaid and classified into four groups and color-coded based on the average MT growth speed and lifetime.
Analysis of cell migration
Cells expressing GFP-EB1 or its mutants were examined under an Inverted fluorescence microscope (Carl Zeiss), and time-lapse images were acquired at 5-min intervals for 5 h. Cell migration trajectories were obtained by using the ImageJ software with the manual-tracking and chemotaxis-migration tool plugins as described25. The velocity of cell migration and the distance from the origin were then calculated.
Statistics
Analysis of statistical significance was performed by the Student's t-test for comparison between two groups and by the ANOVA test for multiple comparisons. Correlation coefficient was calculated by the Spearman's rank correlation test.
Results
Src interacts with and phosphorylates EB1 both in cells and in vitro
To investigate the potential role of Src in regulating EB1 functions during cell migration, we first analyzed whether Src affects the level of EB1 phosphorylation. We transfected HEK293T cells with GFP-Src and HA-EB1, and then assessed the level of phosphorylation in the HA-EB1 immunoprecipitate by immunoblotting with antibodies against phosphorylated tyrosine (pTyr). We found that GFP-Src dramatically increased EB1 phosphorylation (Figure 1A). Conversely, treatment of cells with the Src-specific inhibitors SU6656 and PP2, both of which could inhibit Src activation (as indicated by decreased Src phosphorylation at tyrosine-418), reduced the phosphorylation of endogenous EB1 at comparable levels (Figure 1B). These data suggest that Src promotes EB1 phosphorylation in cells.
Src interacts with and phosphorylates EB1 both in cells and in vitro. (A) HEK293T cells were transfected with HA-EB1 and GFP-Src or the GFP vector. Immunoprecipitation (IP) and immunoblotting (IB) were then performed with the indicated antibodies. (B) HEK293T cells were treated with SU6656 (10 μM), PP2 (10 μM), or equal amount of the vehicle DMSO for 12 h. Immunoprecipitation and immunoblotting were then performed as indicated. (C) In vitro kinase assays were performed with purified His-EB1 and the GFP or GFP-Src immunoprecipitate from HEK293T cells, and examined by immunoblotting. (D) In vitro kinase assays were performed with purified His-EB1 and the GFP-Src immunoprecipitate, in the absence (Mock) or presence of SU6656 (50 μM), PP2 (50 μM), or equal amount of DMSO, and analyzed by immunoblotting. (E) In vitro kinase assays were performed with purified His-EB1 and the GFP-Src immunoprecipitate, in the absence (Mock) or presence of λPPase, and analyzed by immunoblotting. (F) In vitro kinase assays were performed with purified His-EB1 and purified GST or GST-Src and analyzed by immunoblotting. (G) HEK293T cells were transfected with GFP-Src or the GFP vector. Immunoprecipitation and immunoblotting were then performed with the indicated antibodies. (H) Purified His-EB1 was incubated with purified GST or GST-Src. GST pulldown and immunoblotting were then performed as indicated. (I) The lysate of HUVECs was immunoprecipitated with the EB1 antibody or IgG control. The interaction between endogenous Src and EB1 was then examined by immunoblotting of the precipitates. All experiments were replicated three times.
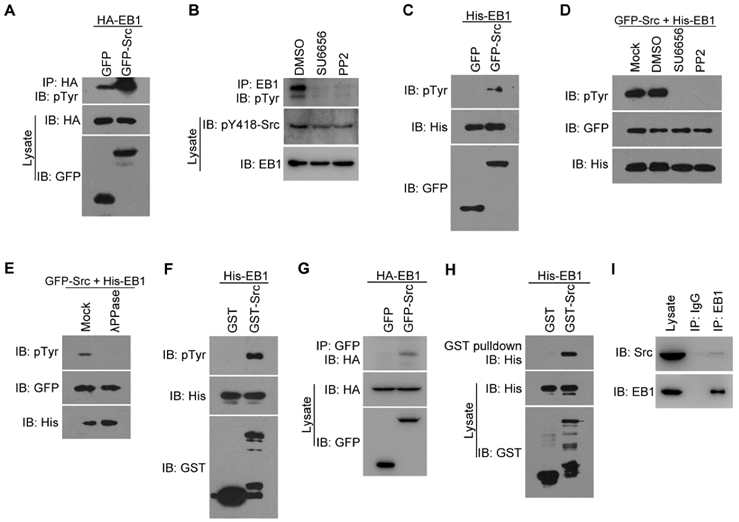
To further examine the role of Src in EB1 phosphorylation, we performed in vitro kinase assays by incubating GFP or GFP-Src immunoprecipitated from HEK293T cells with bacterially purified (His)6-tagged EB1 (His-EB1). Immunoblotting with anti-pTyr antibodies revealed the presence of phosphorylated EB1 when the GFP-Src immunoprecipitate, but not the GFP immunoprecipitate, was added (Figure 1C). This phosphorylation was Src-specific, as it could be inhibited by treatment with Src inhibitors, but not the vehicle DMSO (Figure 1D). Furthermore, treatment with λ phophatase (λPPase) completely abolished the pTyr signal, verifying that the band observed was indeed associated with EB1 phosphorylation (Figure 1E). Use of bacterially purified GST-Src and His-EB1 in in vitro kinase assays also revealed the phosphorylation of EB1 by Src (Figure 1F). Thus, these results indicate that Src can directly phosphorylate EB1.
We then investigated whether Src interacts with EB1 in cells, by transfecting HEK293T cells with GFP-Src and HA-EB1 and performing immunoprecipitation assays. We found that HA-EB1 was co-immunoprecipitated with GFP-Src, but not GFP (Figure 1G). The interaction between Src and EB1 was confirmed by GST pulldown assays with purified GST-Src and His-EB1 (Figure 1H). In addition, we detected an interaction between endogenous Src and EB1 in HUVECs (Figure 1I). Together, these findings demonstrate that EB1 directly interacts with and is a substrate of Src.
Y247 is the primary residue of EB1 phosphorylated by Src
To determine the residue of Src-induced phosphorylation in EB1, we performed in vitro kinase assays with purified GST-Src and His-EB1 wild-type or tyrosine-to-phenylalanine (Y-to-F) or tyrosine-to-alanine (Y-to-A) phospho-deficient mutants. Immunoblotting with anti-pTyr antibodies revealed that the Y247F mutant exhibited the largest decrease in EB1 phosphorylation, suggesting that Y247 is the primary residue in EB1 that is phosphorylated by Src (Figure 2A).
Y247 is the primary residue of EB1 phosphorylated by Src. (A) In vitro kinase assays were performed with purified GST-Src and purified His-EB1 wild-type or mutants. The arrowhead indicates the bands of phosphorylated EB1. (B) HEK293T cells were transfected with the indicated plasmids. Immunoprecipitation and immunoblotting were then performed. (C) In vitro kinase assays were performed with the indicated purified proteins, and immunoblotting were then performed. (D) HUVECs were transfected with GFP-Src and stained with the pY247-EB1 antibody and DAPI. Scale bar, 10 μm. (E) Experiments were performed as in (D), and the relative intensity of pY247-EB1 in the boxed areas was quantified (50 cells were measured for each group). All experiments were replicated three times. ****p < 0.0001. Error bars indicate mean ± SEM.
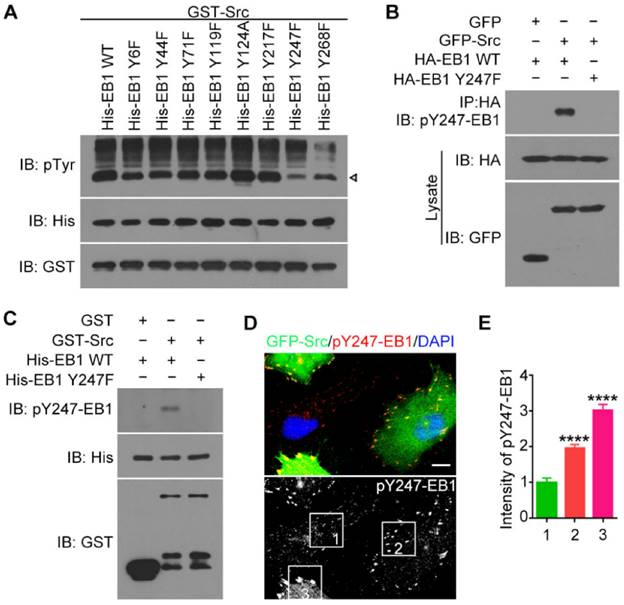
To confirm this finding, we performed immunoblotting with antibodies specifically against EB1 phosphorylated at Y247 (pY247-EB1). We observed a pY247-EB1-specific band in cells transfected with GFP-Src and HA-EB1 wild-type, but not the Y247F mutant (Figure 2B). Similarly, immunoblotting with the pY247-EB1 antibody revealed that in the in vitro kinase assays, GST-Src caused the phosphorylation of His-EB1 wild-type, but not the Y247 mutant (Figure 2C).
To study the physiological relevance of this phosphorylation site, we stained HUVECs expressing GFP-Src with the pY247-EB1 antibody (Figure 2D). We found that pY247-EB1 staining was substantially more intense in HUVECs transfected with GFP-Src compared to untransfected cells (Figure 2D). In addition, the level of pY247-EB1 staining varied in a GFP-Src expression-dependent manner (Figures 2D and 2E). These data suggest that Src phosphorylates endogenous EB1 at Y247.
Activated Src and phosphorylated EB1 co-localize at the centrosome and FAs
We then investigated whether Src-mediated phosphorylation affects the subcellular localization of EB1. By immunofluorescence microscopy, we found that in HUVECs pY247-EB1 primarily localized to the plus ends of MTs (Figure 3A). The pY247-EB1 signal dots partly co-localized with ectopically expressed GFP-EB1, which exhibited robust cytoplasmic localization in addition to tracking MT plus ends (Figure 3B).
Activated Src and phosphorylated EB1 co-localize at the centrosome and FAs. (A) HUVECs were stained with pY247-EB1 (red) and α-tubulin (green) antibodies and DAPI (blue). Scale bar, 10 μm. (B) HUVECs were transfected with GFP-EB1 (green) and stained with the pY247-EB1 antibody (red) and DAPI (blue). Scale bar, 10 μm. (C) HUVECs were stained with DAPI (blue) and antibodies against γ-tubulin (green) and pY247-EB1 or pY418-Src (red). Scale bars, 10 μm. (D) HUVECs were stained with DAPI (blue) and antibodies against vinculin (green) and pY247-EB1 or pY418-Src (red). Scale bars, 10 μm. (E) HUVECs were transfected with GFP-Src Y530F (green) and stained with the pY247-EB1 antibody (red) and DAPI (blue). Scale bar, 10 μm. All experiments were replicated three times.
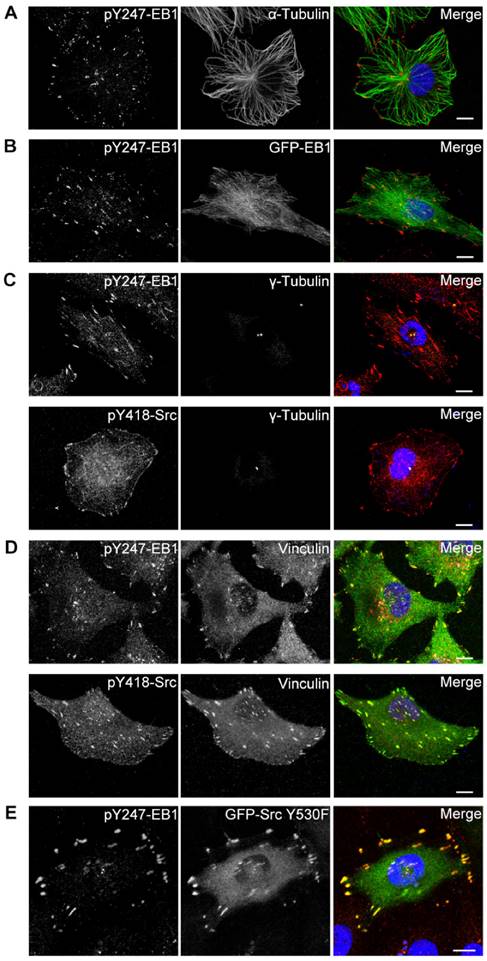
Interestingly, 100% of HUVECs examined showed the localization of pY247-EB1 to the centrosome, where MT growth is initiated, in addition to its localization at MT plus ends (Figures 3A and 3B). This finding was confirmed by staining the centrosomal marker γ-tubulin (Figure 3C). Interestingly, a subset of activated Src (Src phosphorylated at Y418, pY418-Src) also localized at the centrosome (Figure 3C).
We further found that both pY247-EB1 and pY418-Src co-localized with the FA marker vinculin (Figure 3D). We then analyzed the distribution of pY247-EB1 in HUVECs expressing a constitutively active mutant of Src (Y530F). We found that pY247-EB1 and GFP-Src Y530F co-localized with one another and with γ-tubulin (Figure 3E). Together, these findings suggest that Src-mediated phosphorylation of EB1 at Y247 occurs primarily at the centrosome and FAs and may regulate the functions of these cellular structures.
Src-mediated EB1 phosphorylation diminishes its interactions with APC-C and MCAK
Y247 is located in the C-terminal region of EB1, which contains the hydrophobic cavity responsible for binding SxIP motif-containing +TIPs. Therefore, we hypothesized that Src-mediated EB1 phosphorylation at Y247 may regulate the interactions between these proteins. We first examined whether the dimerization of EB1 was affected by its phosphorylation at Y247. Anti-HA immunoblotting revealed that both HA-EB1 wild-type and the Y247D phospho-mimic mutant were efficiently co-immunoprecipitated with GFP-EB1 (Figure 4A), indicating that Y247 phosphorylation had no effect on EB1 dimerization. Next, we examined whether EB1 phosphorylation at Y247 affected its binding to SxIP motif-containing +TIPs, such as the C-terminal EB1-binding domain of APC (APC-C) and MCAK. The Y247D phospho-mimic mutation of EB1 almost abolished its interactions with APC-C (Figure 4B) and MCAK (Figure 4C). Surprisingly, the Y247F phospho-deficient mutation also decreased EB1 binding to APC-C (Figure 4B) and MCAK (Figure 4C).
Src-mediated EB1 phosphorylation diminishes its interactions with APC-C and MCAK. (A) HEK293T cells were transfected with GFP-EB1 and HA-EB1 wild-type or Y247D. Immunoprecipitation and immunoblotting were then performed with the indicated antibodies. (B) HEK293T cells were transfected with GFP-APC-C and HA-EB1 wild-type, Y247F, or Y247D, or the HA vector. Immunoprecipitation and immunoblotting were then performed as indicated. (C) Experiments were performed as in (B) except that GFP-MCAK was used instead of GFP-APC-C. (D) Structural analysis of the interaction between the SxIP motif (violet) and EB1 wild-type, Y247F, or Y247D. (E) HEK293T cells were transfected with GFP-APC-C and treated with SU6656 (10 μM), PP2 (10 μM), or equal amount of DMSO for 12 h or co-transfected with HA-Src or the HA vector. Immunoprecipitation and immunoblotting were then performed. The arrowhead indicates the bands of GFP-APC-C. (F) Experiments were performed as in (D) except that GFP-MCAK was used instead of GFP-APC-C. The arrowhead indicates the bands of GFP-MCAK. All experiments were replicated three times.
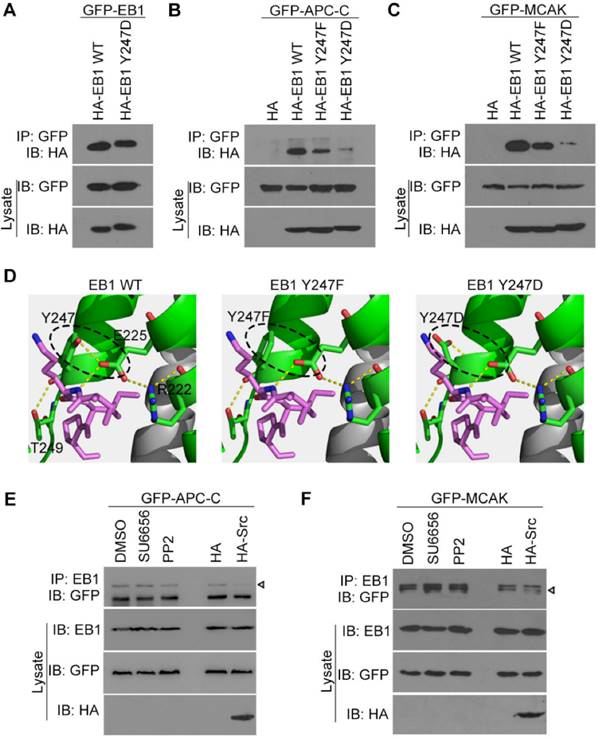
In order to explore the structural basis for these results, we examined the crystal structure of EB1 complexed with the +TIP MACF (Protein Data Bank ID code: 3GJO). In this structure, Y247 was found to form stable hydrogen bonds with E225 and T249 within the EB1 monomer (Figure 4D). Substitution of Y247 by either phenylalanine (F) or aspartic acid (D) could abolish the hydrogen bond between Y247 and E225, potentially disturbing the stability of the hydrophobic cavity in the EB homology (EBH) domain that is responsible for EB1-SxIP interactions (Figure 4D). These data are consistent with the finding that both Y247F and Y247D could decrease EB1 binding to APC-C and MCAK (Figures 4B and 4C).
We then examined the effect of Src on the interactions of EB1 with APC-C and MCAK. We found that Src inhibitors had no obvious effect on the interaction of EB1 with APC-C (Figure 4E), but slightly increased its binding to MCAK (Figure 4F). In contrast, overexpression of Src reduced EB1 binding to both APC-C (Figure 4E) and MCAK (Figure 4F). Together, these data indicate that Src-induced phosphorylation of EB1 decreases its association with SxIP motif-containing +TIPs.
EB1 phosphorylation at Y247 regulates MT dynamics
We next sought to study whether Src-mediated EB1 phosphorylation affects MT dynamics. Time-lapse images of GFP-EB1 comets in HUVECs were recorded and analyzed with the plusTipTracker software. MT growth tracks were overlaid and classified into four color-coded groups based on the average MT growth velocity and lifetime (Figure 5A). We found that both the Y247F mutant and the Y247D mutant of EB1 exhibited increased growth velocity and decreased lifetime relative to wild-type EB1, with Y247D exhibiting the greater change (Figures 5B and 5C).
EB1 phosphorylation at Y247 regulates MT dynamics. (A) MT growth tracks were overlaid and classified into four groups and color-coded based on the average MT growth velocity and lifetime. (B) HUVECs were transfected with GFP-EB1 wild-type, Y247F, or Y247D, and time-lapse videos of GFP-EB1 comets were taken at 2-s intervals. MT growth tracks were then analyzed with the plusTipTracker software. Scale bars, 10 μm. (C) Experiments were performed as in (B), and MT growth velocity and lifetime were measured (20-30 cells were measured for each group). (D) HUVECs were transfected with GFP-EB1 and treated with PP2 (10 μM) or equal amount of DMSO for 6 h. MT growth tracks were then analyzed as in (B). Scale bars, 10 μm. (E) Experiments were performed as in (D), and MT growth velocity and lifetime were then measured (20-30 cells were measured for each group). (F) HUVECs were transfected with GFP-EB1 and HA-Src or the HA vector. MT growth tracks were then analyzed as in (B). Scale bars, 10 μm. (G) Experiments were performed as in (F), and MT growth velocity and lifetime were then measured (20-30 cells were measured for each group). (H) HUVECs were transfected with siControl or siSrc, and immunoblotting was then performed with the indicated antibodies. (I) HUVECs were transfected with GFP-EB1 and siControl or siSrc. MT growth tracks were then analyzed as in (B). Scale bars, 10 μm. (J) Experiments were performed as in (I), and MT growth velocity and lifetime were then measured (20-30 cells were measured for each group). All experiments were replicated three times. *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001. ns, not significant. Error bars indicate mean ± SEM.
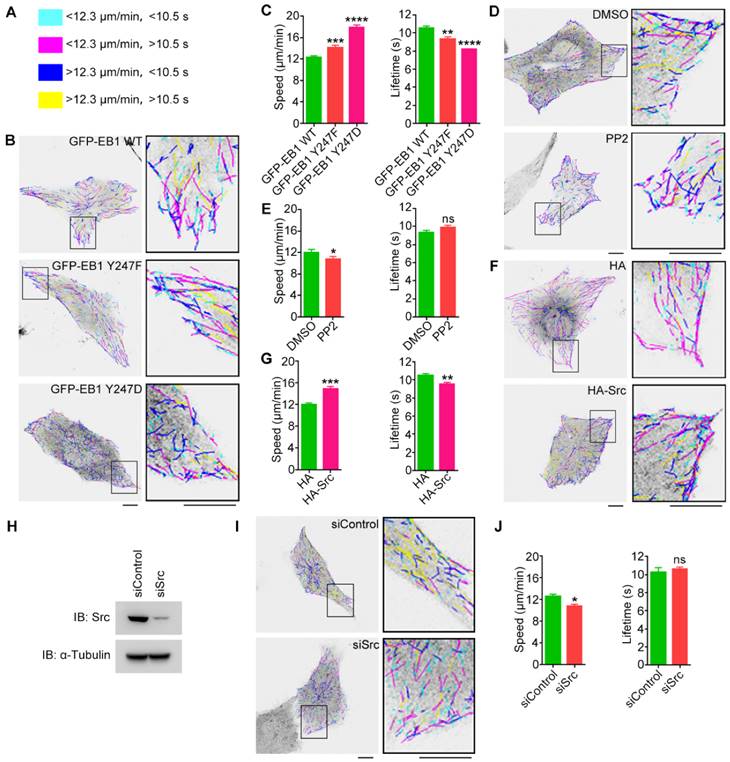
To examine whether changes in Src activity had similar effects, MT dynamics were analyzed in HUVECs transfected with GFP-EB1 and treated with the Src inhibitor PP2. We did not use SU6656 because the intrinsic fluorescence of this Src inhibitor interfered to some extent with the detection of the GFP-EB1 signals. We found that PP2 slightly reduced MT growth velocity, but did not significantly affect MT growth lifetime (Figures 5D and 5E). In contrast, overexpression of HA-Src significantly increased MT growth velocity and decreased MT growth lifetime, similar to those observed for EB1 Y247D (Figures 5F and 5G). We further found that siRNA-mediated knockdown of Src expression reduced MT growth velocity, but did not significantly alter MT growth lifetime (Figures 5H and 5I). Collectively, these results suggest that Src-mediated phosphorylation of EB1 at Y247 increases the rate of MT catastrophe.
Src-mediated phosphorylation of EB1 enhances cell migration
The next question then is whether Src-mediated phosphorylation of EB1 regulates cell migration, which is critically dependent on MT dynamics. The migration trajectories of HUVECs were recorded, and the velocity of cell migration and the distance from the origin were then analyzed with the ImageJ software. Strikingly, the Y247D mutant of EB1 exhibited enhanced migration velocity and distance in comparison to wild-type EB1, but the Y247F mutation did not significantly impede cell migration (Figures 6A-C).
Src-mediated phosphorylation of EB1 enhances cell migration. (A) HUVECs were transfected with GFP-EB1 wild-type, Y247F, or Y247D, and cell migration trajectories were then recorded. Scale bars, 40 μm. (B and C) Experiments were performed as in (A), and the velocity of cell migration (B) and the distance from the origin (C) were measured (20-30 cells were measured for each group). (D) HUVECs were transfected with GFP-EB1 wild-type or Y247D and treated with PP2 or DMSO. Cell migration trajectories were then obtained. Scale bars, 40 μm. (E and F) Experiments were performed as in (D), and the velocity of cell migration (E) and the distance from the origin (F) were measured (20-30 cells were measured for each group). (G) HUVECs were transfected with siControl or siSrc, together with GFP-EB1 wild-type or Y247D. Cell migration trajectories were then obtained. Scale bars, 40 μm. (H and I) Experiments were performed as in (G), and the velocity of cell migration (H) and the distance from the origin (I) were measured (20-30 cells were measured for each group). All experiments were replicated three times. **p < 0.01; ***p < 0.001; ****p < 0.0001. ns, not significant. Error bars indicate mean ± SEM.
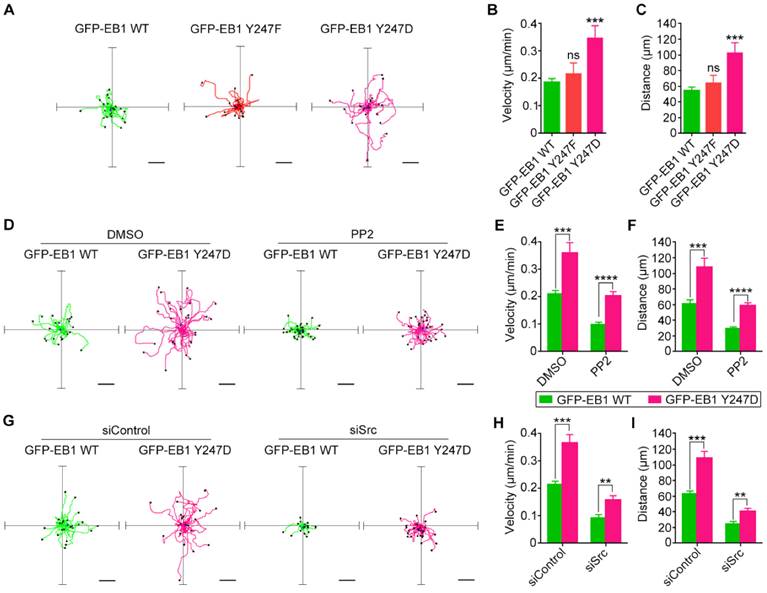
Treatment of HUVECs with the Src inhibitor PP2 decreased both the velocity of cell migration and the distance from the origin (Figures 6D-F). In addition, siRNA-mediated knockdown of Src expression reduced both the velocity of cell migration and the distance from the origin (Figures 6G-I). We found that even in the presence of PP2 or Src siRNA, cell migration velocity and distance for the EB1 Y247D group were greater than those for wild-type EB1 (Figures 6D-I). Based on these observations, we conclude that Src enhances cell migration through regulating EB1 phosphorylation.
Discussion
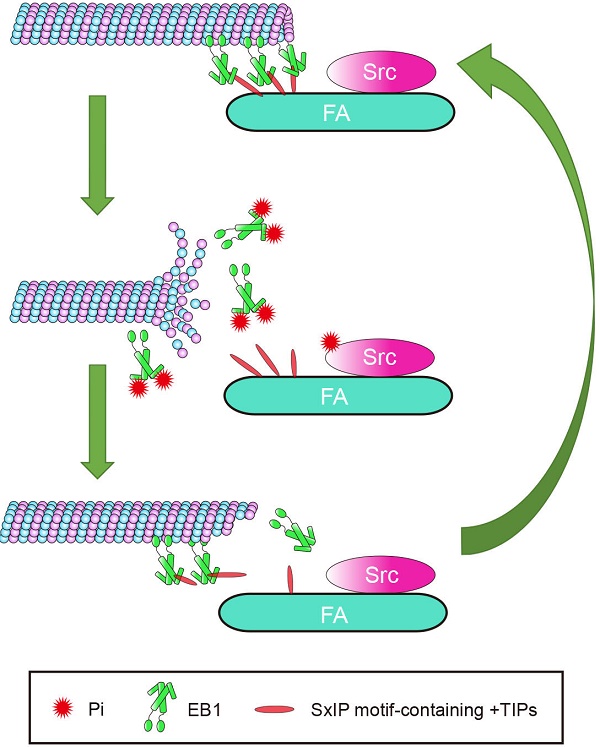
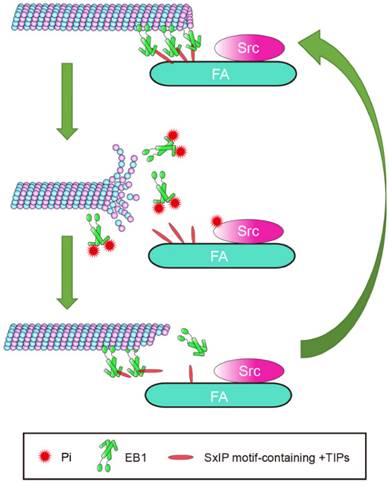
Cell migration is a complex process in which cytoskeletal remodeling must be coordinated with polarization signals to generate polarized cell branches and directed movement. EB1, as a scaffold that binds multiple +TIPs, is well-positioned to efficiently facilitate changes in MT dynamics through regulated associations with other +TIPs. In this study, we demonstrated that phosphorylation of EB1 by Src at Y247 suppresses the interactions between EB1 and SxIP motif-containing +TIPs. Because of the reversible nature of phosphorylation, temporary suppression of these interactions did not abolish MT growth, but instead increased growth velocity and enhanced MT dynamics, resulting in the promotion of cell migration. Based on our findings, we proposed a model describing the effect of Src-mediated EB1 phosphorylation on MT dynamics in the vicinity of FAs (Figure 7). In this model, the phosphorylation of EB1 by Src disrupts the interactions between EB1 and SxIP motif-containing +TIPs and impairs the MT plus end-tracking activity of EB1, rendering MTs to undergo catastrophe and EB1 to dissociate from MTs.
Proposed model for the function of Src-mediated EB1 phosphorylation. EB1 tracks MT plus ends and interacts with SxIP motif-containing +TIPs localized at FAs, thereby temporarily stabilizing MTs. Activation of Src causes its phosphorylation of EB1 at Y247, which perturbs the interactions between EB1 and SxIP motif-containing +TIPs and impairs the MT plus end-tracking ability of EB1. Consequently, MTs undergo catastrophe and EB1 dissociates from MTs. As a result of the reversibility of phosphorylation, EB1 undergoes quick turnover and associates with MTs and +TIPs again, and MTs regrow.
Post-translational modifications of EB1 have previously been reported in a few studies. For example, during anaphase, Bim1p, the budding yeast homolog of EB1, is phosphorylated at serine residues by Aurora B/Ipl1p to ensure timely separation of the spindle midzone26. EB1 is also known to be acetylated at K220 by p300, and this modification regulates its interactions with kinetochore-associated +TIPs during metaphase to prevent anaphase onset until all the chromosomes are correctly attached by MTs27. Our laboratory previously identified with mass spectrometry a few phosphorylated residues in EB1, including Y21728. EB1 Y217D cannot dimerize or interact with other +TIPs, suggesting that phosphorylation of this residue may help maintain the equilibrium between the monomer and dimer forms of EB1, thereby regulating MT dynamics28. In this study, we report that EB1 phosphorylation at Y247 reduces its binding to SxIP motif-containing +TIPs, leading to the promotion of cell migration through an increase in MT dynamics. Different modifications of EB1 may coordinate to provide a means for fine-tuning MT dynamics in specific cellular events.
Our results indicate that Y247 is the primary Src phosphorylation site in EB1. Y247 is located in the EBH domain and forms two hydrogen bonds, with E225 and T249, respectively. Mutation of Y247 to either phenylalanine (F) or aspartic acid (D) abolishes hydrogen bonding with E225, which likely alters the stability of the hydrophobic pocket where SxIP motif-containing +TIPs bind EB1. Thus, the Y247F mutant of EB1 is not simply a nonphosphorylatable mutant. This may help explain why both the Y247F mutant and the Y247D mutant of EB1 impair its binding to APC-C and MCAK (Figures 4B and 4C) and increase the rate of MT catastrophe (Figures 5B and 5C).
Notably, our in vitro kinase assays indicate that in addition to Y247, other tyrosine residues, especially Y268, could potentially be phosphorylated. Y268 is a part of the glutamate-glutamate-tyrosine (EEY) motif of EB1 that is responsible for interactions with CAP-Gly domain-containing +TIPs11. Phosphorylation of this residue might affect the binding of EB1 to CAP-Gly domain-containing +TIPs, such as cytoplasmic linker protein 170 (CLIP-170) and p150Glued, leading to MT destabilization at the cell cortex through a mechanism similar to what we have proposed for Y247 phosphorylation. It will be interesting to examine in the future the effects of the Y268 mutant.
Interestingly, our data revealed that a subset of pY247-EB1 localized to the centrosome, suggesting that Src-mediated EB1 phosphorylation may regulate centrosome functions. Both EB1 and APC have been shown to localize to the centrosome independently of their plus end-tracking property. EB1 knockdown has been demonstrated to suppress MT nucleation and regrowth from the centrosome29. Loss of EB1 and EB3 inhibits the assembly of the primary cilium, likely due to centrosome-associated mechanisms30. In addition, androgen and Src signaling have been reported to regulate MT nucleation and promote the accumulation of γ-tubulin at the centrosome19. Thus, it is tempting to speculate that EB1, as a downstream factor of Src, may play a role in the regulation of MT nucleation and γ-tubulin recruitment to the centrosome.
We observed that the largest proportion of pY247-EB1 localized to FAs. MTs are known to be targeted to FAs through EB1 and its associated +TIPs. Thus, EB1 and other +TIPs are poised to play a central role in coordinating the MT-FA crosstalk. APC, CLASPs, and MACF1/ACF7, three members of the SxIP motif-containing +TIPs, normally capture and stabilize MTs targeting FAs through interactions with EB12,14,15. Despite these stabilizing interactions, the elevated MT catastrophe rate at FAs suggests that FA-associated MTs undergo constant transitions between growth and shrinkage that are induced by a heretofore undefined factor. Src-mediated phosphorylation of EB1 at FAs is a likely candidate for this role, because Y247D-EB1 not only reduces the binding of EB1 to SxIP motif-containing +TIPs, but also decreases the growth lifetime and increases the catastrophe rate of MTs. Consistent with these findings, we also demonstrate that Y247D-EB1 enhances cell migration. Thus, Src-mediated EB1 phosphorylation may promote cell migration through regulating the dynamic crosstalk between MT and FAs.
Abbreviations
MT, microtubule; FA, focal adhesion; EB1, end-binding protein 1; +TIP, MT plus end-tracking protein; CLASP, cytoplasmic linker-associated protein; FAK, FA kinase; CAP-Gly, cytoskeleton-associated protein glycine-rich; APC, adenomatous polyposis coli; MACF1, MT-actin crosslinking factor 1; ACF7, actin crosslinking family 7; MCAK, mitotic centromere associated kinesin; HEK293T, human embryonic kidney 293T; HUVEC, human umbilical vein endothelial cells; KLH, keyhole limpet hemocyanin; CLIP-170, cytoplasmic linker protein 170.
Acknowledgements
We thank Drs. Quan Chen and Yushan Zhu for reagents. This work was supported by grants from the National Natural Science Foundation of China (31130015, 31471262, 31271437, and 31371382) and the National Basic Research Program of China (2012CB945002).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Kaverina I, Straube A. Regulation of cell migration by dynamic microtubules. Semin Cell Dev Biol. 2011;22:968-74
2. Wen Y, Eng CH, Schmoranzer J. et al. EB1 and APC bind to mDia to stabilize microtubules downstream of Rho and promote cell migration. Nat Cell Biol. 2004;6:820-30
3. Efimov A, Schiefermeier N, Grigoriev I. et al. Paxillin-dependent stimulation of microtubule catastrophes at focal adhesion sites. J Cell Sci. 2008;121:196-204
4. Stehbens S, Wittmann T. Targeting and transport: How microtubules control focal adhesion dynamics. J Cell Biol. 2012;198:481-9
5. Gao J, Huo L, Sun X. et al. The tumor suppressor CYLD regulates microtubule dynamics and plays a role in cell migration. J Biol Chem. 2008;283:8802-9
6. Stehbens SJ, Matthew P, Hayley P. et al. CLASPs link focal-adhesion-associated microtubule capture to localized exocytosis and adhesion site turnover. Nat Cell Biol. 2014;16:561-73
7. Ezratty EJ, Partridge MA, Gundersen GG. Microtubule-induced focal adhesion disassembly is mediated by dynamin and focal adhesion kinase. Nat Cell Biol. 2005;7:581-90
8. Kaverina I, Rottner K, Small JV. Targeting, capture, and stabilization of microtubules at early focal adhesions. J Cell Biol. 1998;142:181-90
9. Gouveia SM, Akhmanova A. Cell and molecular biology of microtubule plus end tracking proteins: end binding proteins and their partners. Int Rev Cell Mol Biol. 2010;285:1-74
10. Honnappa S, Gouveia SM, Weisbrich A. et al. An EB1-Binding Motif Acts as a Microtubule Tip Localization Signal. Cell. 2009;138:366-76
11. Bjelic S, De Groot CO, Scharer MA. et al. Interaction of mammalian end binding proteins with CAP-Gly domains of CLIP-170 and p150(glued). J Struct Biol. 2012;177:160-7
12. Li D, Gao J, Yang Y. et al. CYLD coordinates with EB1 to regulate microtubule dynamics and cell migration. Cell Cycle. 2014;13:974-83
13. Li D, Xie S, Ren Y. et al. Microtubule-associated deacetylase HDAC6 promotes angiogenesis by regulating cell migration in an EB1-dependent manner. Protein Cell. 2011;2:150-60
14. Kodama A, Karakesisoglou L, Wong E. et al. ACF7: An essential integrator of microtubule dynamics. Cell. 2003;115:343-54
15. Mimori-Kiyosue Y, Grigoriev I, Lansbergen G. et al. CLASP1 and CLASP2 bind to EB1 and regulate microtubule plus-end dynamics at the cell cortex. J Cell Biol. 2005;168:141-53
16. Sefton BM, Hunter T, Ball EH. et al. Vinculin: A cytoskeletal target of the transforming protein of rous sarcoma virus. Cell. 1981;24:165-74
17. Webb DJ, Donais K, Whitmore LA. et al. FAK-Src signalling through paxillin, ERK and MLCK regulates adhesion disassembly. Nat Cell Biol. 2004;6:154-61
18. Zhang Z, Izaguirre G, Lin SY. et al. The phosphorylation of vinculin on tyrosine residues 100 and 1065, mediated by Src kinases, affects cell spreading. Mol Biol Cell. 2004;15:4234-47
19. Colello D, Reverte CG, Ward R. et al. Androgen and Src signaling regulate centrosome activity. J Cell Sci. 2010;123:2094-102
20. Sun L, Gao J, Dong X. et al. EB1 promotes Aurora-B kinase activity through blocking its inactivation by protein phosphatase 2A. Proc Natl Acad Sci U S A. 2008;105:7153-8
21. Yang Y, Liu M, Li D. et al. CYLD regulates spindle orientation by stabilizing astral microtubules and promoting dishevelled-NuMA-dynein/dynactin complex formation. Proc Natl Acad Sci U S A. 2014;111:2158-63
22. Yang Y, Ran J, Liu M. et al. CYLD mediates ciliogenesis in multiple organs by deubiquitinating Cep70 and inactivating HDAC6. Cell Res. 2014;24:1342-53
23. Gao J, Sun L, Huo L. et al. CYLD regulates angiogenesis by mediating vascular endothelial cell migration. Blood. 2010;115:4130-7
24. Applegate KT, Sebastien B, Alexandre M. et al. plusTipTracker: Quantitative image analysis software for the measurement of microtubule dynamics. J Struct Biol. 2011;176:168-84
25. Braun A, Dang K, Buslig F. et al. Rac1 and Aurora A regulate MCAK to polarize microtubule growth in migrating endothelial cells. J Cell Biol. 2014;206:97-112
26. Zimniak T, Stengl K, Mechtler K. et al. Phosphoregulation of the budding yeast EB1 homologue Bim1p by Aurora/Ipl1p. J Cell Biol. 2009;186:379-91
27. Xia P, Wang ZK, Liu X. et al. EB1 acetylation by P300/CBP-associated factor (PCAF) ensures accurate kinetochore-microtubule interactions in mitosis. Proc Natl Acad Sci U S A. 2012;109:16564-9
28. Chen J, Luo Y, Li L. et al. Phosphoregulation of the dimerization and functions of end-binding protein 1. Protein Cell. 2014;5:795-9
29. Louie RK, Shirin B, Siemers KA. et al. Adenomatous polyposis coli and EB1 localize in close proximity of the mother centriole and EB1 is a functional component of centrosomes. J Cell Sci. 2004;117:1117-28
30. Schroder JM, Larsen J, Komarova Y. et al. EB1 and EB3 promote cilia biogenesis by several centrosome-related mechanisms. J Cell Sci. 2011;124:2539-51
Author contact
Corresponding author: Jun Zhou, State Key Laboratory of Medicinal Chemical Biology, College of Life Sciences, Nankai University, Tianjin 300071, China. Telephone: +86-22-2349-4816; Fax: +86-22-2349-4816; junzhouedu.cn.
 Global reach, higher impact
Global reach, higher impact