Impact Factor
Theranostics 2017; 7(6):1437-1446. doi:10.7150/thno.16558 This issue Cite
Research Paper
Cross-Platform Comparison of Four Leading Technologies for Detecting EGFR Mutations in Circulating Tumor DNA from Non-Small Cell Lung Carcinoma Patient Plasma
1. Key Laboratory of Carcinogenesis and Translational Research (Ministry of Education), Department of Clinical Laboratory, Peking University Cancer Hospital & Institute, Beijing, China;
2. Key Laboratory of Carcinogenesis and Translational Research (Ministry of Education), Department of Thoracic Surgery I, Peking University Cancer Hospital & Institute, Beijing, China;
3. Shanghai AccuraGen Biotechnology Co. Ltd, China.
*Ting Xu and Xiaozheng Kang contributed equally to this study.
#Ke-Neng Chen and Guobing Xu jointly supervised this work.
Received 2016-6-21; Accepted 2017-1-22; Published 2017-4-2
Abstract
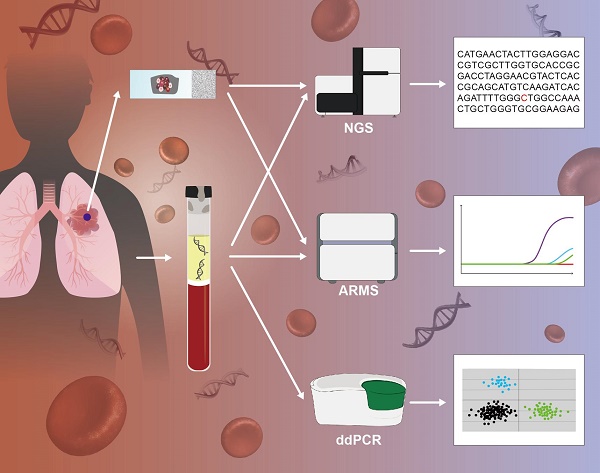
Analysis of circulating tumor DNA (ctDNA) is emerging as a powerful tool for guiding targeted therapy and monitoring tumor evolution in patients with non-small cell lung cancer (NSCLC), especially when representative tissue biopsies are not available. Here, we have compared the ability of four leading technology platforms to detect epidermal growth factor receptor (EGFR) mutations (L858R, exon 19 deletion, T790M and G719X) in ctDNA from NSCLC patients. Two amplification refractory mutation systems (cobas-ARMS and ADx-ARMS), a droplet digital polymerase chain reaction (ddPCR) and a next-generation sequencing (Firefly NGS) platform were included in the comparison. Fifteen EGFR mutations across twenty NSCLC patients were identified. Firefly NGS, cobas-ARMS and ddPCR all displayed superior sensitivity while ADx-ARMS was better suited for the qualitative detection of EGFR mutations with allele frequency higher than 1% in plasma and tissue samples. We observed high coincidence between the plasma and tissue EGFR mutational profiles for three driver mutations (L858R, exon 19 deletion and G719X) that are known targets of first generation EGFR-TKI therapies among patients who relapsed. Discrepancies between tissue and plasma EGFR mutational profiles were mainly attributable to spatial and temporal tumor heterogeneity, mutation inhibition due to therapy response and drug resistance (T790M). This study illustrates the challenges associated with selection of a technology platform for EGFR ctDNA analysis in the context of treatment evaluation and drug resistance detection.
Keywords: non-small cell lung cancer, epidermal growth factor receptor, quantitative real-time polymerase chain reaction, next-generation sequencing.
Introduction
Somatic mutations in epidermal growth factor receptor (EGFR) play a significant role as both biomarkers and rational targets for targeted therapy in non-small cell lung cancer (NSCLC) [1, 2]. New therapies, such as EGFR-TKIs, are emerging as treatment for EGFR-mutated NSCLC [3], and an increasing number of patients are receiving multiple lines of EGFR-targeted therapy. At the same time, the evolutionary trajectory of tumor subclones under selection pressures, such as EGFR inhibition, with either intrinsic or acquired resistance can be selected and drive disease progression. Thus, there is increasing demand for accurate EGFR mutation profiles that will help to inform therapeutic decision-making.
Current recommendations are to detect EGFR mutations in fresh, frozen or formalin-fixed paraffin-embedded (FFPE) tissue from a solid biopsy. However, nearly two-thirds of patients with NSCLC are already at an advanced stage at the time of diagnosis, and surgical biopsy procedures are usually not feasible. Therefore, sufficient tumor tissues for multiplex EGFR mutations analysis are difficult to obtain [4]. Additionally, because of spatial and temporal heterogeneity, mutation analysis from initial tumor tissue samples at diagnosis is not suitable for therapeutic guidance throughout the entire process of treatment, especially after disease progression. For these reasons, new methods for deriving EGFR mutational profiles from alternative sources, such as plasma [5-7], pleural fluid [8], sputum [9], urine [10, 11] and cerebrospinal fluid [12, 13] which collectively are often referred to as 'liquid biopsies', are being developed.
Of the alternative sources tested, plasma has been shown to be most promising [14]. The abundance of circulating tumor DNA (ctDNA) varies from 0.1% to 67% in patients with regional and more advanced cancer, and is even lower (0.01%) in patients with localized cancer [15-18]. Thus, a highly sensitive method for detection of EGFR mutations in ctDNA is essential. Multiple studies have reported that EGFR mutations can be detected in ctDNA using diverse technologies [19]. Compared with the detection of EGFR mutations from tissue samples, ctDNA tests have a sensitivity of 46%-82%, a specificity of 90%-99% and total coincidence rate of 78%-88% in assessing EGFR mutations [20].
Commonly used technologies for plasma EGFR analysis include amplification refractory mutation system (ARMS), droplet digital PCR (ddPCR) and next-generation sequencing (NGS)-based methods [21-23]. These methods differ significantly in sensitivity, specificity and coverage of EGFR mutations [24, 25]. ADx-ARMS (ADx® EGFR 29 Mutations Detection Kit), which can detect 1% mutant DNA in a background of 99% normal DNA [26], has been approved for clinical use in China by the China Federal Drug Administration (CFDA). Cobas-ARMS (cobas® EGFR Mutation Test v2) has been approved by the U. S. Food and Drug Administration (FDA) for the detection of exon 19 deletions and L858R substitution mutations in plasma. In contrast, ddPCR and NGS-based methods have not been approved by the FDA or CFDA, but are widely used in research settings due to their quantitative advantage. While ddPCR has unparalleled sensitivity (0.04%-0.1%) [7, 27], it can detect only one locus per reaction well limiting its use in multiplex tests. NGS-based methods, such as Firefly NGS, have demonstrated performance sensitivity comparable to that of ddPCR without the same limitation in multiplex testing.
A performance comparison of these various technologies in detecting EGFR mutations in ctDNA is crucial for optimizing the use of liquid biopsies for NSCLC in a clinical setting. Here, we report a performance comparison between four techniques (ADx-ARMS, cobas-ARMS, ddPCR and Firefly NGS) in detecting clinically-relevant EGFR mutations in tumor tissue and plasma collected from NSCLC patients.
Materials and Methods
Tissue and blood samples
Tissue and blood samples were obtained from 20 NSCLC patients (age range 37-76 years) treated at the Department of Thoracic Surgery I, Peking University Cancer Hospital & Institute, from July 2009 to November 2015. Patients were considered for inclusion if they met the following criteria: were newly diagnosed with advanced or recurrent or multiple primary NSCLC, had available surgical resected tumor tissue samples and a minimum of 205ng of cfDNA, had complete medical documentation including follow-up records and EGFR status of tissue tested by ADx-ARMS. To include various scenarios a clinician may encounter in daily practice and maximize the representation of all plasma EGFR mutation levels, we enrolled patients with different clinical responses during treatment. To increase representation of drug resistance mutations such as T790M, we included patients that were receiving treatment with TKI. The study was conducted in accordance with Declaration of Helsinki principles, and approved by the Institutional Review Board of Peking University Cancer Hospital & Institute. All subjects gave written informed consent.
Tissue samples included resected primary or metastatic pulmonary, metastatic lymph nodal or pleural lesions. The percentage of tumor cells out of all nucleated cells was assessed in a 4 μm section after hematoxylin and eosin staining by a senior pathologist. Samples with at least 10% tumor cells were considered for further tissue DNA analysis through standard NGS; patients with less than 10% tumor cells in the tissue samples underwent only cfDNA analysis. Treatment efficacy was evaluated by Response Evaluation Criteria in Solid Tumors (RECIST) version 1.1 [28].
In total, 20 ml of venous blood was collected into two 10-mL Streck Cell-Free DNA BCT tubes (Streck Co., Omaha, NE, USA). Within two hours of blood collection, blood was centrifuged at 1,900 g for 10 minutes at 4°C to separate the plasma from the peripheral blood cells. The plasma was then further centrifuged at 16,000 g for 10 minutes at 4°C to pellet any remaining cells. Then, the upper phase from plasma and the peripheral blood cells were collected in several new 5-mL sterile Eppendorf tubes. The peripheral blood cells from the first spin were used for isolation of germline genomic DNA from peripheral blood leukocytes [29]. All tubes were labeled with the corresponding patient name and the date of sample collection and immediately stored at -80°C until DNA isolation.
Tumor tissue, white blood cells genomic DNA and cfDNA extraction
Tumor tissue genomic DNA was extracted from 10 FFPE slides using the QIAamp DNA FFPE Tissue Kit (Qiagen, Venlo, The Netherlands). White blood cells genomic DNA was extracted from 100ul of buffy coat using the QIAamp DNA Blood Maxi Kit (Qiagen, Venlo, The Netherlands). cfDNA was extracted from 8.0 mL - 10.8 mL of plasma using the QIAamp Circulating Nucleic Acid Kit (Qiagen, Venlo, The Netherlands). All DNA was extracted in accordance with manufacturer's instructions and quantified using a Qubit fluorometer (Life Technologies, Carlsbad, CA, USA). Extracted DNA was stored at -80°C until use.
Standard NGS analysis for genomic DNA from white blood cells and tumor tissue
Genomic DNA was sonicated into fragments with a peak length of 200bp. 100ng of fragmented genomic DNA was used for NGS library construction using KAPA sequencing library construction kit (Kapa Biosystems, Boston, MA, USA). Genomic DNA NGS library was then captured by Accu-Act panel (AccuraGen Inc., Shanghai, China) and sequenced with 100 bp paired-end runs on an Illumina HiSeq 2500 system (Illumina, San Diego, CA, USA). The average coverage depth for all probes was at least 500X.
ADx-ARMS for genomic DNA from tumor tissue
EGFR status of tumor tissue using ADx-ARMS assay was obtained from patient records stored at Peking University Cancer Hospital & Institute. ADx-ARMS protocol for tissue genomic DNA was the same as that for cfDNA.
ADx-ARMS, cobas-ARMS, ddPCR and Firefly NGS analysis for cfDNA from plasma
Two AmoyDx® EGFR 29 Mutations Detection Kits (ADx-ARMS) and one cobas® EGFR Mutation Test v2 (cobas-ARMS) were used for plasma EGFR detection in accordance with the manufacturer's instructions. The ddPCR assays (L858R, exon 19 deletion, T790M and G719X) were performed using a QX200 Droplet Digital PCR System (Bio-Rad Laboratories Inc., Hercules, CA, USA). Allele frequencies were analyzed using QuantaSoft v1.6 (Bio-Rad Laboratories Inc., Hercules, CA, USA). Firefly NGS (Accu-Act) assay was performed based on AccuraGen's circularized cfDNA technology (AccuraGen Inc., Shanghai, China). Details of these four methods, determination of the EGFR mutant and wild type loci and coincidence rates are supplied in Supplementary Materials and Methods.
Statistical analysis
Fisher's exact test was used to compare the detection rates of the four platforms and the positive and negative coincidence rates between six pairs. Paired t-test was used to compare the sensitivity and specificity of the four platforms in detecting four types of EGFR mutations. Spearman's rank correlation coefficient was used to determine the strength of association between the allele frequencies of plasma EGFR mutations determined via ddPCR and Firefly NGS. Differences were considered statistically significant when p < 0.05. All calculations were performed with SPSS for Windows (version 19.0; SPSS Inc., Chicago, IL, USA), and figures were created using GraphPad Prism (version 5.0; GraphPad, San Diego, CA).
Results
Patient characteristics
A total of twenty NSCLC patients were enrolled, of whom thirteen (65%) with regional (n = 8) or distant (n = 5) recurrence after surgery, and seven (35%) with distant metastatic disease (n = 6) or multiple primary lung cancer (n = 1). The spatial and temporal characteristics of tissue and plasma sampling are shown in Table 1 and Supplementary Table S1. Blood samples from fifteen patients (75%) were obtained during systemic therapy, and the other five were obtained prior to the initiation of treatment.
The characteristics of twenty NSCLC patients enrolled in the study.
| n (%) | ||
|---|---|---|
| Gender | Male | 10 (50) |
| Female | 10 (50) | |
| Age (median, range) (years) | 60.5, 37-76 | |
| Smoking status | Never smoker | 13 (65) |
| Ever smokers | 6 (30) | |
| Unknown | 1 (5) | |
| Histology | ADC | 18 (90) |
| SCC | 1 (5) | |
| ASC | 1 (5) | |
| Extent of disease | Local regional recurrence | 8 (40) |
| Distant recurrence | 5 (25) | |
| Distant metastatic | 6 (30) 1 (5) | |
| Multiple primary cancer | ||
| Stage (tissue/plasma) a | Stage I | 5 (25)/3 (15) |
| Stage II | 2 (10)/0 (0) | |
| Stage III | 6 (30)/0 (0) | |
| Stage IV | 7 (35)/17 (85) | |
| Time interval between tissue and blood sampling(years) | 0-1 | 7 (35) |
| 1-3 | 5 (25) | |
| 3-6 | 6 (30) | |
| ≥6 | 2 (10) | |
| During systemic treatment | Yes | 14 (70) |
| No | 6 (30) | |
| Response evaluation | PR | 4 (20) |
| SD | 13 (65) | |
| PD | 3 (15) | |
| EGFR status at diagnosis | L858R | 7 (35)b |
| Exon 19 deletion | 9 (45)b,c | |
| T790M | 1 (5)c | |
| Negative | 5 (25) |
a, 7th edition AJCC/UICC TNM staging system for NSCLC was applied accordingly. The stage of tissue and plasma sampling time-point was recorded respectively; b, the L858R/exon 19 deletion double mutation was found in one case; c, the exon 19 deletion/T790M double mutation was found in one case.
Detection rates of ADx-ARMS, cobas-ARMS, ddPCR and Firefly NGS platforms
Fifteen EGFR mutations were found among 20 patients across four platforms. The overall detection rate of ADx-ARMS, cobas-ARMS, ddPCR and Firefly NGS was 46.7% (7/15), 73.3% (11/15), 66.7% (10/15) and 80% (12/15), respectively (χ2 = 3.972, p = 0.318) (Figure 1A).
The overall (A) and detection rate in lower (allele frequency <1%) (B) and higher allele frequency subgroup (allele frequency ≥1%) (C). If a positive result was obtained from any one of the four platforms, the sample was categorized as positive. Two cases, in which the allele frequencies were unknown, were excluded from Figure 1B and 1C (S2 ADx-ARMS and S9 cobas-ARMS results). Values represent n. * denotes statistically significant difference (p < 0.05).
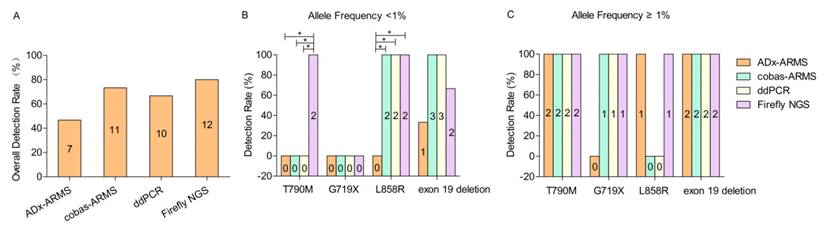
According to the allele frequency (< 1% vs. ≥ 1%), thirteen EGFR alterations were divided into two subgroups (7 vs. 6), and the other two EGFR alterations with unknown allele frequencies tested solely by ADx-ARMS (S2 in table 2) or cobas-ARMS (S9 in table 2) were excluded. In the lower allele frequency subgroup, the detection rate of ADx-ARMS, cobas-ARMS, ddPCR and Firefly NGS was 14.3% (1/7), 71.4% (5/7), 71.4% (5/7) and 85.7% (6/7), respectively (χ2 = 8.138, p = 0.047). Though the detection rate of ADx-ARMS was significantly lower than that of the other three platforms, there was no significant performance difference between cobas-ARMS, ddPCR and Firefly NGS. In the higher allele frequency subgroup, the detection rate of ADx-ARMS, cobas-ARMS, ddPCR and Firefly NGS was 83.3% (5/6), 83.3% (5/6), 83.3% (5/6) and 100% (6/6), respectively. There was no statistically significant difference between the ability of all four platforms to detect each of the four EGFR mutation types (Figure 1C).
EGFR mutation profiles in tissue and plasma.
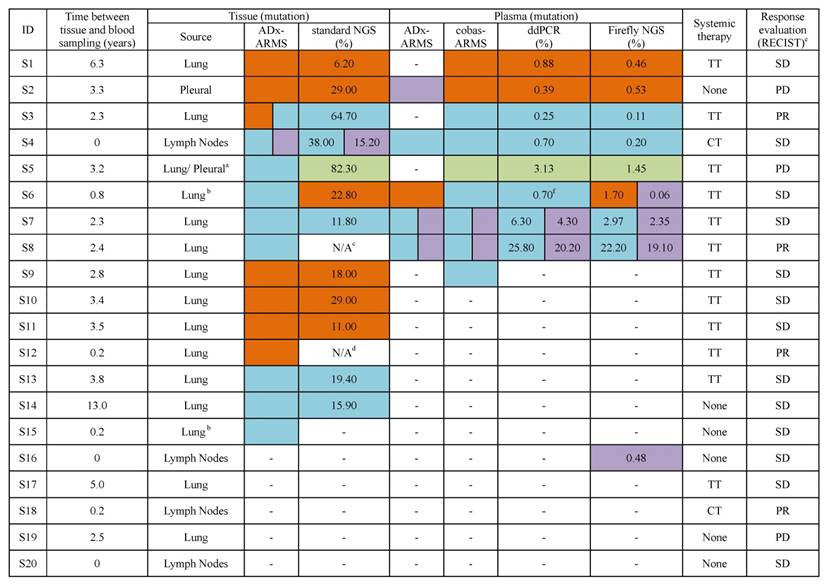
 L858R mutation, the number represents the allele frequency of the L858R mutation;
L858R mutation, the number represents the allele frequency of the L858R mutation;  Exon 19 deletion mutation, the number represents the allele frequency of the exon 19 deletion mutation;
Exon 19 deletion mutation, the number represents the allele frequency of the exon 19 deletion mutation;  T790M mutation, the number represents the allele frequency of the T790M mutation;
T790M mutation, the number represents the allele frequency of the T790M mutation;  G719X mutation, the number represents the allele frequency of the G719X mutation. a, the primary tumor tissue was used for ADx-ARMS, but the pleural metastasis sample was used for standard NGS; b, the patient had multiple primary tumors; c, the amount of DNA was not sufficient for standard NGS; d, there was no tumor content in the slide; e, the first radiologic evaluation after this plasma sampling; f, L858R was positive in an additional ddPCR testing.
G719X mutation, the number represents the allele frequency of the G719X mutation. a, the primary tumor tissue was used for ADx-ARMS, but the pleural metastasis sample was used for standard NGS; b, the patient had multiple primary tumors; c, the amount of DNA was not sufficient for standard NGS; d, there was no tumor content in the slide; e, the first radiologic evaluation after this plasma sampling; f, L858R was positive in an additional ddPCR testing.
N/A, result of standard NGS was not available; TT, targeted therapy, targeted drugs that patients took in this study are first generation EGFR-TKIs; CT, chemotherapy; None, the patient took no systemic therapy when blood was collected; RECIST, Response Evaluation Criteria in Solid Tumors; PR, partial response; SD, stable disease; PD, progressive disease.
Positive and negative coincidence rates across platforms
In the lower allele frequency subgroup, the positive coincidence rate of each pair-wise platform comparison ranged from 16.7% to 100%. Those three pairs, ADx-ARMS vs. cobas-ARMS, ADx-ARMS vs. ddPCR and ADx-ARMS vs. Firefly NGS, presented the lowest positive coincidence with rates of 20% (1/5), 20% (1/5) and 16.7% (1/6), respectively (χ2 = 0.484, p = 1.000). With the exclusion of sample S9 due to lack of allele frequency data, cobas-ARMS and ddPCR demonstrated the highest positive coincidence rate (100%, 5/5) (Figure 2A). In the higher allele frequency subgroup, the positive coincidence rate of each pair-wise platform comparison ranged from 66.7% to 100%. No statistically significant difference was found among the six pairs. The platform pair with the greatest concordance was again cobas-ARMS and ddPCR, yielding identical results (100%, 5/5) (Figure 2A).
The positive (A) and negative (B) coincidence rates between six paired comparisons. Values represent n. * denotes statistically significant difference (p < 0.05).
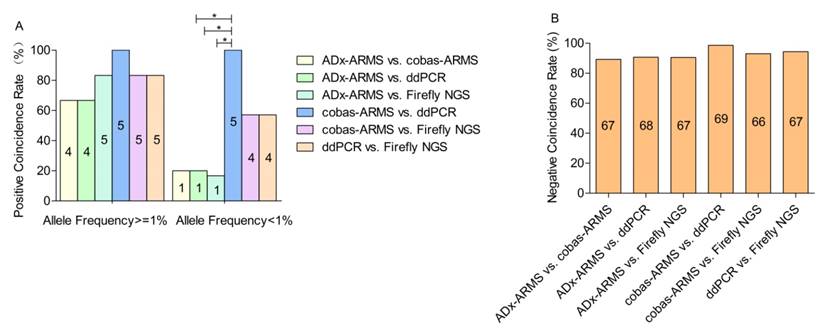
A total of seventy-five EGFR wild-type loci were detected in this study. ADx-ARMS, cobas-ARMS, ddPCR and Firefly NGS uncovered 73, 69, 70 and 68 EGFR wild-type loci, respectively. The negative coincidence rate of each pair-wise comparison was all higher than 90%. No statistically significant difference was found among the six paired comparisons (Figure 2B).
Coincidence between ADx-ARMS and standard NGS in tumor tissue analysis
EGFR mutation analysis was conducted in tumor tissue using two methods: ADx-ARMS and standard NGS. ADx-ARMS analysis was performed at the time of initial tumor biopsy, and patients had been treated based on this result. Standard NGS was performed at a later date on subsequent slides for all patients (S1- S20); two of which had insufficient tissue genomic DNA (S8) or no tumor content in the slide (S12) for standard NGS analysis, and three of which (S5, S6 and S15) had spatially (S6 and S15) or temporally (S5) different tissues sampling for ADx-ARMS and standard NGS assay. Thus, we analyzed the coincidence between the historical ADx-ARMS and the current NGS-based EGFR mutation results in fifteen patients. We observed a coincidence of 93.3% (14/15) between the two methods (Table 2); however, in one case (S3), L858R and exon 19 deletion double mutations were identified by ADx-ARMS, but only the exon 19 deletion was identified by standard NGS.
Comparison of EGFR mutational profiles from plasma and tumor tissue
We compared the EGFR mutational profiles observed in plasma and tumor tissue samples for all twenty cases. The time between tissue and blood sampling ranged from 0 to 13 years, with seven (35%) patients sampled 0-1 year apart, five (25%) sampled 1-3 years apart, six (30%) sampled 3-6 years apart, and two (10%) sampled ≥ 6 years apart (Table 2).
The four platforms detected a total of three L858Rs, six exon 19 deletions, five T790Ms and one G719X across the cohort. Meanwhile, three L858Rs, five exon 19 deletions and one G719X were definitely diagnosed in the matched tissues. In order to determine the coincidence between tissue and plasma EGFR mutational profiles, we assessed how often EGFR mutations were observed in plasma but not in tumor tissue. For cobas-ARMS, ddPCR and Firefly NGS, the L858R, exon 19 deletion and G719X mutations observed in plasma were also seen in tumor tissue samples, demonstrating a high coincidence between plasma and tumor tissue mutational profiles. There was one disagreement of the exon 19 deletion, which was identified only by cobas-ARMS in plasma, whereas the tissue assay result was L858R (S9). In addition, the plasma mutational profiles varied across the three technology platforms in only one case (S6).
A different pattern was observed for the T790M mutation, a well-characterized drug resistance mutation that has been shown to arise in response to treatment [30]. There were five disagreements of T790M, of which one was detected by ADx-ARMS only (S2), two were detected by Firefly NGS only (S6, S16), and two were detected by all four platforms (S7, S8). Of note, patients S7 and S8 were both offered first generation EGFR-TKIs. Within five months post-sampling, S7 developed brain metastasis, and S8 died of lung cancer.
Finally, ten patients with EGFR wild-type in plasma, carried L858R (S10 - S12), exon 19 deletion (S13 - S15) or wild-type (S17 - S20) in their paired tissue samples (Table 2).
Total coincidence rate of EGFR variants in tumor tissue and plasma
Seventeen patients with stage IV disease at blood sampling were enrolled to determine the total coincidence rates of EGFR mutational profiles between tissue and plasma. For patients S5 and S6, their tissue test results from ADx-ARMS and standard NGS were inconsistent because the samples were collected at different times and from different tumors. ADx-ARMS and standard NGS data of patient S5 and S6 was both used as the reference, but one case with two different EGFR mutation types using ADx-ARMS and standard NGS was considered as two cases with only one EGFR mutation type each. For the other fifteen patients, the tissue test results obtained from ADx-ARMS were used as the reference. However, two patients with double mutations (S3, S4) were also considered as four cases with only one EGFR mutation type each. As a result, twenty-one cases were analyzed. Using the tissue EGFR mutational profiles for three driver mutations (L858R, exon 19 deletion and G719X) as a reference, the sensitivity, specificity and total coincidence rate of ctDNA assays were 26.7% - 53.3%, 100% and 45% - 65%, respectively (Supplementary Table S2). Of note, the sensitivity of the exon 19 deletion assay (50% - 83.3%) was higher than that of the L858R (12.5% - 37.5%, p = 0.008) and T790M (0%, p = 0.003) assays. Furthermore, the T790M assay had a lower specificity (80.0% - 90.0%), compared with the L858R (100%, p = 0.010) and exon 19 deletion (93.3% - 100%, p = 0.043) assays (Table 3).
Performance of four ctDNA test platforms for detection of EGFR mutation profiles in twenty-one* plasma samples.
| ADx-ARMS | cobas-ARMS | ddPCR | Firefly NGS | |
|---|---|---|---|---|
| L858R | ||||
| Sensitivity | 12.5% (1/8) | 25% (2/8) | 25% (2/8) | 37.5% (3/8) |
| Specificity | 100% (13/13) | 100% (13/13) | 100% (13/13) | 100% (13/13) |
| Total Coincidence Rate | 66.7% (14/21) | 71.4% (15/21) | 71.4% (15/21) | 76.2% (16/21) |
| Exon19 deletion | ||||
| Sensitivity | 50% (3/6) | 83.3% (5/6) | 83.3% (5/6) | 66.7% (4/6) |
| Specificity | 100% (15/15) | 93.3% (14/15) | 100% (15/15) | 100% (15/15) |
| Total Coincidence Rate | 85.7% (18/21) | 90.5% (19/21) | 95.2% (20/21) | 90.5% (19/21) |
| T790M | ||||
| Sensitivity | 0% (0/1) | 0% (0/1) | 0% (0/1) | 0% (0/1) |
| Specificity | 85.0% (17/20) | 90.0% (18/20) | 90.0% (18/20) | 80.0% (16/20) |
| Total Coincidence Rate | 81.0% (17/21) | 85.7% (18/21) | 85.7% (18/21) | 76.2% (16/21) |
| G719X | ||||
| Sensitivity | 0% (0/1) | 100% (1/1) | 100% (1/1) | 100% (1/1) |
| Specificity | 100% (20/20) | 100% (20/20) | 100% (20/20) | 100% (20/20) |
| Total Coincidence Rate | 95.2% (20/21) | 100% (21/21) | 100% (21/21) | 100% (21/21) |
*Seventeen patients with stage IV disease at blood sampling were enrolled to determine the total coincidence rates of EGFR mutation profiles between tissue and plasma. For patients S5 and S6, their tissue test results from ADx-ARMS and standard NGS were both used as the reference, but one case with two different EGFR mutation types using ADx-ARMS and standard NGS was considered as two cases with only one EGFR mutation type each. For the other fifteen patients, the tissue test results obtained from ADx-ARMS were used as the reference. However, two patients with double mutations (S3, S4) were also considered as four cases with only one EGFR mutation type each. As a result, twenty-one cases were analyzed.
Discussion
In this study, we conducted a performance comparison of four leading platforms for detecting EGFR mutations with different allele frequencies. We demonstrated the superior sensitivity and specificity of cobas-ARMS, ddPCR and Firefly NGS platforms. In contrast, ADx-ARMS was better suited for the qualitative detection of EGFR mutations with allele frequency exceeding 1% in plasma and tissue samples. To our knowledge, the current study is the first comparison of two ARMS, a ddPCR and a NGS-based platform in this field.
We first compared the sensitivity of each technology platform in the higher allele frequency (≥1%) and lower allele frequency (<1%) subgroups. For the higher allele frequency variants, the four technology platforms demonstrated similar plasma detection rates ranging between 83.3% and 100%. However, there was a clear separation between detection rates when analyzing the lower allele frequency subgroup. The detection rates for the cobas-ARMS, ddPCR and Firefly NGS platforms ranged between 71.4% and 85.7%. In contrast, the detection rate of ADx-ARMS was only 14.3%. These results demonstrate that cobas-ARMS, ddPCR and Firefly NGS methods have higher detection limits compared with ADx-ARMS.
A high coincidence rate (91%, 10/11) was observed between cobas-ARMS and ddPCR in our study, in keeping with previous investigations [24]. Moreover, the two ctDNA quantitation methods, Firefly NGS and ddPCR, yielded excellent positive coincidence rates in this study, even for the analysis of low-template copies. The frequencies of ctDNA calculated by the two methods were highly correlated (Spearman Correlation, R2 = 0.98, p < 0.0001, Supplementary Figure S1), consistent with previous reports [31]. Thus, the Firefly NGS platform demonstrates equivalent sensitivity to ddPCR, the most sensitive platform for EGFR mutational profiling.
We compared the EGFR mutational profiles between the plasma and tumor tissue samples and observed diverging platform-specific behavior, which must be interpreted within the context of the assay's sensitivity and the inherent biological properties of the EGFR mutation of interest. First, among the three higher-sensitivity assays (cobas-ARMS, ddPCR and Firefly NGS), we observed a high coincidence between the plasma and tissue mutational profiles for three driver mutations that can be identified in primary tumor and are known targets of first generation EGFR-TKI therapies (L858R, exon 19 deletion and G719X). All L858R, exon 19 deletion and G719X mutations observed in plasma were confirmed in tissue samples. There was one discordance for the exon 19 deletion, which was identified only by cobas-ARMS in plasma, whereas only the L858R was detected in tumor tissue by both ADx-ARMS and standard NGS (S9). This result can be explained by sampling error, but is likely a false positive because it was not detected by the assays with higher sensitivity.
A second pattern of discordances emerged in the analysis of T790M mutation. We observed five discordances for T790M, of which two were detected by all four platforms (S7, S8), two were detected by Firefly NGS only (S6, S16), and one was detected by ADx-ARMS only (S2). The T790M mutation has been shown to appear in response to first generation EGFR-TKI therapies [30]. In Patients S7 and S8, the T790M mutations were observed in plasma across all four platforms, and is likely reflective of acquired first generation EGFR-TKI resistance, as described previously [32].
Two patients (S6, S16) were characterized as T790M-positive using Firefly NGS, but T790M-negative using the other three methods. In the case of S16, T790M might represent a false-positive based on a subsequent ddPCR-negative result. This false positive may be explained by either the inherent biological properties of the sample, as T790M transition mutations have been shown to arise from the spontaneous deamination of cytosine and 5-mC [33], or a technical artifact caused by assay or algorithmic features. In the case of S6, insufficient quantities of cfDNA inhibited the verification of T790M through additional ddPCR testing. Given the high background signal (0.5%) of T790M in peripheral blood specimens from normal donors [34], the detection of T790M may be mitigated via threshold optimization. The patient for S6 received treatment with first-generation EGFR-TKI creating the possibility that a rare event - a newly emerged drug-resistant mutation - was detected. Sampling bias resulting from limited ctDNA access may have also contributed to the negative results reported by the other platforms. These results suggest that positive T790M results should be followed-up with a confirmatory test both at the assay level as well as an additional blood draw at a later time point, prior to clinical action.
Moreover, S6 presented with multiple primary lung cancer and presented either the L858R (standard NGS) or exon 19 deletion (ADx-ARMS) in two different spatially separated tissues suggesting the spatial tumor heterogeneity [35, 36]. For S6's cfDNA analysis, ddPCR and cobas-ARMS only reported exon 19 deletion from the initial test, and Firefly NGS and ADx-ARMS reported L858R mutation. Accordingly, the plasma might contain L858R and exon 19 deletion double mutations. The L858R in plasma was reported to be positive in an additional ddPCR confirmation test, but the exon 19 deletion in plasma was not validated due to insufficient quantities of cfDNA. This discrepancy between the results of ddPCR and Firefly NGS on L858R and exon 19 deletion in the initial experiment may be the result of random Poisson sampling errors given the low abundance of ctDNA [37, 38].
In the final T790M discordance, ADx-ARMS found a T790M-positive case (S2) that was identified to be L858R-positive in tumor tissue and plasma using the other three platforms. Given the relative sensitivities of the platforms, this is likely a false-positive.
Finally, six patients with EGFR wild-type in plasma carried L858R (S10 - S12) or exon 19 deletion (S13 - S15) in their tissue. Of the six, four (S10 - S13) were responsive to treatment thus explaining the absence of mutations in plasma samples. The remaining two patients (S14, S15) were Stage IA treatment naïve, and the discrepancy between the initial tissue biopsy and plasma EGFR mutation profiles may have arisen from the limited availability of ctDNA in the blood stream of early stage patients (Supplementary Table S1).
Compared with the EGFR mutational profiles of tissue samples, ctDNA tests had a total coincidence rate of 45% - 65%, which was lower than that found in earlier studies (57% - 97%) [17, 24, 39]. One possible explanation can be found in the temporal tumor heterogeneity, such as sampling time interval variations between peripheral blood and tumor tissue. An additional explanation may be a change in EGFR mutation frequency previously observed in patients undergoing chemotherapy [40] or EGFR-TKI [41] therapy. 70% (14/20) patients included in this study were undergoing at least one of these two forms of treatment.
In addition to temporal heterogeneity, spatial heterogeneity within a tumor sample is a key factor in assessing the clinical relevance of ctDNA testing. The limitations associated with sampling heterogeneous tumors have been well-studied and are a major drawback of tissue biopsy-based mutation testing [36]. Studies have shown that primary and metastatic sites have varying genetic compositions [42], which is consistent with our findings. In patient S5, we identified an exon 19 deletion in primary lung tumor tissue and a G719X mutation in pleural lesions and blood samples post-metastasis. After three years of first generation EGFR-TKIs treatment, the disease progressed. Importantly, all three higher sensitivity technology platforms only detected the G719X mutation in the plasma at the time of disease progression. These results suggest that ctDNA testing offers more accurate mutational profiles after treatment, compared with the initial tissue biopsy, which has clear implications for clinical decision-making.
In addition, we evaluated whether mutational profiles in ctDNA were predictive of imminent relapse, as shown by other groups [41-43]. Tumor tissue and plasma samples from two patients (S7, S8) were observed. Tissue sample profiling detected only the exon 19 deletion, while exon 19/T790M double mutation was detected in plasma samples obtained approximately two years later. Both patients deteriorated significantly within five months post-sampling despite being diagnosed as having either stable disease or partial response. These results provide further support for the role of liquid biopsy testing in relapse prediction and clinical decision-making.
Another critical consideration in evaluating the comparative advantages of one technology over another is its regulatory status. Among the four platforms tested, only ADx-ARMS and cobas-ARMS are approved for EGFR mutation testing. Operationally, both methods have rapid turnaround times and simple workflows which require limited technical expertise. ADx-ARMS is the more affordable platform between the two though with a lower sensitivity. Another consideration is the testing panel's breadth. Cobas-ARMS and ddPCR also have fast turnaround times and high performance sensitivity, but neither is compatible with a comprehensive mutation assay which presents a challenge especially when the quantity of cfDNA sample is limited. At present, cobas-ARMS can only test for EGFR variants, and ddPCR can test one variant at a time and cannot assay EGFR mutations such as the exon 20 insertion.
In contrast, NGS-based techniques are both highly-sensitive and able to cover a breadth of mutations including all EGFR variants and other clinically-important genes such as ALK, ROS, RET, KRAS and BRAF [44]. Furthermore, NGS-based methods can be multiplexed and can be easily expanded to accommodate an increasing number of druggable mutations [45]. NGS-based methods, however, are more costly than ARMS and ddPCR methods with longer turnaround times. Additional studies are needed to investigate the clinical relevance of ultra-sensitive variant data and inform the cost-benefit analysis of platform selection.
There were several limitations to this study. First, parallel analysis of tumor tissue and plasma was not performed and most patients were enrolled while undergoing treatment. Second, our sample size was small and a larger follow-up study is needed to validate our findings. Third, only a small fraction of the assays was repeated due to limitations in cfDNA availability.
In conclusion, our study provides a comparative evaluation of four major technologies, ADx-ARMS, cobas-ARMS, ddPCR and Firefly NGS, for EGFR mutation detection in ctDNA from NSCLC patients. Despite the limited sample size, the cases are representative of the larger patient population and therefore can provide valuable insight into the role of ctDNA analysis in the treatment of cancer. This small data set illustrates some of the many advantages that liquid biopsy provides within the context of treatment evaluation and drug resistance detection. The cases that we reported here will contribute to the growing body of knowledge of the role of ctDNA detection in the treatment of cancer patients. Further study is needed to establish guidelines on how the detection of cancer related variants - such as EGFR mutational profiling - in ctDNA can be incorporated into existing clinical treatment paradigms.
Abbreviations
NSCLC, non-small cell lung cancer; ADC, adenocarcinoma; SCC, squamous cell carcinoma; ASC, adenosquamous carcinoma; EGFR, epidermal growth factor receptor; TKI, tyrosine kinase inhibitor; cfDNA, cell-free DNA; ctDNA, circulating tumor DNA; PCR, polymerase chain reaction; ARMS, amplification refractory mutation system; ddPCR, droplet digital polymerase chain reaction; NGS, next-generation sequencing; ADx, AmoyDx; CFDA, China Federal Drug Administration; FDA, U.S. Food and Drug Administration; TT, targeted therapy; CT, chemotherapy; PEM, pemetrexed; PTX, paclitaxel; DDP, cisplatin; GEM, gemcitabine; NDP, nedaplatin; TXT, docetaxel; CAP, carboplatin; RECIST, Response Evaluation Criteria in Solid Tumors; PR, partial response; SD, stable disease; PD, progressive disease; COSMIC, Catalogue of Somatic Mutations in Cancer.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
We would like to thank our patients and their families for their contributions to this research.
Financial support
This study was funded by National High Technology Research and Development Program of China (2015AA020403); Beijing Municipal Administration of Hospitals Clinical Medicine Development of Special Funding Support (ZYLX201509); Science Fund for Creative Research Groups of the National Natural Science Foundation of China (GrantIRT13003); Prior Sci-Tech Startup Project for Overseas Chinese Talents 2016 of the Beijing Bureau of Human Resource and Social Security.
Competing Interests
G. Q. Zhao and S. Lin are employees and stockholders of AccuraGen Inc. All other authors declare that there are no conflicts of interest.
References
1. Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW. et al. Activating Mutations in the Epidermal Growth Factor Receptor Underlying Responsiveness of Non-Small-Cell Lung Cancer to Gefitinib. N Engl J Med. 2004;350:2129-39
2. Paez JG, Janne PA, Lee JC, Tracy S, Greulich H, Gabriel S. et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science. 2004;304:1497-500
3. Mok TS, Wu YL, Thongprasert S, Yang CH, Chu DT, Saijo N. et al. Gefitinib or carboplatin-paclitaxel in pulmonary adenocarcinoma. N Engl J Med. 2009;361:947-57
4. Goto K, Ichinose Y, Ohe Y, Yamamoto N, Negoro S, Nishio K. et al. Epidermal growth factor receptor mutation status in circulating free DNA in serum: from IPASS, a phase III study of gefitinib or carboplatin/paclitaxel in non-small cell lung cancer. J Thorac Oncol. 2012;7:115-21
5. Sacher AG, Paweletz C, Dahlberg SE, Alden RS, O'Connell A, Feeney N. et al. Prospective Validation of Rapid Plasma Genotyping for the Detection of EGFR and KRAS Mutations in Advanced Lung Cancer. JAMA Oncol. 2016;2:1014-22
6. Que D, Xiao H, Zhao B, Zhang X, Wang Q, Xiao H. et al. EGFR mutation status in plasma and tumor tissues in non-small cell lung cancer serves as a predictor of response to EGFR-TKI treatment. Cancer Biol Ther. 2016;17:320-7
7. Zhu G, Ye X, Dong Z, Lu YC, Sun Y, Liu Y. et al. Highly Sensitive Droplet Digital PCR Method for Detection of EGFR-Activating Mutations in Plasma Cell-Free DNA from Patients with Advanced Non-Small Cell Lung Cancer. J Mol Diagn. 2015;17:265-72
8. Zhang X, Zhao Y, Wang M, Yap WS, Chang AY. Detection and comparison of epidermal growth factor receptor mutations in cells and fluid of malignant pleural effusion in non-small cell lung cancer. Lung Cancer. 2008;60:175-82
9. Hubers AJ, Heideman DA, Yatabe Y, Wood MD, Tull J, Taron M. et al. EGFR mutation analysis in sputum of lung cancer patients: a multitechnique study. Lung Cancer. 2013;82:38-43
10. Reckamp KL, Melnikova VO, Karlovich C, Sequist LV, Camidge DR, Wakelee H. et al. A Highly Sensitive and Quantitative Test Platform for Detection of NSCLC EGFR Mutations in Urine and Plasma. J Thorac Oncol. 2016;11:1690-700
11. Berz D, Raymond VM, Garst JH, Erlander MG. Non-invasive urine testing of EGFR activating mutation and T790M resistance mutation in non-small cell lung cancer. Exp Hematol Oncol. 2016;5:24-9
12. Sasaki S, Yoshioka Y, Ko R, Katsura Y, Namba Y, Shukuya T. et al. Diagnostic significance of cerebrospinal fluid EGFR mutation analysis for leptomeningeal metastasis in non-small-cell lung cancer patients harboring an active EGFR mutation following gefitinib therapy failure. Respir Investig. 2016;54:14-9
13. Kawahara A, Azuma K, Sumi A, Taira T, Nakashima K, Aikawa E. et al. Identification of non-small-cell lung cancer with activating EGFR mutations in malignant effusion and cerebrospinal fluid: rapid and sensitive detection of exon 19 deletion E746-A750 and exon 21 L858R mutation by immunocytochemistry. Lung Cancer. 2011;74:35-40
14. Yung TK, Chan KC, Mok TS, Tong J, To KF, Lo YM. Single-molecule detection of epidermal growth factor receptor mutations in plasma by microfluidics digital PCR in non-small cell lung cancer patients. Clin Cancer Res. 2009;15:2076-84
15. Diehl F, Schmidt K, Choti MA, Romans K, Goodman S, Li M. et al. Circulating mutant DNA to assess tumor dynamics. Nat Med. 2008;14:985-90
16. Holdhoff M, Schmidt K, Donehower R, Diaz LA Jr. Analysis of circulating tumor DNA to confirm somatic KRAS mutations. J Natl Cancer Inst. 2009;101:1284-5
17. Xu S, Lou F, Wu Y, Sun DQ, Zhang JB, Chen W. et al. Circulating tumor DNA identified by targeted sequencing in advanced-stage non-small cell lung cancer patients. Cancer Lett. 2016;370:324-31
18. Jamal-Hanjani M, Wilson GA, Horswell S, Mitter R, Sakarya O, Constantin T. et al. Detection of ubiquitous and heterogeneous mutations in cell-free DNA from patients with early-stage non-small-cell lung cancer. Ann Oncol. 2016;27:862-7
19. Huang WL, Wei F, Wong DT, Lin CC, Su WC. The Emergent Landscape of Detecting EGFR Mutations Using Circulating Tumor DNA in Lung Cancer. Biomed Res Int. 2015;2015:340732-41
20. Yang X, Zhuo M, Ye X, Bai H, Wang Z, Sun Y. et al. Quantification of mutant alleles in circulating tumor DNA can predict survival in lung cancer. Oncotarget. 2016;7:20810-24
21. Xu F, Wu J, Xue C, Zhao Y, Jiang W, Lin L. et al. Comparison of different methods for detecting epidermal growth factor receptor mutations in peripheral blood and tumor tissue of non-small cell lung cancer as a predictor of response to gefitinib. Onco Targets Ther. 2012;5:439-47
22. Taniguchi K, Uchida J, Nishino K, Kumagai T, Okuyama T, Okami J. et al. Quantitative detection of EGFR mutations in circulating tumor DNA derived from lung adenocarcinomas. Clin Cancer Res. 2011;17:7808-15
23. Forshew T, Murtaza M, Parkinson C, Gale D, Tsui DW, Kaper F. et al. Noninvasive identification and monitoring of cancer mutations by targeted deep sequencing of plasma DNA. Sci Transl Med. 2012;4:1-14
24. Thress KS, Brant R, Carr TH, Dearden S, Jenkins S, Brown H. et al. EGFR mutation detection in ctDNA from NSCLC patient plasma: A cross-platform comparison of leading technologies to support the clinical development of AZD9291. Lung Cancer. 2015;90:509-15
25. Suzuki S, Ono N, Furusawa C, Ying BW, Yomo T. Comparison of sequence reads obtained from three next-generation sequencing platforms. PLoS One. 2011;6:1-6
26. Kimura H, Kasahara K, Kawaishi M, Kunitoh H, Tamura T, Holloway B. et al. Detection of epidermal growth factor receptor mutations in serum as a predictor of the response to gefitinib in patients with non-small-cell lung cancer. Clin Cancer Res. 2006;12:3915-21
27. Watanabe M, Kawaguchi T, Isa S, Ando M, Tamiya A, Kubo A. et al. Ultra-Sensitive Detection of the Pretreatment EGFR T790M Mutation in Non-Small Cell Lung Cancer Patients with an EGFR-Activating Mutation Using Droplet Digital PCR. Clin Cancer Res. 2015;21:3552-60
28. Eisenhauer EA, Therasse P, Bogaerts J, Schwartz LH, Sargent D, Ford R. et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer. 2009;45:228-47
29. Newman AM, Bratman SV, To J, Wynne JF, Eclov NC, Modlin LA. et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med. 2014;20:548-54
30. Kobayashi S, Boggon TJ, Dayaram T, Janne PA, Kocher O, Meyerson M. et al. EGFR mutation and resistance of non-small-cell lung cancer to gefitinib. N Engl J Med. 2005;352:786-92
31. Dawson SJ, Tsui DW, Murtaza M, Biggs H, Rueda OM, Chin SF. et al. Analysis of circulating tumor DNA to monitor metastatic breast cancer. N Engl J Med. 2013;368:1199-209
32. Wu SG, Liu YN, Tsai MF, Chang YL, Yu CJ, Yang PC. et al. The mechanism of acquired resistance to irreversible EGFR tyrosine kinase inhibitor-afatinib in lung adenocarcinoma patients. Oncotarget. 2016;7:12404-13
33. Fujii A, Harada T, Iwama E, Ota K, Furuyama K, Ijichi K. et al. Hypermethylation of the CpG dinucleotide in epidermal growth factor receptor codon 790: implications for a mutational hotspot leading to the T790M mutation in non-small-cell lung cancer. Cancer Genet. 2015;208:271-8
34. Lin MT, Mosier SL, Thiess M, Beierl KF, Debeljak M, Tseng LH. et al. Clinical validation of KRAS, BRAF, and EGFR mutation detection using next-generation sequencing. Am J Clin Pathol. 2014;141:856-66
35. Ye C, Wang J, Li W, Chai Y. Novel Strategy for Synchronous Multiple Primary Lung Cancer Displaying Unique Molecular Profiles. Ann Thorac Surg. 2016;101:e45-7
36. Zhang J, Fujimoto J, Zhang J, Wedge DC, Song X, Zhang J. et al. Intratumor heterogeneity in localized lung adenocarcinomas delineated by multiregion sequencing. Science. 2014;346:256-9
37. Bosman KJ, Nijhuis M, van Ham PM, Wensing AM, Vervisch K, Vandekerckhove L. et al. Comparison of digital PCR platforms and semi-nested qPCR as a tool to determine the size of the HIV reservoir. Sci Rep. 2015;5:13811-9
38. Irwin PL, Nguyen LH, Chen CY. The relationship between purely stochastic sampling error and the number of technical replicates used to estimate concentration at an extreme dilution. Anal Bioanal Chem. 2010;398:895-903
39. Li X, Ren R, Ren S, Chen X, Cai W, Zhou F. et al. Peripheral blood for epidermal growth factor receptor mutation detection in non-small cell lung cancer patients. Transl Oncol. 2014;7:341-8
40. Bai H, Wang Z, Chen K, Zhao J, Lee JJ, Wang S. et al. Influence of chemotherapy on EGFR mutation status among patients with non-small-cell lung cancer. J Clin Oncol. 2012;30:3077-83
41. Oxnard GR, Paweletz CP, Kuang Y, Mach SL, O'Connell A, Messineo MM. et al. Noninvasive detection of response and resistance in EGFR-mutant lung cancer using quantitative next-generation genotyping of cell-free plasma DNA. Clin Cancer Res. 2014;20:1698-705
42. Kim EY, Cho EN, Park HS, Kim A, Hong JY, Lim S. et al. Genetic heterogeneity of actionable genes between primary and metastatic tumor in lung adenocarcinoma. BMC Cancer. 2016;16:27-35
43. Yanagita M, Redig AJ, Paweletz CP, Dahlberg SE, O'Connell A, Feeney N. et al. A prospective evaluation of circulating tumor cells and cell-free DNA in EGFR mutant non-small cell lung cancer patients treated with erlotinib on a phase II trial. Clin Cancer Res. 2016;22:6010-20
44. Chabon JJ, Simmons AD, Lovejoy AF, Esfahani MS, Newman AM, Haringsma HJ. et al. Circulating tumour DNA profiling reveals heterogeneity of EGFR inhibitor resistance mechanisms in lung cancer patients. Nat Commun. 2016;7:11815-28
45. Deeb KK, Hohman CM, Risch NF, Metzger DJ, Starostik P. Routine Clinical Mutation Profiling of Non-Small Cell Lung Cancer Using Next-Generation Sequencing. Arch Pathol Lab Med. 2015;139:913-21
Author contact
Corresponding authors: Ke-Neng Chen, chenkenengedu.cn; Guobing Xu, 18813186407edu.cn
 Global reach, higher impact
Global reach, higher impact