13.3
Impact Factor
Theranostics 2018; 8(8):2251-2263. doi:10.7150/thno.23877 This issue Cite
Research Paper
Cyclin D1-mediated microRNA expression signature predicts breast cancer outcome
1. Pennsylvania Cancer and Regenerative Medicine Research Center, Baruch S. Blumberg Institute, Pennsylvania Biotechnology Center and Lankenau Institute for Medical Research, 100 East Lancaster Avenue, Suite, 222, Wynnewood, PA. 19096
2. Research Center for Translational Medicine, East Hospital, Tongji University School of Medicine, Shanghai 200120, China
3. Department of Cancer Biology, Thomas Jefferson University, 233 South 10 th St. Philadelphia PA 19107
4. Center for Integrated Bioinformatics, Drexel University, Philadelphia, PA 19104
5. School of Biomedical Engineering, Systems and Health Sciences, Drexel University, Philadelphia, PA 19104
6. Shanghai Gongli Hospital, the Second Military Medical University, Shanghai 200120, China.
7. Lee Kong Chian School of Medicine, Nanyang Technological University, Singapore 637551, Singapore.
# Contributed equally to this paper.
Abstract
Background: Genetic classification of breast cancer based on the coding mRNA suggests the evolution of distinct subtypes. Whether the non-coding genome is altered concordantly with the coding genome and the mechanism by which the cell cycle directly controls the non-coding genome is poorly understood.
Methods: Herein, the miRNA signature maintained by endogenous cyclin D1 in human breast cancer cells was defined. In order to determine the clinical significance of the cyclin D1-mediated miRNA signature, we defined a miRNA expression superset from 459 breast cancer samples. We compared the coding and non-coding genome of breast cancer subtypes.
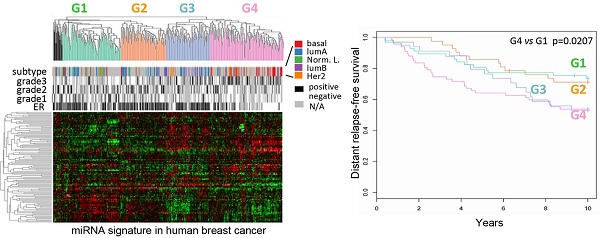
Results: Hierarchical clustering of human breast cancers defined four distinct miRNA clusters (G1-G4) associated with distinguishable relapse-free survival by Kaplan-Meier analysis. The cyclin D1-regulated miRNA signature included several oncomirs, was conserved in multiple breast cancer cell lines, was associated with the G2 tumor miRNA cluster, ERα+ status, better outcome and activation of the Wnt pathway. The coding and non-coding genome were discordant within breast cancer subtypes. Seed elements for cyclin D1-regulated miRNA were identified in 63 genes of the Wnt signaling pathway including DKK. Cyclin D1 restrained DKK1 via the 3'UTR. In vivo studies using inducible transgenics confirmed cyclin D1 induces Wnt-dependent gene expression.
Conclusion: The non-coding genome defines breast cancer subtypes that are discordant with their coding genome subtype suggesting distinct evolutionary drivers within the tumors. Cyclin D1 orchestrates expression of a miRNA signature that induces Wnt/β-catenin signaling, therefore cyclin D1 serves both upstream and downstream of Wnt/β-catenin signaling.
Keywords: cyclin D1, miRNA, breast cancer
 Global reach, higher impact
Global reach, higher impact