13.3
Impact Factor
Theranostics 2019; 9(13):3723-3731. doi:10.7150/thno.33980 This issue Cite
Research Paper
One-step colorimetric genotyping of single nucleotide polymorphism using probe-enhanced loop-mediated isothermal amplification (PE-LAMP)
1. Natural Products Research Center, Chengdu Institute of Biology, Chinese Academy of Sciences, Chengdu 610041, P. R. China
2. University of Chinese Academy of Sciences, Beijing 100049, P. R. China
3. Chengdu University of Traditional Chinese Medicine, Chengdu 611137, P. R. China.
Abstract
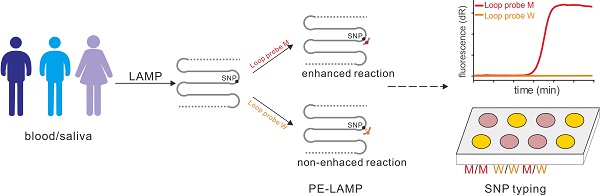
Single nucleotide polymorphism (SNP) is the most abundant molecular marker associated with many physiologic and pathologic phenotypes. An isothermal, accurate and cost-effective SNP detection could make a great difference in point-of-care testing (POCT) or on-site diagnosis. However, there are two challenges, the expensive instrument and labor-intensive process, faced by the development of on-site SNP detection. We reported a novel SNP typing method based on the probe-enhanced loop-mediated isothermal amplification (PE-LAMP), which combines the oligonucleotide probe with a conventional LAMP to realize the SNP discrimination by analyzing the great discrepancy in amplification efficiency.
Methods: We firstly constructed the genotyping method by combining the hybridization of the specific probe with the powerful amplification of LAMP. Then we validated the method by genotyping the SNP rs3741219 and we sought to realize one-step visualized typing. Finally, we applied the method to pharmacogenomic testing by genotyping CYP2C19*2 and MDR1 C3435T.
Results: The PE-LAMP was successfully constructed to detect SNP and the sensitivity of our method is as low as 1000 copies of target DNA, which is sufficient to routine diagnosis. The high specificity in detecting mutant in the presence of excess wild-type allele could be achieved. It has shown good performance in helping predict the individual response of antiplatelet drug Clopidogrel through typing simply treated saliva samples.
Conclusions: The proposed method is one-step, colorimetric, specific and sensitive enough to detect crudely treated samples, showing great potential in the pharmacogenomic study and POCT use.
Keywords: SNP, LAMP, colorimetric detection, pharmacogenomic
 Global reach, higher impact
Global reach, higher impact