Impact Factor
Theranostics 2011; 1:371-380. doi:10.7150/thno/v01p0371 This volume Cite
Research Paper
Trackable and Targeted Phage as Positron Emission Tomography (PET) Agent for Cancer Imaging
1. Molecular Imaging Center, Department of Radiology, University of Southern California, Los Angeles 90033, USA.
2. Biosciences Division, Argonne National Laboratory, 9700 S. Cass Avenue, , Argonne, IL 60439, USA.
# Jin and Huang contributed equally to the research.
Received 2011-10-22; Accepted 2011-11-9; Published 2011-11-18
Abstract
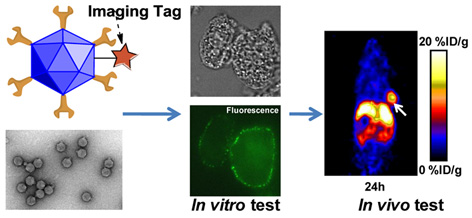
The recent advancement of nanotechnology has provided unprecedented opportunities for the development of nanoparticle enabled technologies for detecting and treating cancer. Here, we reported the construction of a PET trackable organic nanoplatform based on phage particle for targeted tumor imaging. Method: The integrin αvβ3 targeted phage nanoparticle was constructed by expressing RGD peptides on its surface. The target binding affinity of this engineered phage particle was evaluated in vitro. A bifunctional chelator (BFC) 1,4,7,10-tetraazadodecane-N,N',N",N"'-tetraacetic acid (DOTA) or 4-((8-amino-3,6,10,13,16,19-hexaazabicyclo [6.6.6] icosane-1-ylamino) methyl) benzoic acid (AmBaSar) was then conjugated to the phage surface for 64Cu2+ chelation. After 64Cu radiolabeling, microPET imaging was performed in U87MG tumor model and the receptor specificity was confirmed by blocking experiments. Results: The phage-RGD demonstrated target specificity based on ELISA experiment. According to the TEM images, the morphology of the phage was unchanged after the modification with BFCs. The labeling yield was 25 ± 4% for 64Cu-DOTA-phage-RGD and 46 ± 5% for 64Cu-AmBaSar-phage-RGD, respectively. At 1 h time point, 64Cu-DOTA-phage-RGD and 64Cu-AmBaSar-phage-RGD have comparable tumor uptake (~ 8%ID/g). However, 64Cu-AmBaSar-phage-RGD showed significantly higher tumor uptake (13.2 ± 1.5 %ID/g, P<0.05) at late time points compared with 64Cu-DOTA-phage-RGD (10 ± 1.2 %ID/g). 64Cu-AmBaSar-phage-RGD also demonstrated significantly lower liver uptake, which could be attributed to the stability difference between these chelators. There is no significant difference between two tracers regarding the uptake in kidney and muscle at all time points tested. In order to confirm the receptor specificity, blocking experiment was performed. In the RGD blocking experiment, the cold RGD peptide was injected 2 min before the administration of 64Cu-AmBaSar-phage-RGD. Tumor uptake was partially blocked at 1 h time point. Phage-RGD particle was also used as the competitive ligand. In this case, the tumor uptake was significantly reduced and the value was kept at low level consistently. Conclusion: In this report, we constructed a PET trackable nanoplatform based on phage particle and demonstrated the imaging capability of these targeted agents. We also demonstrated that the choice of chelator could have significant impact on imaging results of nano-agents. The method established in this research may be applicable to other receptor/ligand systems for theranostic agent construction, which could have an immediate and profound impact on the field of imaging/therapy and lay the foundation for the construction of next generation cancer specific theranostic agents.
Keywords: phage particle, positron emission tomography, integrin αvβ3, RGD, Cu-64.
INTRODUCTION
During the last decade, tremendous progress has been made in the development of new molecular imaging probes and therapeutic agents targeting cancer [1-4]. However, cancer still remains a major fatal disease around the world. There is clearly a need to develop innovative diagnostic and therapeutic methods beyond tradition and convention. Recently, the advancement of nanotechnology has provided unprecedented opportunities for the development of nanoparticle enabled technologies for detecting and treating cancer [5-8]. For example, surface functionalized organic/inorganic nanoparticles hold the great promises for the eradication of cancer by creating the theranostic systems (therapy + multifunctional diagnosis) that enable i) early detection of the disease, ii) monitoring therapeutic response, and iii) targeted delivery of therapeutic agents, based on the “whole in one approach” [9, 10]. However, applications of these nanotechniques were limited by the potential toxicity from inorganic core and the lack of well controlled surface modification method [11, 12]. The development of well defined and biodegradable delivery systems is therefore preferred.
As shown in Figure 1, phage particles are unique platforms for imaging probes or drug carriers in that 1) they could be genetically modified to display target specific ligands; 2) they can be economically and effectively produced with absolute uniformity controlled by nature; 3) they can be covalently attached to radiometal chelators while simultaneously expressing multiple copies of cancer targeting peptides; 4) they are physically well-characterized, resistant to harsh conditions, biocompatible and nonpathogenic (compared with virus based particles); 5) the genetic material inside the capsid can be removed without affecting the integrity of capsid [13]. Such empty container is ideal for encapsulating other therapeutic reagents which can be released after reaching specific cellular sites. These factors suggest that functionalized bacteriophage particles hold great potential as novel advanced imaging agents and targeting systems for drug and DNA delivery, and will likely complement existing organic/inorganic nanoplatforms [14-17]. Although target specific radiolabeled phage particles obtained from either high throughput screen (phage display) or rational design (the expression of established affinity reagents on the phage surface) have been reported in molecular imaging of variety of diseases, most of the imaging results are still suboptimal [18, 19]. Both two-step and three-step pretargeted strategies have been explored for SPECT imaging of cancer [19, 20]. However, its application in theranostic approach is limited in some extend due to the complicated procedures. In this report, we developed a set of phage particle-based novel imaging agents with great therapeutic potential due to their high carrying capacity and superior targeted delivery ability [21-23].
(A) Rationale for developing PET trackable phage nanoparticles. (B) The synthetic scheme of DOTA-/AmBaSar-T7-RGD.
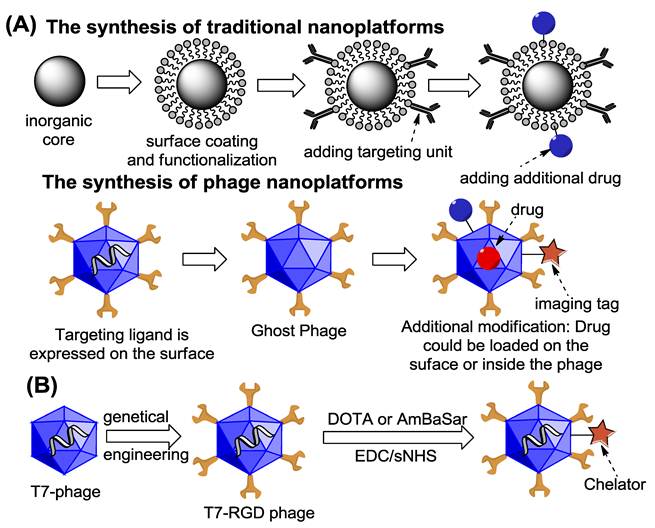
Positron emission tomography (PET) is a non-invasive functional imaging technique with good resolution, high sensitivity, and accurate quantification [24]. An important advantage of PET is that it provides quantitative information of physiological, biochemical and pharmacological processes in living subjects. The vitronectin receptor integrin αvβ3 has been the focus of intensive research because of its major role in several distinct processes, particularly osteoclast mediated bone resorption, angiogenesis and pathological neovascularization, and tumor metastasis [25, 26]. A variety of RGD peptide and antibody based probes have been reported for multimodality imaging of integrin αvβ3 expression in vivo [27-30]. In our approach, we developed integrin αvβ3 targeted phage particle for PET imaging.
MATERIALS AND METHODS
All commercially available reagents were used without further purification. DOTA was purchased from Macrocyclics, Inc. AmBaSar was prepared in house as reported in reference [31]. PD-10 column was purchased from GE Healthcare. 1-ethyl-3-[3-(dimethylamino)propyl]carbodiimide (EDC), N-hydroxysulfonosuccinimide (SNHS), and Chelex 100 resin (50 - 100 mesh) were purchased from Aldrich. Water and all buffers were passed through a Chelex 100 column (1 × 15 cm) before radiolabeling. 64Cu was purchased from University of Wisconsin-Madison or obtained from Shawn Chen's group at NIBIB/NIH.
Production of T7-RGD
To construct T7 bacteriophages with RGD-containing peptide displayed on the surface, two primers, 5'- aattctggcggcggcagctgcCGTGGTGATTTTAAAtgcTAA-3' and 5'- AGCTTTAgcaTTTAAAATCACCACGgcagctgccgccgccag-3' were synthesized, self-annealed and inserted into T7select415-1b vector previously digested by EcoRI and Hind III following the instructions provided by manufacturer (EMD4Biosciences, T7Select® 415-1 Cloning Kit, Gibbstown, NJ 08027) . The ligated products were then subjected for in vitro packaging according to the protocol provided by manufacturer. The resulting recombinant phages were identified by PCR screen and named T7-RGD. The inserted DNA sequence was confirmed by DNA sequencing and led to the expression of the following peptide NSGGGSCRGDFKC where RGD sequence was constrained by two cysteins and led by GGGS flexible linker.
Preparation of Cy5.5-Phage-RGD
The preparation of Cy5.5 conjugated Phage-RGD was achieved using a water-soluble procedure. In brief, 1 mg of Cy5.5-NHS (1.5 μmol) was dissolved in 100 μl of DMSO and saved as stock solution. The Cy5.5-NHS and Phage-RGD was added at 1:5 ratio and the reaction mixture was incubated overnight at pH 8.5. The Cy5.5-Phage-RGD was purified by passing through a PD-10 column. The eluate was washed with water three times using centricon filter (10K cut-off). The resulting solution was aliquoted for future use.
Chemistry and radiochemistry
DOTA and AmBaSar were activated and conjugated to the phage particle using a water-soluble procedure reported earlier [32]. For example: 0.75 mg of AmBaSar (1.5 μmol) dissolved in 20 μL of water and 0.29 mg of EDC (1.5 μmol) dissolved in 10 μL of water were mixed at pH 5.0. SNHS (0.26 mg, 1.2 μmol) in 10 μL of water was then added to the stirring mixture on ice-bath, and the pH was adjusted to 5.5. The reaction was incubated for 30 min at 4 °C. The Phage-RGD was then added to the AmBaSar-OSSu reaction mixture at pH 8.5. The reaction was incubated at 4 °C for overnight. The AmBaSar-Phage-RGD was purified by passing through a PD-10 column. The eluate was washed with water three times using centricon filter. The resulting solution was aliquoted for future use.
The 64Cu-DOTA-Phage-RGD or 64Cu-AmBaSar-Phage-RGD was prepared using a method developed in our laboratory with minor modifications[32]: [64Cu]Acetate (64Cu(OAc)2) was prepared by adding 37-111 MBq of 64CuCl2 into pH 5.0~5.5 ammonium acetate buffer (0.1 N, 300 µL). The DOTA-phage-RGD or AmBaSar-Phage-RGD was then added to the above 64Cu(OAc)2 solution, and the resulting mixture was incubated at 23~37 °C for 30 min, followed by purification with PD-10 column. The radioactive peak containing 64Cu-DOTA-Phage-RGD or 64Cu-AmBaSar-Phage-RGD was collected for the future use.
In vitro assay
Purified T7 and T7-RGD phages were fluorescently labeled and used for the affinity detection with integrin αvß3 in a standard ELISA assay. The wells were first coated with different concentration of αvß3, and then washed, blocked with 5% milk and coated with different amount of Alexa Fluor-488 (Invitrogen) labeled T7-RGD phage or wild type T7 phage. After washed, the fluorescence intensity was measured under appropriate excitation and emission setting. To test whether the RGD displayed on T7 phage surface can interact or mediate the internalization of phage through the interaction with integrin expressed on cancer cell surfaces, both prefixed (cells were fixed first then incubated with phage) and living ovarian cancer cells (cells were incubated with phage first then detached and fixed) were used. When SKOV3 or U87MG (high expression of integrin αvβ3)[33-35] grew to 70% confluent, they were detached by treatment with trypsin and fixed by 4% glutaraldehyde, then incubated with labeled T7 or T7-RGD phage. Alternatively, labeled phages were directly added to the attached living SKOV3 or U87MG cells without any fixation. After incubation for 1 h, cells were dissociated, washed, fixed, and observed under microscope.
Animal Models
Athymic nude mice (about 10-20 weeks old, with a body weight of 20-25 g) were obtained from Harlan (Charles River, MA). All animal experiments were performed according to a protocol approved by University of Southern California Institutional Animal Care and Use Committee. The U87MG human glioma xenograft model was generated by subcutaneous injection of 5 × 106 U87MG human glioma cells into the front flank of athymic nude mice. The tumors were allowed to grow 3-5 weeks until 200-500 mm3 in volume. Tumor growth was followed by caliper measurements of the perpendicular dimensions.
In Vivo Near-infrared optical imaging of tumors
In vivo fluorescence imaging was performed with an IVIS 200 small animal imaging system (Xenogen, Alameda, CA). The system was equipped with 21 emmission filter sets that can be used to image optical signals from green to near-infrared. A Cy5.5 filter set was used for acquiring the fluorescence of Cy5.5-Phage-RGD in vivo. Identical illumination settings (lamp voltage, filters, f/stop, field of views, binning) were used for acquiring all images. Fluorescence emission was normalized to photons per second per centimeter squared per steradian (p/s/cm2/sr). Images were analyzed using Living Image 4.0 software (Xenogen) to allow the spectral unmixing, reduce the tissue autofluorescence and crosstalk. The experimental mice (n=3) were injected with Cy5.5 conjugated RGD-phages via the tail vein, anesthetized with 2% isoflurane, and followed with static scans. All near-infrared fluorescence images were acquired using a 1-second exposure time (f/stop= 4). For each imaging scan, regions of interest (ROIs) were drawn over each tumor xenograft site to semi-quantify the signals. The mice were euthanized at 72 hours post-injection. The tumor and major organs were dissected, and ex vivo fluorescence images were acquired to confirm the in vivo results.
microPET Studies
Micro-PET scans were performed using a rodent scanner (microPET R4; Siemens Medical Solutions) as previously reported [36]. About 11.1 MBq of 64Cu-DOTA-Phage-RGD or 64Cu-AmBaSar-Phage-RGD was intravenously injected into each mouse under isoflurane anesthesia. Ten minute static scans were acquired at desired time points. The images were reconstructed by a 2-dimensional ordered-subsets expectation maximum (OSEM) algorithm. For each microPET scan, regions of interest were drawn over the tumor, normal tissue, and major organs on the decay-corrected whole-body coronal images. The radioactivity concentration (accumulation) within the tumor, muscle, liver, and kidneys were obtained from the mean value within the multiple regions of interest and then converted to %ID/g. For the RGD blocking experiment, mice bearing U87MG tumors were scanned (10 min static) at desired time points after the coinjection of 11.1 MBq 64Cu-AmBaSar-Phage-RGD with 30 mg/kg RGD per mouse. For phage-RGD blocking, 100 times phage-RGD was injected 3 min before the administration of 11.1 MBq 64Cu-AmBaSar-Phage-RGD.
Statistical Analysis
Quantitative data were expressed as mean ± SD. Means were compared using One-way ANOVA and student's t-test. P values < 0.05 were considered statistically significant.
RESULTS
Chemistry and Radiochemistry
Wild type T7 bacteriophages were purified from clarified lysates by precipitation with 10% PEG 8000 followed by banding in a CsCl step gradient. However, the yield of T7-RGD was found too low by using the same procedure. Majority of T7-RGD particles became insoluble. By lowering the concentration of PEG from 10% to 3% with the addition of 2-mecapataethanol, the insolubility of T7-RGD was greatly diminished and could be readily purified to a concentration as high as 1013 pfu/ml. After the conjugation to chelators, the labeling yield was 25±5% for DOTA-Phage-RGD and 48±6% for AmBaSar-Phage-RGD.
Cell Adhesion assay
The interaction between T7-RGD and αvβ3 integrin is first demonstrated by ELISA experiment. As shown in Figure 2A, compared to unmodified wild type T7 phage, T7-RGD phage is able to bind to the coated integrin in a dose dependent manner. When fibronectin, a demonstrated binding ligand of integrin via RGD peptide, was introduced, the interaction between integrin and T7-RGD phage is largely inhibited (data not shown), indicating the interaction between the phage and integrin is indeed mediated by RGD peptide displayed on the phage surface. To test whether the RGD displayed on T7-RGD phage surface can interact or mediate the internalization of phage through the binding with integrin expressed on cancer cell surfaces, both prefixed and living SKOV3 cells were used.
(A) Binding between T7-RGD phage and integrin αvβ3. (B) Confocal microscopic images of fixed SKOV3 ovarian cancer cells labeled with 488 conjugated wild type T7 (left, the exposure of control sample was further extended to study the weak binding with wild type phage) and T7-RGD (right) phage. (C) TEM images of DOTA and AmBaSar cage modified phage particles.
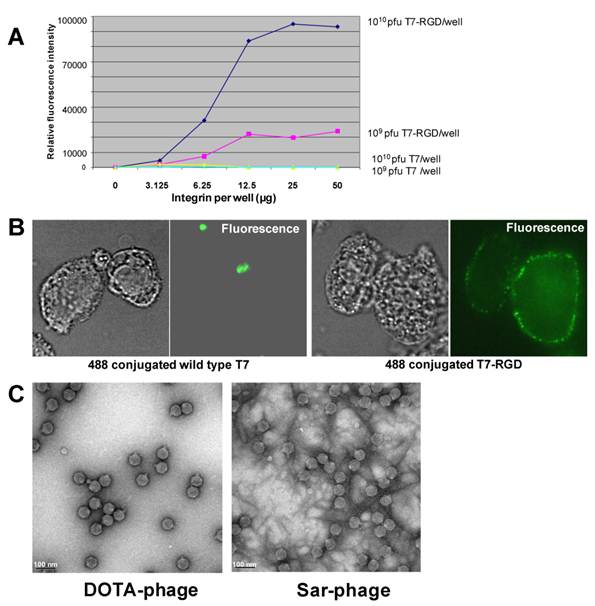
As shown in Figure 2B, when cells were fixed, the T7-RGD phage was found only binding to cell membrane where integrins were located. In contrast, almost no fixed cells were labeled by wild-type T7 phage. When living cells were used, about 80% cells were either weakly or strongly labeled by T7-RGD, while only 10-20% SKOV3 cells were labeled by T7. Furthermore, the localization of T7-RGD phage seemed to be both cytoplasm and membrane-bound, whereas T7 was only found intracellular (Supplementary Material: Figure S1). The U87MG cell behaves similarly and the T7-RGD demonstrated ~ 4 fold higher uptake compared with wild type T7 phage in living cells. These results clearly demonstrated that the RGD displayed on T7-RGD phage surface could mediate the internalization of phage through the interaction with integrin expressed on cancer cell surfaces.
In vivo Imaging of U87MG Tumor-Bearing Mice
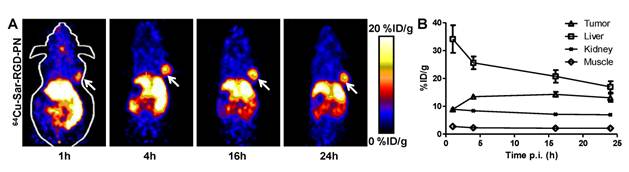
As shown in Figure 2C, the morphology of the phage was unchanged after the modification based on the TEM images. After the phage was radiolabeled with 64Cu (25±5% labeling yield), whole body imaging experiments were performed (Figure 3). The U87MG tumor (integrin αvβ3 positive) could be clearly visualized in microPET images and the tumor uptake reached a plateau after 4 h post injection (~9%ID/g). When Cy5.5-Phage-RGD was imaging using optical machine, much prominent tumor uptake and lower liver uptake was obtained (Fig. 3C). In addition to DOTA-phage-RGD particle, we introduced the sarcophagine (Sar) cage to the surface of the phage particle according to our previous procedure. Based on the TEM images, the morphology of phage was not changed after the modification with Sar cage (Figure 2C). After 64Cu radiolabeling (45±7% labeling yield), microPET imaging was performed in U87MG tumor model. At 1 h time point, 64Cu-DOTA-phage-RGD and 64Cu-AmBaSar-Phage-RGD have comparable tumor uptake (Figure 4, 7.9±2.3%ID/g and 8.8±0.7%ID/g, respectively). However, 64Cu-AmBaSar-Phage-RGD showed high tumor uptake at late time points (13.4±0.5, 14.3±0.9, and 13.2±1.5%ID/g, at 4, 16, and 24 h, respectively). 64Cu-AmBaSar-Phage-RGD also demonstrated significantly lower liver uptake (51.2±4.1, 44.9±3.4, and 32.3±2.6%ID/g for 64Cu-DOTA-phage-RGD; compared with 34.1±4.9, 25.6±2.3, and 20.7±2.2%ID/g 64Cu-AmBaSar-Phage-RGD at 1, 4, and 16 h, respectively), which could be attributed to the stability difference between two chelators. There is no significant difference between two tracers regarding the uptake in kidney and muscle at all time points tested (Supplementary Material: Figure S2).
(A) Coronal microPET images of nude mouse bearing U87MG tumor at 1 h, 4 h, and 16 h p.i. of 64Cu-DOTA-Phage-RGD. (B) Time activity curve derived from multi-time point microPET study (n=3). (C) Optical images of U87MG tumor at 4, 24, 48, 72 h post-injection. The ex vivo experiments were performed at 72 h post-injection.
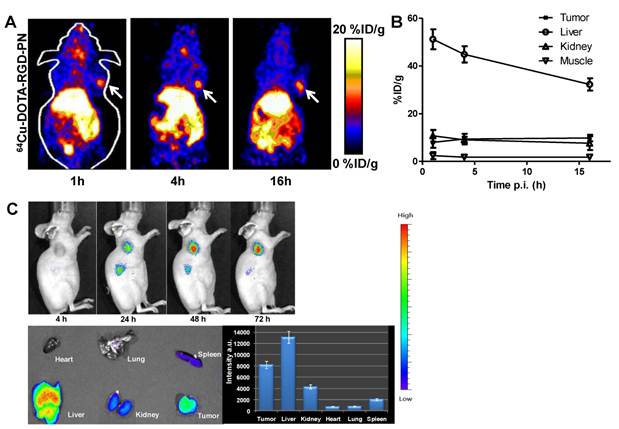
Coronal microPET images of nude mouse bearing U87MG tumor at 1 h, 4 h, 16 h, and 24 h p.i. of 64Cu-AmBaSar-Phage-RGD. (B) Time activity curve derived from multi-time point microPET study (n=3).
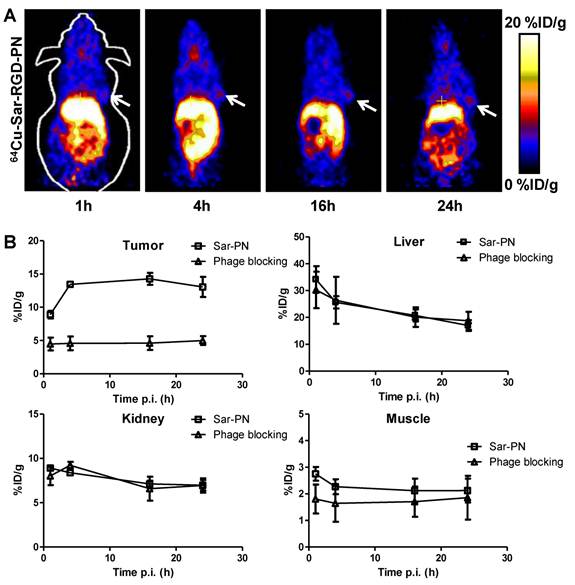
In order to confirm the receptor specificity, blocking experiment was performed. In the RGD blocking experiment, cold RGD peptide was injected 2 min before the administration of 64Cu-AmBaSar-T7-RGD. Tumor uptake was partially blocked at 1h time point and the blocking became less effective at late time points (Supplementary Material: Figure S3). The tracer uptake in other major organs has no significant difference between blocked and unblocked mice. We also performed the blocking experiment by using phage particle as the competitive ligand (Figure 5). In this case, the tumor uptake was significantly reduced and the value was kept at low level consistently. The more effective blocking may be attributed to the matched half-life between phage and 64Cu-AmBaSar-phage. Nonetheless, a more systematic study would be needed before a conclusion is drawn. The tracer uptake in other major organs has no significant difference between the blocked and unblocked mice.
(A) Coronal microPET images of U87MG tumor bearing nude mice after injection of 64Cu-AmBaSar-Phage-RGD with a blocking dose Phage-RGD. (B) The unlocked/ blocked tumor and major organ uptake derived from microPET study (n=3).
DISCUSSION
Phage display technology is emerging as a powerful approach to develop novel peptide ligands for a specific molecular target [13]. However, the screening process could be time consuming and the success is not guaranteed. Although target specific radiolabeled phage particles obtained from either high throughput screen (phage display) or rational design (the expression of established affinity reagents on the phage surface) have been reported in molecular imaging of variety of diseases, most of the imaging results are still suboptimal [18, 19]. In this report, we demonstrated that the phage based organic particle could be used as an imaging agent.
By using standard molecular cloning technique, a DNA fragment which encodes CRGDFKC peptide sequence was introduced into T7 genome. The resulting recombinant T7 phage (T7-RGD) enabled the expression and display of 415 copies of RGD-containing peptides on phage surface. The introduction of two cysteins at both ends might facilitate the formation of cyclic structure, which could possibly lead to stronger binding to integrins. The T7-RGD particles can be readily purified under reducing environment to a concentration as high as 1 × 1013 pfu/ml.
After confirming the receptor binding in vitro, we then evaluated the T7-RGD phage as PET agent for cancer imaging. Copper-64 (t1/2 = 12.7 h) decays by β+ (20%) and β- emission (37%), as well as electron capture (43%), making it well suited for radiolabeling proteins, antibodies and peptides, both for PET imaging (β+) and therapy (β+ and β-) [24, 37]. In vivo stable attachment of 64Cu2+ to targeted biomolecules generally requires the use of a bifunctional chelator (BFC). In order to label our integrin targeted phage particle, 1,4,7,10-tetraazadodecane-N,N',N",N"'-tetraacetic acid (DOTA) was conjugated to the particle. Although we have obtained promising imaging result for DOTA-phage-RGD particle, the liver uptake of this tracer was rather high with ~50%ID/g at 1h and ~30%ID/g at 16h. It is well known that the DOTA, TETA, and their derivatives, for 64Cu2+ labeling have limited stability in vivo due to the dissociation of 64Cu2+ from these BFCs, leading to high retention in liver. Recently, we have developed a carboxylate functionalized sarcophagine chelator AmBaSar for 64Cu2+ radiolabeling, which shows improved pharmacokinetics and dynamics due to the increased stability [32, 38, 39]. We then introduced the sarcophagine cage to the surface of the phage particle according to our previous procedure. 64Cu-AmBaSar-Phage-RGD showed high tumor uptake at late time points. 64Cu-AmBaSar-Phage-RGD also demonstrated significantly lower liver uptake, which could be attributed to the stability difference between two chelators. This information is important for nanoparticle based imaging studies. It clearly demonstrated that the choice of chelator could have significant impact on imaging results. As the imaging tags were an insignificant component compared to the whole T7 phage, the distribution pattern of T7-RGD phage should be mainly determined by the particle itself. However, the 64Cu-DOTA-Phage-RGD demonstrated more than 50% higher liver uptake compared with 64Cu-AmBaSar-Phage-RGD. The difference in these nanoparticle based PET agents could be mainly caused by the release of free Cu when an unstable chelator was selected. In fact, when Cy5.5 was tagged to the Phage-RGD particle (Fig. 3C), the particle distribution pattern correlates well with 64Cu-AmBaSar-Phage-RGD. Therefore, when high liver uptake was observed for 64Cu labeled nanoparticle agents, it could not only be caused by the reticuloendothelial clearance, but also by the free 64Cu fell off from the particle.
Unlike peptide based imaging agent, the retention time of phage particle was shown to be high and persistent in tumors. Due to its large size (~50 nm), high liver uptake was observed at early time point, and the value keep decreasing at late time points. These characters make the phage particles more suitable to be used as theranostic agent instead of pure imaging agent. Although these newly developed phage agents were only labeled with 64Cu for PET imaging in this application, the same construct may also be labeled with therapeutics isotopes (such as 67Cu/90Y) for targeted radio-immunotherapy of cancer. Moreover, for future phage based therapy applications (including radio, chemo and/or gene therapy), optimization of this newly developed phage platform could be performed through molecular imaging (using PET to select the best carrier with optimal tumor up-take and in vivo distribution) before the lengthy and costly therapy experiments. In summary, a PET trackable imaging platform was developed based on phage particle, which are physically well characterized, trackable after surface modification, and dictate its own target specific production.
CONCLUSION
While theranostic approach has become an attractive strategy for in vivo cancer imaging and therapy studies, there is a significant shortage of established platforms/methods for biodegradable theranostic agent construction. In this report, we constructed a PET trackable nanoplatform based on phage particle and demonstrated the imaging capability of these targeted agents. We also demonstrated that the choice of chelator could have significant impact on imaging results. Moreover, these newly developed agents hold great therapeutic potential due to their high loading capacity and superior targeted delivery ability, which could turn them into novel theranostic agents. The method established in this research should be applicable to other receptor/ligand systems for theranostic agent construction, which could have an immediate and profound impact on the field of imaging/therapy and lay the foundation for the construction of next generation cancer specific theranostic agents.
SUPPLEMENTARY MATERIAL
Figs. S1 - S3.
Acknowledgements
This work was supported by the USC Department of Radiology, the Department of Energy (DE-SC0002353), the National Cancer Institute (P30A014089), and the USC Biomedical Imaging Science Initiative.
Conflict of Interest
The authors have declared that no conflict of interest exists.
References
1. Mitsunaga M, Ogawa M, Kosaka N, Rosenblum LT, Choyke PL, Kobayashi H. Cancer cell-selective in vivo near infrared photoimmunotherapy targeting specific membrane molecules. Nat Med. 2011 [Epub ahead of print]
2. Conde J, Doria G, Baptista P. Noble metal nanoparticles applications in cancer. J Drug Deliv. 2012 [Epub ahead of print]
3. Luo S, Zhang E, Su Y, Cheng T, Shi C. A review of NIR dyes in cancer targeting and imaging. Biomaterials. 2011;32:7127-38
4. Liu TW, Chen J, Burgess L. et al. Multimodal bacteriochlorophyll theranostic agent. Theranostics. 2011;1:354-62
5. Huang P, Xu C, Lin J. et al. Folic Acid-conjugated Graphene Oxide loaded with Photosensitizers for Targeting Photodynamic Therapy. Theranostics. 2011;1:240-50
6. Allegra A, Penna G, Alonci A, Rizzo V, Russo S, Musolino C. Nanoparticles in Oncology: The New Theragnostic Molecules. Anticancer Agents Med Chem. 2011 [Epub ahead of print]
7. Shubayev VI, Pisanic TR 2nd, Jin S. Magnetic nanoparticles for theragnostics. Adv Drug Deliv Rev. 2009;61:467-77
8. Ye Y, Chen X. Integrin targeting for tumor optical imaging. Theranostics. 2011;1:102-26
9. Xie J, Lee S, Chen X. Nanoparticle-based theranostic agents. Adv Drug Deliv Rev. 2010;62:1064-79
10. Jeong H, Huh M, Lee SJ. et al. Photosensitizer-conjugated human serum albumin nanoparticles for effective photodynamic therapy. Theranostics. 2011;1:230-9
11. Johnston HJ, Hutchison G, Christensen FM, Peters S, Hankin S, Stone V. A review of the in vivo and in vitro toxicity of silver and gold particulates: particle attributes and biological mechanisms responsible for the observed toxicity. Crit Rev Toxicol. 2010;40:328-46
12. Sayes CM, Warheit DB. Characterization of nanomaterials for toxicity assessment. Wiley Interdiscip Rev Nanomed Nanobiotechnol. 2009;1:660-70
13. Deutscher SL. Phage display in molecular imaging and diagnosis of cancer. Chem Rev. 2010;110:3196-211
14. Clark JR, Bartley K, Jepson CD, Craik V, March JB. Comparison of a bacteriophage-delivered DNA vaccine and a commercially available recombinant protein vaccine against hepatitis B. FEMS Immunol Med Microbiol. 2011;61:197-204
15. Huang RK, Steinmetz NF, Fu CY, Manchester M, Johnson JE. Transferrin-mediated targeting of bacteriophage HK97 nanoparticles into tumor cells. Nanomedicine (Lond). 2011;6:55-68
16. Godin B, Tasciotti E, Liu X, Serda RE, Ferrari M. Multistage nanovectors: from concept to novel imaging contrast agents and therapeutics. Acc Chem Res. 2011;44:979-89
17. Yacoby I, Bar H, Benhar I. Targeted drug-carrying bacteriophages as antibacterial nanomedicines. Antimicrob Agents Chemother. 2007;51:2156-63
18. Rusckowski M, Gupta S, Liu G, Dou S, Hnatowich DJ. Investigation of four 99mTc-labeled bacteriophages for infection-specific imaging. Nucl Med Biol. May. 2008;35:433-40
19. Newton JR, Miao Y, Deutscher SL, Quinn TP. Melanoma imaging with pretargeted bivalent bacteriophage. J Nucl Med. 2007;48:429-36
20. Newton-Northup JR, Figueroa SD, Quinn TP, Deutscher SL. Bifunctional phage-based pretargeted imaging of human prostate carcinoma. Nucl Med Biol. 2009;36:789-800
21. Petrenko V. Evolution of phage display: from bioactive peptides to bioselective nanomaterials. Expert Opin Drug Deliv. 2008;5:825-36
22. Rodriguez-Carmona E, Villaverde A. Nanostructured bacterial materials for innovative medicines. Trends Microbiol. 2010;18:423-30
23. Yacoby I, Benhar I. Targeted filamentous bacteriophages as therapeutic agents. Expert Opin Drug Deliv. 2008;5:321-9
24. Li Z, Conti PS. Radiopharmaceutical chemistry for positron emission tomography. Adv Drug Deliv Rev. 2010;62:1031-51
25. Giancotti FG, Ruoslahti E. Integrin signaling. Science. 1999;285:1028-32
26. Chen K, Chen X. Integrin targeted delivery of chemotherapeutics. Theranostics. 2011;1:189-200
27. Li ZB, Cai W, Cao Q. et al. 64Cu-labeled tetrameric and octameric RGD peptides for small-animal PET of tumor αvβ3 integrin expression. J Nucl Med. 2007;48:1162-71
28. Li ZB, Wu Z, Chen K, Chin FT, Chen X. Click chemistry for 18F-labeling of RGD peptides and microPET imaging of tumor integrin αvβ3 expression. Bioconjug Chem. 2007;18:1987-94
29. Wu Z, Li ZB, Cai W. et al. 18F-labeled mini-PEG spacered RGD dimer (18F-FPRGD2): synthesis and microPET imaging of αvβ3 integrin expression. Eur J Nucl Med Mol Imaging. 2007;34:1823-31
30. Zhang Y, Yang Y, Cai W. Multimodality Imaging of Integrin αvβ3 Expression. Theranostics. 2011;1:135-48
31. Cai H, Li Z, Huang CW, Park R, Shahinian AH, Conti PS. An improved synthesis and biological evaluation of a new cage-like bifunctional chelator, 4-((8-amino-3,6,10,13,16,19-hexaazabicyclo[6.6.6]icosane-1-ylamino)methyl)benzoic acid, for 64Cu radiopharmaceuticals. Nucl Med Biol. 2010;37:57-65
32. Cai H, Fissekis J, Conti PS. Synthesis of a novel bifunctional chelator AmBaSar based on sarcophagine for peptide conjugation and 64Cu radiolabelling. Dalton Trans. 2009;27:5395-400
33. Cannistra SA, Ottensmeier C, Niloff J, Orta B, DiCarlo J. Expression and function of β1 and αvβ3 integrins in ovarian cancer. Gynecol Oncol. 1995;58:216-25
34. Lang L, Li W, Jia HM. et al. New Methods for Labeling RGD Peptides with Bromine-76. Theranostics. 2011;1:341-53
35. Niu G, Chen X. Why integrin as a primary target for imaging and therapy. Theranostics. 2011;1:30-47
36. Li ZB, Cai W, Cao Q. et al. 64Cu-labeled tetrameric and octameric RGD peptides for small-animal PET of tumor αvβ3 integrin expression. J Nucl Med. 2007;48:1162-71
37. Kumar R, Dhanpathi H, Basu S, Rubello D, Fanti S, Alavi A. Oncologic PET tracers beyond [18F]FDG and the novel quantitative approaches in PET imaging. Q J Nucl Med Mol Imaging. 2008;52:50-65
38. Voss SD, Smith SV, DiBartolo N. et al. Positron emission tomography (PET) imaging of neuroblastoma and melanoma with 64Cu-SarAr immunoconjugates. Proc Natl Acad Sci U S A. 2007;104:17489-93
39. Cai H, Li Z, Huang CW. et al. Evaluation of Copper-64 Labeled AmBaSar Conjugated Cyclic RGD Peptide for Improved MicroPET Imaging of Integrin αvβ3 Expression. Bioconjug Chem. 2010;21:1417-24
Author contact
Corresponding author: Zibo Li, PhD. Molecular Imaging Center, Department of Radiology, University of Southern California. Fax: (+1) 323-442-3253 E-mail: ziboliedu or Liaohai Chen, PhD. Biosciences Division, Argonne National Laboratory. E-mail: lhchengov
 Global reach, higher impact
Global reach, higher impact