Impact Factor
Theranostics 2017; 7(16):3824-3841. doi:10.7150/thno.19890 This issue Cite
Research Paper
Metabolomic Profiling of Extracellular Vesicles and Alternative Normalization Methods Reveal Enriched Metabolites and Strategies to Study Prostate Cancer-Related Changes
1. Institute for Molecular Medicine Finland FIMM, University of Helsinki, Finland;
2. EV group, Division of Biochemistry and Biotechnology, Department of Biosciences and Division of Pharmaceutical Biosciences, Faculty of Pharmacy, University of Helsinki, Finland;
3. Finnish Red Cross Blood Service, Helsinki, Finland;
4. Division of Pharmaceutical Biosciences, Centre for Drug Research, Faculty of Pharmacy, University of Helsinki, Finland;
5. Metabolomics Unit, Institute for Molecular Medicine Finland FIMM, Helsinki, Finland;
6. Department of Pathology (HUSLAB), Helsinki University Hospital;
7. Science for Life Laboratory, Department of Oncology and Pathology, Karolinska Institutet, Solna, Sweden;
8. Department of Urology, Helsinki University Central Hospital, Helsinki, Finland;
9. Orion Corporation, Orion Pharma, Finland.
* equal contribution
Abstract
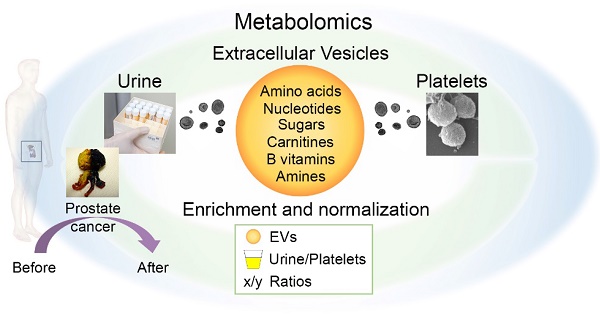
Body fluids are a rich source of extracellular vesicles (EVs), which carry cargo derived from the secreting cells. So far, biomarkers for pathological conditions have been mainly searched from their protein, (mi)RNA, DNA and lipid cargo. Here, we explored the small molecule metabolites from urinary and platelet EVs relative to their matched source samples. As a proof-of-concept study of intra-EV metabolites, we compared alternative normalization methods to profile urinary EVs from prostate cancer patients before and after prostatectomy and from healthy controls.
Methods: We employed targeted ultra-performance liquid chromatography-tandem mass spectrometry to profile over 100 metabolites in the isolated EVs, original urine samples and platelets. We determined the enrichment of the metabolites in the EVs and analyzed their subcellular origin, pathways and relevant enzymes or transporters through data base searches. EV- and urine-derived factors and ratios between metabolites were tested for normalization of the metabolomics data.
Results: Approximately 1 x 1010 EVs were sufficient for detection of metabolite profiles from EVs. The profiles of the urinary and platelet EVs overlapped with each other and with those of the source materials, but they also contained unique metabolites. The EVs enriched a selection of cytosolic metabolites including members from the nucleotide and spermidine pathways, which linked to a number of EV-resident enzymes or transporters. Analysis of the urinary EVs from the patients indicated that the levels of glucuronate, D-ribose 5-phosphate and isobutyryl-L-carnitine were 2-26-fold lower in all pre-prostatectomy samples compared to the healthy control and post-prostatectomy samples (p < 0.05). These changes were only detected from EVs by normalization to EV-derived factors or with metabolite ratios, and not from the original urine samples.
Conclusions: Our results suggest that metabolite analysis of EVs from different samples is feasible using a high-throughput platform and relatively small amount of sample material. With the knowledge about the specific enrichment of metabolites and normalization methods, EV metabolomics could be used to gain novel biomarker data not revealed by the analysis of the original EV source materials.
Keywords: extracellular vesicles, exosomes, metabolomics, urine, platelets, prostate cancer
 Global reach, higher impact
Global reach, higher impact