Impact Factor
Theranostics 2018; 8(7):1752-1765. doi:10.7150/thno.23180 This issue Cite
Research Paper
Genes involved in prostate cancer progression determine MRI visibility
1. Cedars Sinai Medical Center, Department of Surgery
2. Cedars Sinai Medical Center, Department of Biomedical Sciences
3. Cedars Sinai Medical Center, Biomedical Imaging Research Institute
4. Cedars Sinai Medical Center, Department of Pathology
5. Cedars Sinai Medical Center, Department of Radiology
6. Baylor College of Medicine, Cell Biology
*equal contribution
Abstract
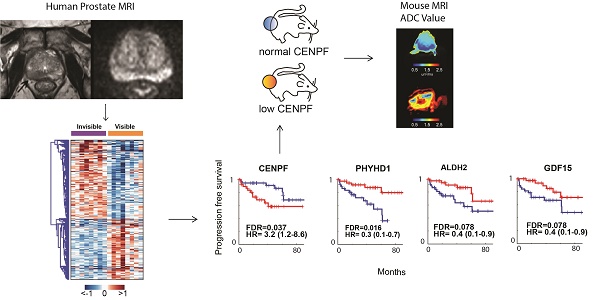
MRI is used to image prostate cancer and target tumors for biopsy or therapeutic ablation. The objective was to understand the biology of tumors not visible on MRI that may go undiagnosed and untreated.
Methods: Prostate cancers visible or invisible on multiparametric MRI were macrodissected and examined by RNAseq. Differentially expressed genes (DEGs) based on MRI visibility status were cross-referenced with publicly available gene expression databases to identify genes associated with disease progression. Genes with potential roles in determining MRI visibility and disease progression were knocked down in murine prostate cancer xenografts, and imaged by MRI.
Results: RNAseq identified 1,654 DEGs based on MRI visibility status. Comparison of DEGs based on MRI visibility and tumor characteristics revealed that Gleason score (dissimilarity test, p<0.0001) and tumor size (dissimilarity test, p<0.039) did not completely determine MRI visibility. Genes in previously reported prognostic signatures significantly correlated with MRI visibility suggesting that MRI visibility was prognostic. Cross-referencing DEGs with external datasets identified four genes (PHYHD1, CENPF, ALDH2, GDF15) that predict MRI visibility, progression free survival and metastatic deposits. Genetic modification of a human prostate cancer cell line to induce miR-101 and suppress CENPF decreased cell migration and invasion. As prostate cancer xenografts in mice, these cells had decreased visibility on diffusion weighted MRI and decreased perfusion, which correlated with immunostaining showing decreased cell density and proliferation.
Conclusions: Genes involved in prostate cancer prognosis and metastasis determine MRI visibility, indicating that MRI visibility has prognostic significance. MRI visibility was associated with genetic features linked to poor prognosis.
Keywords: prostate cancer, MRI, RNAseq, diffusion weighted imaging, prognosis
 Global reach, higher impact
Global reach, higher impact