Impact Factor
Theranostics 2019; 9(2):311-323. doi:10.7150/thno.29755 This issue Cite
Research Paper
Microhexagon gradient array directs spatial diversification of spinal motor neurons
1. Biomedical Institute for Global Health Research and Technology, National University of Singapore, 117599, Singapore
2. Institute of Molecular and Cell Biology, Agency for Science, Technology and Research, 138673, Singapore
3. Department of Biological Sciences, Faculty of Science, National University of Singapore, 117543, Singapore
4. Department of Physiology, Yong Loo Lin School of Medicine, National University of Singapore, 117456, Singapore
5. National Neuroscience Institute, 308433, Singapore
6. Key Laboratory for Major Obstetric Diseases of Guangdong Province, The Third Affiliated Hospital of Guangzhou Medical University, Guangzhou, 510150, China
7. Department of Biomedical Engineering, Faculty of Engineering, National University of Singapore, 117583, Singapore
8. Department of Surgery, Yong Loo Lin School of Medicine, National University of Singapore, 119228, Singapore
* These authors contributed equally
Abstract
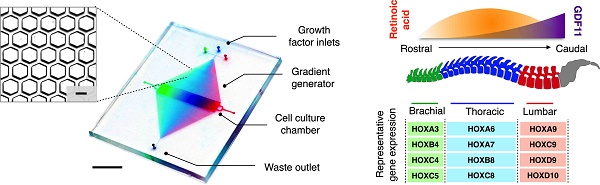
Motor neuron diversification and regionalization are important hallmarks of spinal cord development and rely on fine spatiotemporal release of molecular cues. Here, we present a dedicated platform to engineer complex molecular profiles for directed neuronal differentiation.
Methods: The technology, termed microhexagon interlace for generation of versatile and fine gradients (microHIVE), leverages on an interlocking honeycomb lattice of microstructures to dynamically pattern molecular profiles at a high spatial resolution. By packing the microhexagons as a divergent, mirrored array, the platform not only enables maximal mixing efficiency but also maintains a small device footprint.
Results: Employing the microHIVE platform, we developed optimized profiles of growth factors to induce rostral-caudal patterning of spinal motor neurons, and directed stem cell differentiation in situ into a spatial continuum of different motor neuron subtypes.
Conclusions: The differentiated cells showed progressive RNA and protein signatures, consistent with that of representative brachial, thoracic and lumbar regions of the human spinal cord. The microHIVE platform can thus be utilized to develop advanced biomimetic systems for the study of diseases in vitro.
Keywords: Microfluidics, molecular gradient, stem cell, motor neuron, spinal cord
 Global reach, higher impact
Global reach, higher impact