Impact Factor
Theranostics 2019; 9(7):1965-1979. doi:10.7150/thno.30958 This issue Cite
Research Paper
Acidic Microenvironment Up-Regulates Exosomal miR-21 and miR-10b in Early-Stage Hepatocellular Carcinoma to Promote Cancer Cell Proliferation and Metastasis
1. Sun Yat-sen University Cancer Center, State Key Laboratory of Oncology in South China, Collaborative Innovation Center of Cancer Medicine, Guangzhou, China
2. Department of Reproductive Medicine, The Second Affiliated Hospital of Guangzhou University of Chinese Medicine, Guangzhou, China.
3. Department of Pathology, Sun Yat-sen University Cancer Center, Guangzhou, China
4. Department of Pharmacology, College of Pharmacy, Jinan University, Guangzhou, China
5. Department of Medical Oncology, Sun Yat-sen University Cancer Center, Guangzhou, China
* These authors contributed equally.
Received 2018-10-25; Accepted 2019-1-27; Published 2019-3-16
Abstract
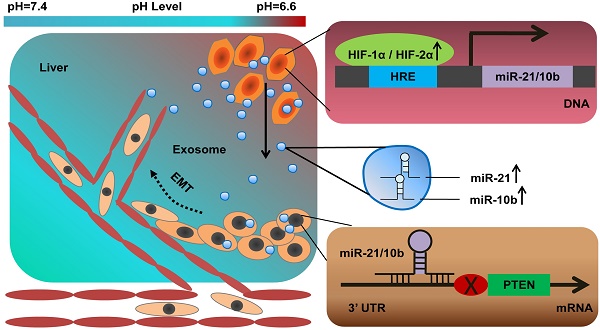
Rationale: The incidence of hepatocellular carcinoma is rising worldwide. It is predicted that nearly half of the early-stage hepatocellular carcinoma (E-HCC) patients will develop recurrence. Dysregulated pH, a hallmark of E-HCC, is correlated with poor prognosis. The acidic microenvironment has been shown to promote the release of exosomes, the membrane vesicles recognized as intercellular communicators associated with tumor progression, recurrence, and metastasis. We, therefore, aimed to identify exosomes induced by acidic microenvironment that may regulate E-HCC progression and to explore their mechanisms and clinical significance in E-HCCs.
Methods: miRNA microarray analysis and LASSO logistic statistic model were used to identify the main functional exosomal miRNAs. Invasion and scratch assays were performed to examine the migration and invasion of HCC cells. Immunoblotting and immunofluorescence were employed to detect the epithelial-to-mesenchymal transition (EMT) in HCC cells. Chromatin immunoprecipitation (ChIP) was used to analyze the binding of HIF-1α and HIF-2α to promoter regions of miR-21 and miR-10b.
Results: The acidic microenvironment in HCC was correlated with poor prognosis of patients. Exosomes from HCC cells cultured in the acidic medium could promote cell proliferation, migration, and invasion of recipient HCC cells. We identified miR-21 and miR-10b as the most important functional miRNAs in acidic HCC-derived exosomes. Also, the acidic microenvironment triggered the activation of HIF-1α and HIF-2α and stimulated exosomal miR-21 and miR-10b expression substantially promoting HCC cell proliferation, migration, and invasion both in vivo and in vitro. In E-HCC patients, serum exosomal miR-21 and miR-10b levels were associated with advanced tumor stage and HIF-1α and HIF-2α expression and were independent prognostic factors for disease-free survival of E-HCC patients. Most importantly, we developed a nano-drug to target exosomal miR-21 and/or miR-10b and examined its therapeutic effects against HCC in vivo.
Conclusion: Our findings suggested that the exosomal miR-21 and miR-10b induced by acidic microenvironment in HCC promote cancer cell proliferation and metastasis and may serve as prognostic molecular markers and therapeutic targets for HCC.
Keywords: acidic microenvironment, early-stage hepatocellular carcinoma, miR-21, miR-10b, epithelial- mesenchymal transition
Introduction
Hepatocellular carcinoma (HCC) is one of the most devastating malignancies worldwide [1]. During the past 20 years, the detection of patients with early-stage hepatocellular carcinoma (E-HCC) has increased, and surgery has greatly improved patients' survival, with a 5-year survival of 70% [2]. However, about half of E-HCC patients relapse after surgery [3] and metastatic spread is responsible for most E-HCC- associated morbidity and mortality [4]. Investigation of molecular mechanisms of E-HCC metastasis could identify prognostic biomarkers and/or useful therapeutic targets. E-HCC metastasis is a multistep process and is known to be regulated by the tumor microenvironment [5]. Although hypoxia in E-HCC has been studied extensively, dysregulated pH is less recognized in E-HCC, especially in oxygenated conditions. It is believed that dysregulation of the pH gradient in and around cancer cells, with intracellular pH (pHi) higher than 7.4 and extracellular pH (pHe) between 6.6 -7.2, creates a favorable environment for metastatic progression [6].
Extracellular acidification around E-HCC can be attributed to increased glycolysis in cancer cells. HCC cells in both primary and early malignant lesions consume glucose at a high rate even in oxygenated environments, a phenomenon known as Warburg effect [7]. Abnormal blood perfusion and hypoxia, coupled with a glycolytic phenotype, generate a pH gradient from the inner tumor region to adjacent normal tissues. This dysregulated pH gradient promotes cancer cell proliferation, metabolic adaptation, and migration and/or invasion both in vitro and in vivo [8, 9]. Treatments with pH-balancing reagents, such as bicarbonate, imidazole, and lysine, to neutralize tumor-derived acid can inhibit spontaneous and experimental metastasis [10, 11]. Extracellular acidification not only alters expression of metabolism-related genes, such as glucose transporter 1 (GLUT-1), carbonic anhydrase (CA-IX), or sodium-hydrogen exchanger 1 (NHE-1), it also activates some pH-dependent proteins that are involved in pro-metastatic transformation [12]. Recent studies revealed that normoxic acidosis triggers pH-dependent nucleolar sequestration of von Hippel-Lindau (VHL) protein, leading to activation of hypoxia-inducible factors (HIFs) [13], both of which regulate the transcription of numerous genes that are involved in metabolism, angiogenesis, cell proliferation, and metastasis. [14, 15].
Exosomes are membrane vesicles that are 30-100 nm in size and are released into the extracellular environment by various cell types. Although initially considered as “garbage transporters” from parental cells, exosomes are now recognized as a new category of intercellular communicators. Cells constitutively sort envelope proteins and RNAs into exosomes, a process that can be stimulated by a variety of pathophysiologic stimuli [16]. Exosomes bear surface molecules that render their attachment to target cells and activate signaling systems via ligand-receptor interactions or endocytosis. Through receiving a specific set of exosomes into the cytosol, a recipient cell modulates its physiological state [17]. Many studies have demonstrated that tumor-secreted exosomes can promote angiogenesis, modulate the immune system, support tumor progression, and remodel surrounding parenchymal tissues to promote pro-metastatic adaptation [18]. Exosomes isolated from the serum of cancer patients have been associated with tumor metastasis and/or relapse [19]. Taken together, tumor-derived exosomes are critical mediators of cancer progression. A recent report indicated that the acidic microenvironment promotes the release of exosomes [20]. In the present study, we investigated the relationship between the acidic microenvironment and exosomes with the aim to identify recurrence and/or metastasis-related biomarkers for designing new strategies to treat E-HCC.
Materials and Methods
Cell culture and development of acidic microenvironment culture
The human HCC cell lines SMMC-7721 and Hep3B were purchased from the China Center for Type Culture Collection (Wuhan, China) and the American Type Culture Collection (Manassas, VA, USA), respectively. Cells were cultured in Dulbecco's modified Eagle's medium (Invitrogen, Carlsbad, CA, USA) with 10% newborn calf serum at 37℃ with 5% CO2. Vesicle-depletion was performed by centrifuging cell culture medium at 100,000 g overnight. To simulate the tumor pH microenvironment, the cell culture medium was supplemented with 25 mmol/L PIPES (pH adjusted to 7.4) and 25 mmol/L of HEPES (pH adjusted to 6.6) as described previously [21]. Low pH-adapted cells were cultured and passaged directly in pH 6.6 medium for at least 4 weeks.
Exosome isolation and labeling
HCC cells (1×106) were cultured in vesicle- depleted medium for 48-72 hours. The medium was gathered and subjected to gradient centrifugation. Briefly, the medium was first centrifuged at 1000g for 10 min, at 3000g for 30 min, at 10,000g for 60 min, at 100,000g for 4 hours. All centrifugal steps were performed at 4 ℃. Exosomes from sera of HCC patients were isolated using ExoQuick exosome precipitation solution (SBI System Biosciences) according to the manufacturer's instructions [22]. The extracted exosomes were suspended in 300 μL PBS. The exosomes were quantified by BCA protein assay. A total of 10 µg exosomes were used for each in vitro experiment. For in vivo growth and metastasis assays, 1 µg/g exosomes were applied three times a week. We also used fluorescent dye Dil (Sigma) to label exosomes. Briefly, exosomes were incubated with Dil (1:2000) for 2 hours and then washed with PBS. The endocytosis of recipient cells was visualized using a confocal fluorescence microscope (Zeiss).
Patients
One hundred twenty-four surgery patients diagnosed with E-HCC at Sun Yat-sen University Cancer Center from January 1, 2009 to December 30, 2012 were screened in this study. E-HCC was defined as solitary tumors with diameters of < 5 cm and without vascular invasion [23]. A patient was excluded from the study if he or she had transarterial chemoembolization (TACE), radiotherapy, ablation, or liver transplant before resection. Patients were also excluded if they had no definitive diagnosis or follow-up data. The tumor stage and clinical stage were established using the 2003 Union for International Cancer Control/American Joint Committee on Cancer criterion. Healthy volunteers' blood samples were used as a control. All samples were obtained with informed consent. The study protocol was approved by the Institutional Review Board. The clinical-pathologic characteristic of the E-HCC patients are summarized in Table 1.
Correlation of clinical-pathologic characteristics of serum exosomal miR-21 and miR-10b in 124 E-HCC patients.
| Variable | All cases | Exosomal miR-21 | P * | Exosomal miR-10b | P * | ||
|---|---|---|---|---|---|---|---|
| Low | High | Low | High | ||||
| Age(years) | |||||||
| ≥48 | 61 | 28 | 33 | 0.479 | 34 | 27 | 0.289 |
| <48 | 63 | 33 | 30 | 29 | 34 | ||
| Sex | |||||||
| Male | 115 | 59 | 56 | 0.164 | 59 | 56 | 0.741 |
| Female | 9 | 2 | 7 | 4 | 5 | ||
| Hepatitis history | |||||||
| Yes | 108 | 51 | 57 | 0.293 | 50 | 58 | 0.014 |
| No | 16 | 10 | 6 | 13 | 3 | ||
| AFP(ng/ml) | |||||||
| ≤20 | 35 | 21 | 14 | 0.164 | 20 | 15 | 0.428 |
| >20 | 89 | 40 | 49 | 43 | 46 | ||
| Liver cirrhosis | |||||||
| yes | 96 | 48 | 48 | 0.831 | 49 | 47 | 0.923 |
| no | 28 | 13 | 15 | 14 | 14 | ||
| Tumor size(cm) | |||||||
| ≤3 | 44 | 37 | 17 | 0.005 | 31 | 13 | 0.001 |
| >3 | 80 | 34 | 46 | 32 | 48 | ||
| Tumor multiplicity | |||||||
| Signal | 98 | 49 | 49 | 0.826 | 52 | 46 | 0.255 |
| Multiple | 26 | 12 | 14 | 9 | 15 | ||
| Differentiation | |||||||
| Well-moderate | 67 | 40 | 27 | 0.012 | 40 | 27 | 0.047 |
| Poor-undifferentiated | 57 | 21 | 36 | 23 | 34 | ||
| Recurrence | |||||||
| No | 62 | 43 | 19 | 0.000 | 45 | 17 | 0.000 |
| Yes | 62 | 18 | 44 | 18 | 44 | ||
| HIF-1α | |||||||
| Negative | 65 | 44 | 21 | 0.001 | 49 | 16 | 0.000 |
| Positive | 69 | 27 | 42 | 24 | 45 | ||
| HIF-2α | |||||||
| Negative | 72 | 46 | 26 | 0.000 | 59 | 23 | 0.000 |
| Positive | 52 | 15 | 37 | 14 | 38 | ||
AFP: alpha fetoprotein; HIF: hypoxia-inducible factor.
Tissue pH measurement
The tissue pH was measured by microelectrode. Briefly, tissues were obtained from HCC patients or xenografted nude mice after surgery. The tissue pH was measured by inserting the glass electrode (LabSen 252, Sanxin, Shanghai, China) into the center of tumor tissues, as directed by the manufacturer's instruction. The center part from each tumor was selected and extracted for analyzing the mRNA expression of GLUT-1, CA9, MMP3, and MMP9 by Q-PCR and their correlation with the pH value in the same region of HCC tissues. The value of tissue pH was measured in triplicate for each sample.
Electron microscopy
The exosomes were examined by transmission electron microscopy as previously described [24]. Briefly, the samples were fixed with 2% glutaraldehyde and loaded on to Formvar carbon at room temperature. Subsequently, the samples were negatevely stained with 1% uranyl acetate for 3 minutes at 4 ℃, and dried under an electric incandescent lamp for 10 minutes. Photographs were taken using the JEM-1400 transmission electron microscope (JEOL, Tokyo, Japan) at 120 kV.
RNA isolation and quantitative real-time PCR
Total RNA was extracted by using TRIzol reagent (Invitrogen). cDNA was synthesized using the PrimeScript RT reagent Kit (Promega, Madison, WI, USA) as previously described [25]. Real-time PCR was performed using an ABI 7900HT Fast Real-time PCR system (Applied Biosystems, Foster City, California, USA) as previously described [25].
Oligonucleotide transfection, lentiviral packaging
GLUT-1 siRNA, CA9 siRNA, miR-21-5p siRNA, miR-10b-5p siRNA, miR-21 mimics, miR-10b mimics, HIF-1α shRNA and HIF-2α shRNA were synthesized by Kangcheng biotechnology company (Guangzhou, China). The pCDH-CMV-MCS-EF1-coGFP plasmid was used to construct virus particles. This plasmid, together with packaging plasmids pCMV/pVSVG, pRSV/pREV, and pMDLG/pRRE, were transfected into 293FTcells using Lipofectamine 2000 reagent (Invitrogen). After 48 hours, virus particles were harvested from the cell supernatant. SMMC-7721 and Hep3B cells were transfected with centrifuged lentivirus plus 8 mg/ml polybrene (Sigma, St Louis, MO, USA). Oligonucleotide transfection was performed with Lipofectamine 2000 reagent (Invitrogen). U6 snRNA was used as a positive control, and reactions without reverse transcriptase or RNA template were included as negative controls.
PDCM system
The nanoparticles (PDCM) are composed of 1-palmitoyl-2-oleoyl-snglycero-3-phosphocholine (POPC), 1,2-dioleoyl-snglycero-3-phosphoethanolamine (DOPE), cholesteryl hemisuccinate (CHEMS), and cholesteryl-4-((2-(4-morpholinyl) ethyl) amino)-4-oxoburanoate (MOCHOL). The designed nanoparticles have “pH-tunable” characteristic, which facilitates efficient delivery relying on the surrounding pH [26]. Preliminary experiments and clinical trials showed good systemic biodistribution of the PDCM system [26-28]. PCDM-GLUT-1, PDCM-CA9, PDCM-miRNA- 21, and PDCM-miRNA-10b were synthesized and assembled by Kangcheng biotechnology company (Guangzhou, China).
miRNA microarray
Sample preparation and miRNA microarrays (Human miRNA Microarray, Release 21.0, Agilent) were performed at Biomedlab Company (Shanghai, China). Briefly, total RNA (2.5 μg) extracted from exosomes was labelled with pCp-DY647 (Dharmacon, Lafayette, CO, USA), hybridized onto microarray, scanned using a LuxScan 10K Microarray Scanner (CapitalBio, Beijing, China) and analyzed using GenePix Pro 6.0 software (Axon Instruments, Foster City, CA, USA). The microarray data have been deposited online under accession number GSE118647.
MTT assay
MTT proliferation assay (Sigma) was performed according to the manufacturer's protocol. A total of 10 µg exosomes was used for each group according to the experimental design. Viability was measured at different time points from 12 h to 72 h after treatment.
Scratch assay
The scratch assay was used to measure cell migration in vitro. In brief, cells were seeded in 60mm dishes and cultured until they formed a fused monolayer. A P200 pipette tip was used to create a scratch following which cells were incubated with different exosomes (10 µg). After 48 h, wound closure was photographed by a microscope. The migration rate was measured by the fraction of cell coverage across the scratch. The migration rate (%) = migration area/total area ×100%.
Invasion assay
Matrigel invasion chambers (BD Biosciences, NJ, USA) were used to measure cell invasion. A total of 10 µg exosomes was used for each group according to the experimental design. Briefly, 1×104 cells were added to the upper chamber, and FBS was added to the lower chamber. After 48 h, the chambers were fixed and stained with 0.05% crystal violet for 2h. The cells on the upper surface were gently scraped. The stained cells were counted and photographed under a microscope (×200).
Immunohistochemistry and immunofluorescence
The slides were dried, dewaxed, rehydrated, and blocked. Next, the sliders were boiled and then incubated with the anti-HIF-1α antibody (1:500, ProteinTech Group, Inc.) and anti-HIF-2α antibody (1:500, ProteinTech Group, Inc.) in a moist chamber. After 3 washes, the slides were incubated with a secondary antibody (Envision, Dako, Glostrup, Denmark) and then stained with DAB (3,3-diaminobenzidine). Nuclei were stained with hematoxylin. The cutoff value (45% positive cells) of HIF-1α/ HIF-2α immunoreactivity was determined by receiver operator curve (ROC) analysis. For immunofluorescence, slides were incubated with rabbit anti-E- cadherin (1:200, Abcam) and mouse anti-vimentin (1:200, Abcam) for 2 hours. Slides were incubated for 1 hour with Alexa Fluor 488 goat anti-mouse IgG (1:800, Cell Signaling Technology) and Alexa Fluor 594 goat anti-rabbit IgG (1:800, Cell Signaling Technology).
Western blot analysis
Western blotting was performed using standard procedures. Briefly, protein samples (25mg) were separated on polyacrylamide-SDS gels and electrophoretically transferred to polyvinylidene difluoride membranes (Millipore). Subsequently, membranes were blocked and incubated with mouse anti-CD81 (1:1000, Abcam), mouse anti-CD63 (1:1000, Abcam), mouse anti-HIF-1α (1:1500, ProteinTech Group, Inc.), mouse HIF-2α (1:1500, ProteinTech Group, Inc.), rabbit anti-PTEN (1500, Abcam), rabbit anti-E-cadherin (1:1000, Abcam), mouse anti-vimentin (1:1000, Abcam), rabbit anti-mTOR (1:1000, Abcam), rabbit anti-AKT (1:1000, Abcam), rabbit anti-pAKT (1:1000, Abcam), and rabbit anti-tubulin (1:1000 Abcam) antibodies. Anti-rabbit or anti-mouse IgG was used as the secondary antibody (1:8000). The signal intensity was evaluated using Quantity One 4.4.0 software.
In vivo growth and metastasis assay
The In vivo growth and metastasis assay were performed as previously described [29] and are illustrated in Supplementary Figure S14. Briefly, for in vivo growth assays, 2×106 SMMC-7721 cells were injected subcutaneously into the flanks of BALB/ C-nu/nu athymic nude mice. The tumor size was measured using Vernier calipers every 7 days. For in vivo metastasis assay, the subcutaneous tumors were resected from growth assays and formed into 1mm3 cubes. These tumor cubes were implanted in the left lobes of the livers of nude mice. Exosomes (three times a week, 1 µg/g for each time) were used for in vivo growth and metastasis assay. On day 50, the liver and lung were dissected, fixed by formalin, and stained with haematoxylin & eosin. Consecutive tissue sections were used to examine pulmonary metastatic nodules. All experimental procedures involving animals were performed according to the institutional ethical guidelines.
Chromatin immunoprecipitation (ChIP) assay
The ChIP Assay Kit (Abcam) was used according to the manufacturer's protocol. Briefly, cells were lysed in cell lysis buffer to generate DNA fragments. Nonspecific IgG (1:200, Sigma), mouse anti-HIF-1α (1:200, Abcam), and rabbit anti-HIF-2α antibodies (1:200, Abcam) were incubated with lysates to precipitate target regions. The promoter fragment (contain HIF response element, HRE) was extracted and visualized by gel. The region of the miR-130b promoter was used as a negative control.
LASSO Logistic regression
The least absolute shrinkage and selection operator (LASSO) is an alternative method for fitting sparse models for regression with high-dimensional predictors. To eliminating variables, the L1 penalty could force coefficient estimates to zero [30, 31]. The logistic regression model with LASSO penalty was used to achieve variable shrinkage and selection. Optimal values of λ were determined by repeated cross-validations. The optimal λ is the largest value for which the partial likelihood deviance is within 1-s.e. of the smallest value of partial likelihood deviance. R software version 3.0.1 (R Foundation for Statistical Computing) and the 'glmnet' package were used to perform the LASSO regression analysis.
Statistical analysis
SPSS software (SPSS Standard v 17.0) was used to perform statistical analysis. The most functional miRNAs with non-zero coefficients were determined by LASSO logistic regression in TCGA dataset. The correlation between serum exosomal miR-21 and miR-10b levels and clinical-pathologic features was assessed with the Chi-square test. Survival curves were calculated by Kaplan-Meier analysis and compared by the log-rank test. Multiple Cox proportional hazards regression analysis was performed to assess the survival variables. Data are presented as mean ± SE and assessed by the student's test. P values < 0.05 were considered significant.
Results
The acidic microenvironment in HCC is correlated with poor survival of patients
We first analyzed the TCGA-HCC database to screen for genes that are related to the acidic microenvironment and correlated with HCC patients' survival (Supplementary Figure S1). We found that GLUT-1, CA9, GCK, NHE1, and CA12 expression was significantly increased in HCC samples compared to adjacent normal tissues (P < 0.05, Figure 1A and Supplementary Figure S2). Furthermore, Kaplan- Meier and ROC analyses showed that the expression of GLUT-1 and CA9 was significantly correlated with poor overall survival of HCC patients (P < 0.001, Figure 1B and Supplementary Figure S3 and S4). We examined GLUT-1 and CA9 expression in fresh tissues from 20 E-HCC patients and found that they were negatively correlated with extracellular pH (P < 0.05, Figure 1C). The activity of MMP-3 (matrix metalloproteinase-3) and MMP-9 (matrix metalloproteinase- 9) was reported to be high in acidic microenvironment [32-35]. Therefore, we first confirmed significantly increased expression of MMP-3 and MMP-9 in the acidic microenvironment in HCC cell lines (P < 0.05, Figure 1D). The expression of these two metalloproteases was also negatively correlated with extracellular pH in fresh tissues of 20 E-HCC patients (Figure 1E). Also, MMP-3 and MMP-9 expression was significantly correlated with poor overall and disease-free survival (DFS) in TCGA-HCC database and SYSUCC E-HCC patients (P < 0.05, Figure 1F and 1G).
Acidic microenvironment correlated with poor survival. A. Expression of GLUT-1 and CA9 in HCC tissues and adjacent normal tissues in TCGA datasets. B. Correlation of overall survival and GLUT-1/CA9 gene expression by Kaplan-Meier analysis in HCC patients in TCGA datasets. C. Correlation of GLUT-1/CA9 expression and extracellular pH in 20 E-HCC patients' fresh tissues in vitro. D. Expressions of MMP3 and MMP9 in SMMC-7721 cells cultured under pH 6.6 and pH 7.4 conditions. E. Correlation of MMP3/MMP9 expression and extracellular pH in the 20 E-HCC patients' fresh tissues. F. Correlation of overall survival and MMP3/MMP9 expression by Kaplan-Meier analysis in HCC patients in TCGA datasets. G. Correlation of disease-free survival and MMP3/MMP9 gene expression by Kaplan-Meier analysis in E-HCC patients in SYSUCC datasets. H. Tumor volumes in PBS, NaHCO3, PDCM-ctrl, PDCM-CA9, PDCM-GLUT-1, and PDCM-CA9 + PDCM-GLUT-1 groups. I. Numbers of lung metastases numbers of cell groups as in (H). J. Extracellular pH of cell groups as in (H). *, P<0.05. **, P<0.01.
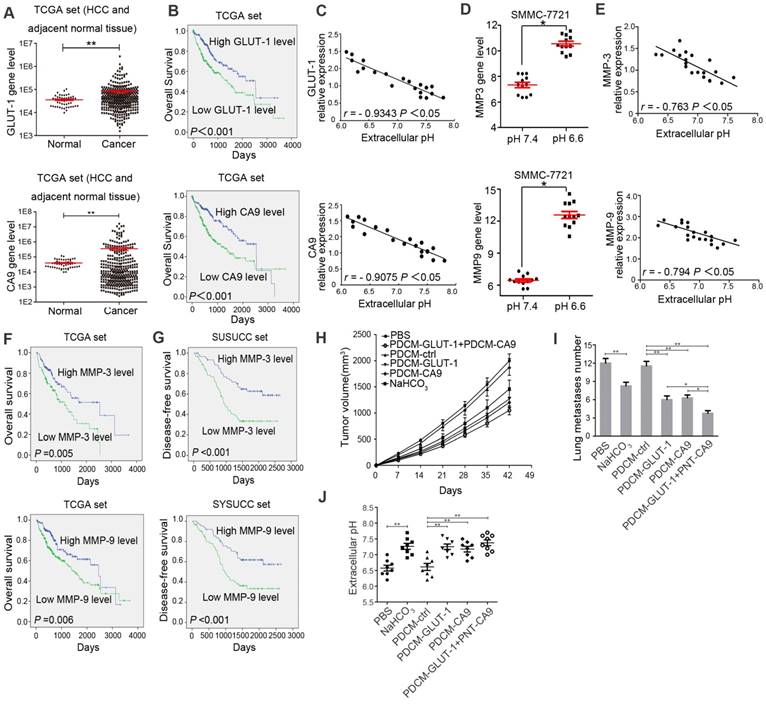
To evaluate the role of extracellular pH on HCC cell proliferation and metastasis in vivo, we took advantage of specialized liposomes (PDCM, trade name SMARTICLES), which facilitate transmembrane diffusion in a pH-dependent manner. Based on PDCM molecular properties, PDCM-CA9 and PDCM- GLUT-1 could significantly decrease CA9 and GLUT-1 levels in HCC cells (P < 0.05, Supplementary Figure S5A and S5B). Knocking-down expression of CA9 and GLUT-1, as well as NaHCO3 injection, could elevate the acidic pHe [12, 36, 37]. NaHCO3, PDCM-CA9, PDCM-GLUT-1, and PDCM-CA9 + PDCM-GLUT-1 could increase extracellular pH and decrease tumor growth and lung metastases, indicating that low extracellular pH may play a role in tumor proliferation and invasion (P < 0.05, Figure 1H, 1I, and 1J).
Exosomes derived from HCC cells cultured under acidic condition promote cell migration and invasion of recipient HCC cells cultured under the normal condition
We isolated extracellular vesicles from the medium of HCC cells (SMMC-7721 and Hep-3B) cultured under pH 6.6 and pH 7.4 by multistep ultracentrifugation. Western blot analysis showed that the nanovesicles expressed exosome markers CD63 and CD81 (Figure 2A). Transmission electron microscopy showed diameters of extracellular vesicles ranging from 50 to 200 nm (Figure 2C). These nanovesicles were homogeneous in appearance and showed features of exosomes.
Acidic HCC cell-derived exosomes promote migration and invasion of the non-acidic recipient HCC cells. A. Western blots of CD63 and CD81 in exosomes derived from SMMC-7721 and Hep3B cells cultured under different pHs. B. Western blot analysis of HIF-1α and HIF-2α in normoxia. C. Transmission electron microscopy of exosomes isolated from the cell culture supernatant. D. Confocal microscopy of the Dil-labeled exosomes (red). E. MTT assays of the proliferation of recipient cells treated with acidic HCC-derived exosomes. F. Invasion of recipient cells treated with acidic HCC-derived exosomes. G. Migration of recipient cells treated with acidic HCC-derived exosomes. Experiments performed in triplicates. Non-Ac exo, the exosomes derived from HCC cells cultured in the non-acidic medium. Ac mock exo, exosomes derived from HCC cells cultured in the acidic medium. Ac ctrl shRNA exo, exosomes derived from HCC cells transfected by empty vector cultured in the acidic medium. Ac HIF-1 KD exo, exosomes derived from HIF-1α KD HCC cells cultured in the acidic medium. Ac HIF-2 KD exo, exosomes derived from HIF-2α KD HCC cells cultured in the acidic medium. Ac HIF-1+2 KD exo, exosomes derived from HIF-1α and HIF-2α KD HCC cells cultured in the acidic medium. *, P < 0.05. **, P < 0.01.
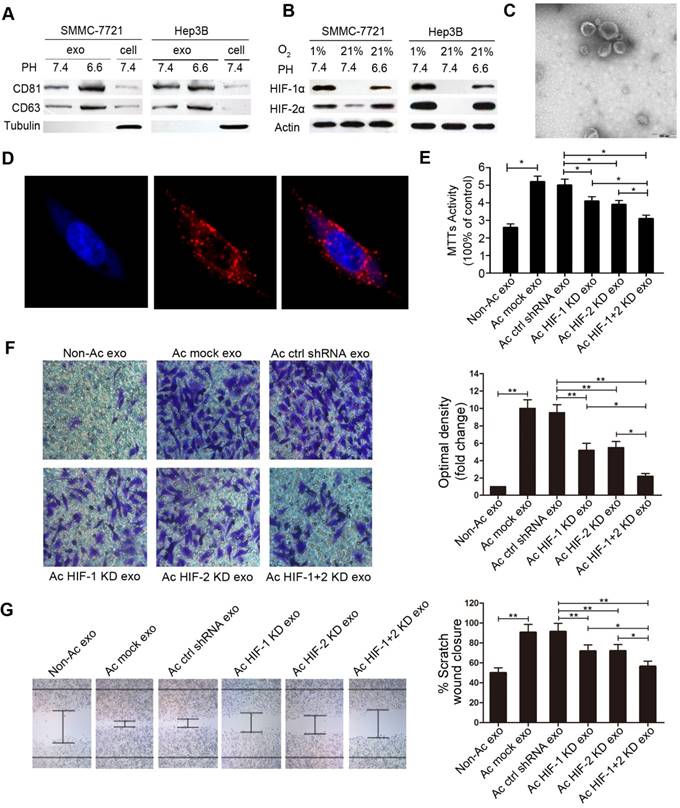
To test whether exosomes derived from HCC cells cultured under acidic condition could influence the biological behavior of recipient cells, we first performed fluorescence microscopy to visualize the endocytosis of exosomes by recipient cells. As shown in Figure. 2D, Dil-labeled exosomes derived from HCC cells cultured under acidic condition were endocytosed by normal HCC cells after incubation for 2 hours. Subsequently, we investigated the effects of acidic condition-derived exosomes on oncogenic properties of non-acidic HCC cells, including cell proliferation, migration, and invasion. MTT, invasion and migration assays showed that acidic HCC- derived exosomes significantly increased cell proliferation, migration, and invasion of recipient cells compared with non-acidic HCC-derived exosomes (Figure 2E, 2F, 2G, and Supplementary Figure S6). As previously reported, we verified that acidifying the extracellular milieu of HCC cells could trigger the activation of HIF-1α and HIF-2α in normoxia (Figure 2B). Intriguingly, exosomes derived from HIF-1α and HIF-2α knockdown (KD) cells cultured under acidic condition could not increase cell proliferation, migration, and invasion (Figure 2E, 2F, 2G, and Supplementary Figure S6).
Identification of candidate functional miRNAs in acidic HCC cell-derived exosomes
It is well known that exosomes could deliver both mRNAs and miRNAs to recipient cells and affect cellular functions [38]. To identify exosomal miRNAs that may play an oncogenic role, we performed comparative microarray profiling of miRNAs derived from exosomes from the supernatant of SMMC-7721 cells cultured under normal (pH 7.4) or acidic conditions (pH 6.6) for at least 4 weeks.
Five hundred forty-five and 802 known miRNAs were identified in acidic and non-acidic exosomes, respectively, of which 535 miRNAs were shared by both (Figure 3A). We compared the expression levels of miRNAs between acidic and non-acidic exosomes. Using a threshold cutoff of 3-fold and P < 0.05, we discovered that expression of 81 miRNAs was significantly different between the two kinds of exosomes (Supplementary Table S1), of which 70 miRNAs were upregulated in acidic exosomes compared with non-acidic exosomes. Gene ontology (GO) and KEGG analysis showed that genes regulating EMT (epithelial-mesenchymal transition) and PI3K- AKT signaling were among top upregulated clusters. (Figure 3B, Supplementary Figure S7 and S8).
Identification of functional miRNAs based on miRNA expression profiles of exosomes derived from acidic HCC cells and TCGA dataset. A. The pie chart shows different miRNAs from acidic and non-acidic exosomes. B. Gene ontology (GO) analysis and KEGG analysis of miRNAs identified in (A). C. The volcano plot shows 84 differentially expressed miRNAs from 49 pairs of samples of TCGA dataset. D. 18 miRNAs identified by the LASSO logistic regression model with non-zero coefficients. E. Schematic illustration for the selection of candidate miRNAs.
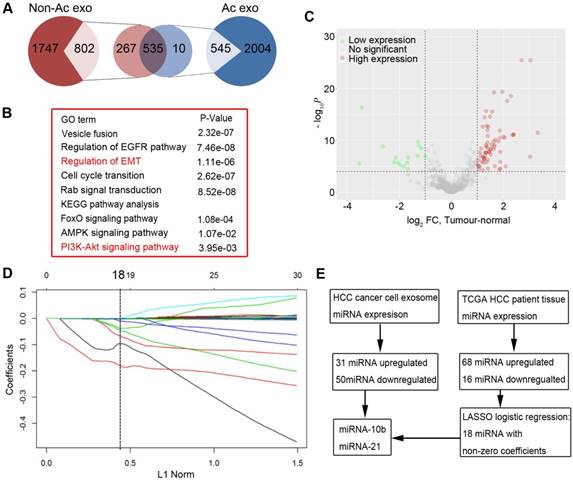
To further identify the functional miRNAs in acidic HCC-derived exosomes, we analyzed 49 paired HCC and adjacent normal tissues in the TCGA data set to identify differentially expressed miRNAs in these samples. The volcano plot showed that the 84 miRNAs showed more than 2-fold change in the 49 pairs of samples (t-test, all P < 0.0001, Figure 3C). We used a LASSO (the least absolute shrinkage and selection operator) logistic regression model to analyze the 84 miRNAs and found that 18 had non-zero coefficients (Figure 3D and Supplementary Figure S9). miR-21 and miR-10b were upregulated in both TCGA dataset and exosome microarray analyses and therefore were considered as the most important functional miRNAs in acidic HCC-derived exosomes (Figure 3E).
Acidic condition stimulates exosomal miR-21 and miR-10b expression in HIF-1α and/or HIF-2α-dependent manner
We further confirmed the miRNA microarray results by real-time PCR. Consistent with the microarray results, exosomes secreted from HCC cells cultured under pH 6.6 had higher levels of miR-10b, miR-21, miR-223, and miR-940 than those derived from cells cultured under pH 7.4 condition (Figure 4A). We focused on miR-21 and miR-10b for further study because they were extracted from the LASSO logistic model and were higher in acidic exosomes (Figure 3E). Under normal condition, cellular and exosomal levels of miR-21 and miR-10b were not influenced by knocking-down either HIF-1α or HIF-2α (Figure 4B and 4C). However, under acidic condition, knocking-down HIF-1α or HIF-2α led to decreased levels of both cellular and exosomal miR-21 and miR-10b (Figure 4B and 4C). We also used an additional HCC cell line Hep3B to confirm these results (Supplementary Figure S10).
Acidic microenvironment induces exosomal miR-21 and miR-10b expression through HIF-1α or HIF-2α. A. Comparison of the levels of miR-10b, miR-21, miR-223 and miR-940 from acidic and non-acidic media by RT-PCR. B and C. Comparison of the levels of cellular and exosomal miR-21 and miR-10b in HIF-1α or HIF-2α knockdown cells by RT-PCR. D. Binding of HIF-1α and HIF-2α to the predicted HRE regions in miR-21 and miR-10b promoters by ChIP assays. Input DNA was used as positive control and miR-130b was used as a negative control. E. Expression of cellular miR-21 and miR-10b in SMMC-7721 and Hep3B cells cultured under pH 6.6 and pH 7.4. F. The relationship between exosomal miR-21/miR-10b levels and pH in normal HCC cells and miR-21/miR-10b mimics-transfected HCC cells. Experiments performed in triplicates. *, P<0.05. **, P<0.01. Non-Ac condition, SMMC-7721 cells were cultured in pH 7.4 medium. Ac condition, SMMC-7721 cells were cultured in pH 6.6 medium. HIF-1 KD, HIF-1α KD SMMC-7721 cells. HIF-2 KD, HIF-2α KD SMMC-7721 cells. HIF-1+2 KD, HIF-1α and HIF-2α KD SMMC-7721 cells.
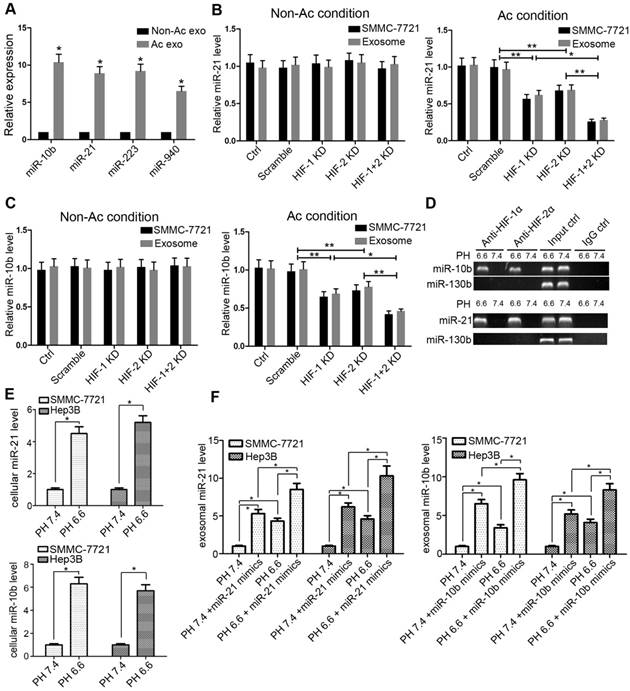
HIF-1α and HIF-2α bind to the HRE sites in promoters. We next performed ChIP assay to validate the binding of HIF-1α and HIF-2α to the HRE regions of miR-21 and miR-10b promoters. As shown in Fig 4D, anti-HIF-1α or HIF-2α antibodies immunoprecipitated genomic DNAs containing miR-21 and miR-10b HRE-containing promoter regions in cells cultured under pH 6.6 conditions. In contrast, the miR-130 promoter region (without HRE region) did not show recruitment of HIF-1α or HIF-2α under acidic condition. The levels of miR-21 and miR-10b were higher in SMMC-7721 and Hep3B cell lines cultured under acidic condition than non-acidic condition (P < 0.05, Figure 4E). Also, after transfection of miR-21/10b into both HCC cell lines, we found that the levels of exosomal miR-21 and miR-10b were substantially increased in cells cultured under acidic condition (Figure 4F). These data provided evidence that in HCC cells cultured under acidic condition, increased HIF-1α and HIF-2α could directly bind to the HRE-containing promoter regions of miR-21 and miR-10b and up-regulate their cellular expression. The increased cellular expression of miR-21 and miR-10b may result in the elevated levels of these miRNAs in exosomes. However, the precise mechanism for increased loading of these miRNAs in exosomes remains unclear and needs to be investigated. Furthermore, silencing HIF-1α and/or HIF-2α in SMMC-7721 and Hep3B cells under acidic condition resulted in decreased levels of cellular miR-210 (another target of HIF-1α), whereas the levels of exosomal miR-210 were not altered (Supplementary Figure S11). These results indicated complex regulation of expression for individual miRNAs and their loading into exosomes.
miR-21 and miR-10b mediate acidic exosome-induced HCC cell proliferation, migration, and invasion
Knockdown of miR-10b and miR-21 in acidic exosomes significantly decreased cell proliferation, migration, and invasion of recipient cells compared with non-acidic-induced exosomes (Figure 5A-5E, Supplementary Figure S12). To study whether cell migration and invasion induced by miR-21 and miR-10b from acidic exosomes depend on HIF-1α or HIF-2α, we overexpressed miR-21 and miR-10b in HIF-1α- or HIF-2α knocked-down cells. Transfection of miR-21 and miR-10b mimics in HIF-1α- or HIF-2α knocked-down cells restored miR-21 and miR-10b levels in exosomes (Supplementary Figure S13). Furthermore, restoration of miR-21 and miR-10b levels in acidic exosomes increased recipient cell proliferation, migration, and invasion (Figure 5F-5J). We also examined whether exosomal miR-21 and miR-10b induced EMT-related changes in HCC cells, and found that exosomal miR-21 and miR-10b increased Vimentin and Snail expression while decreased PTEN and E-cadherin levels in treated cells (Figure 6A). These results suggested that acidic exosome-induced cell migration and invasion were dependent on miR-21 and miR-10b and these two exosomal miRNAs could also induce EMT in the recipient cells.
miR-21 and miR-10b mediated acidic exosome-induced cell proliferation, migration, and invasion. A, F, MTT assay; B, D, G and I, invasion assay; C, E, H and J, scratch assay of cancer cells modified as depicted in the Figure. Experiments performed in triplicates. *, P < 0.05.
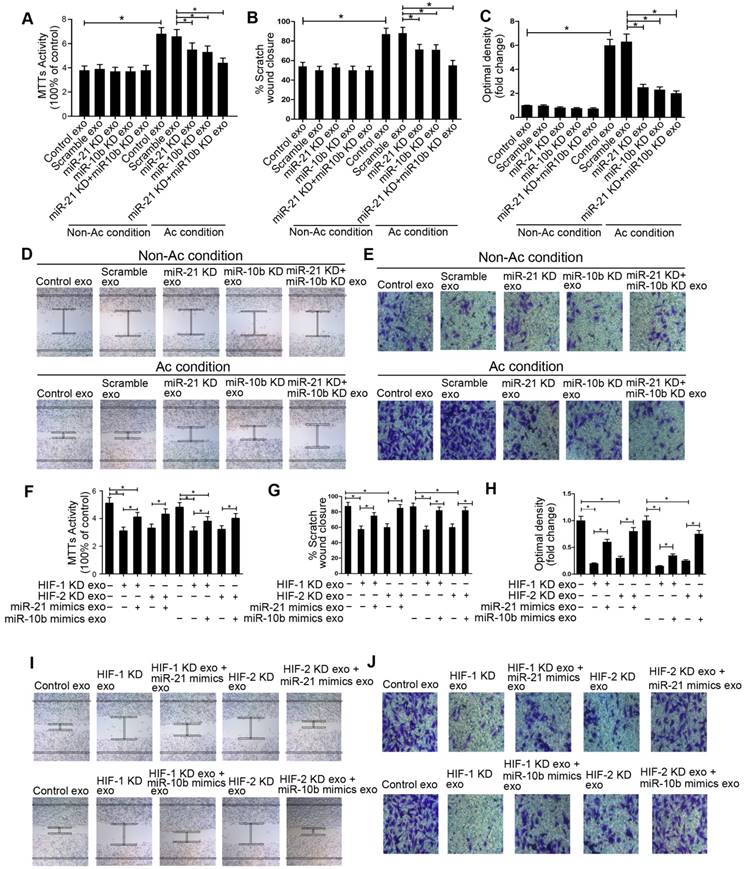
Acidic exosomal miR-21 and miR-10b promote tumor growth and metastasis in xenograft models. A. miR-21 and miR-10b could bind to 3' UTR of PTEN. Western blot analysis of PTEN, p-AKT, E-cadherin, mTOR, Snail and Vimentin. B. Tumor volumes in PBS, non-acidic exosome, miR-21 KD exosome, miR-10b KD exosome, miR-21 mimics exosome, miR-10b mimics exosome, and acidic exosome groups of cancer cells. C. Lung metastases numbers of groups of cancer cells as in (B). D. Western blot analysis of PTEN, E-cadherin, and Vimentin of cancer cells. E. Immunofluorescence staining of E-cadherin and Vimentin.
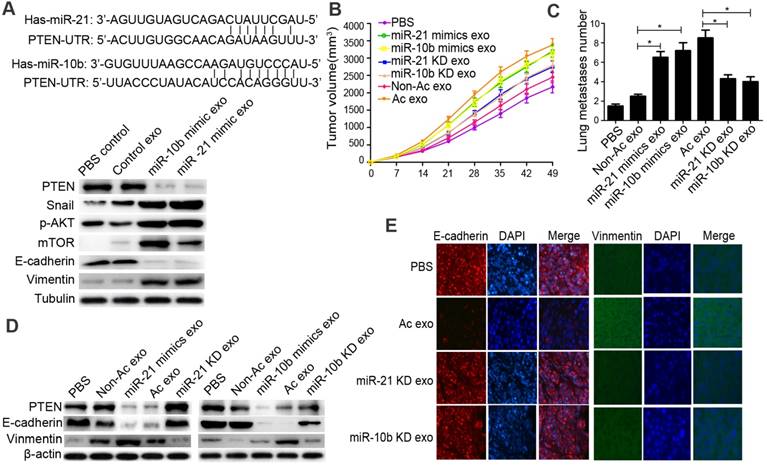
Acid-derived exosomal miR-21 and miR-10b promote HCC cell growth and metastasis in a xenograft model
We used a xenograft model and a lung metastasis model to investigate the effects of tumor- derived exosomes on tumor growth and metastasis (Supplementary Figure S14). Overexpression of miR-21 and miR-10 in acid-derived exosomes increased tumor growth and tumor volume. The tumor volume and numbers of lung metastases were significantly increased in mice injected with acid-treated or miR- 21/10b-overexpressing exosomes-treated cells (Figure 6B and 6C). Knock-down of miR-21 and miR-10b expression in acidic exosomes significantly decreased the tumor volume and numbers of lung metastases (Figure 6B and 6C). We also investigated whether exosomal miR-21 and miR-10b regulated EMT in vivo. Consistent with the in vitro experiments, the expression of Vimentin was increased, while the expression of E-cadherin was decreased in acid-treated and miR-21/10b-overexpressing exosomes (Figure 6D and 6E, Supplementary Figure S15).
miR-21 and miR-10b derived from exosomes are correlated with disease-free survival in E-HCC patients
We next isolated and examined the exosomes from sera of patients with E-HCC and healthy volunteers. Electron microscopy showed that the precipitated vesicles had the same size distribution consistent with exosomes (50-200 nm) (Figure 7A). The expression of miR-21 and miR-10b in E-HCC patient exosomes was significantly increased compared with that in exosomes from healthy volunteers (Figure 7B). We also found that the exosomes derived from the sera of E-HCC patients could increase cell proliferation, migration, and invasion compared with exosomes from healthy volunteers (Figure 7C and 7D). Furthermore, the expression of miR-21 and miR-10b was negatively correlated with extracellular pH in fresh tissues from 20 E-HCC patients (Supplementary Figure S16).
Acidic serum exosomal miR-21 and miR-10b correlate with disease-free survival in E-HCC patients. A. Electron micrograph of exosomes isolated from E-HCC patients' sera. B. Levels of serum exosomal miR-21 and miR-10b between E-HCC patients and healthy volunteers. C. Proliferation of recipient cells treated with exosomes derived from E-HCC patients' sera by the MTT assay. D. Migration and invasion of recipient cells treated with exosomes derived from E-HCC patients' sera using the invasion assay and scratch assay. E. Kaplan-Meier analysis of the association between exosomal miR-21/miR-10b and E-HCC patients' disease-free survival. F. ROC analysis of the sensitivity and specificity of exosomal miR-21 and miR-10b compared with tissue miR-21/miR-10b and AFP in 124 E-HCC patients. G. Tumor volumes of PBS, PDCM-ctrl, PDCM-miR-21, PDCM-miR-10b, PDCM-miR-21+PDCM-miR-10b groups of cancer cells. H. Numbers of lung metastases of groups of cells as in G. Experiments were performed in triplicates. *, P < 0.05. **, P < 0.01.
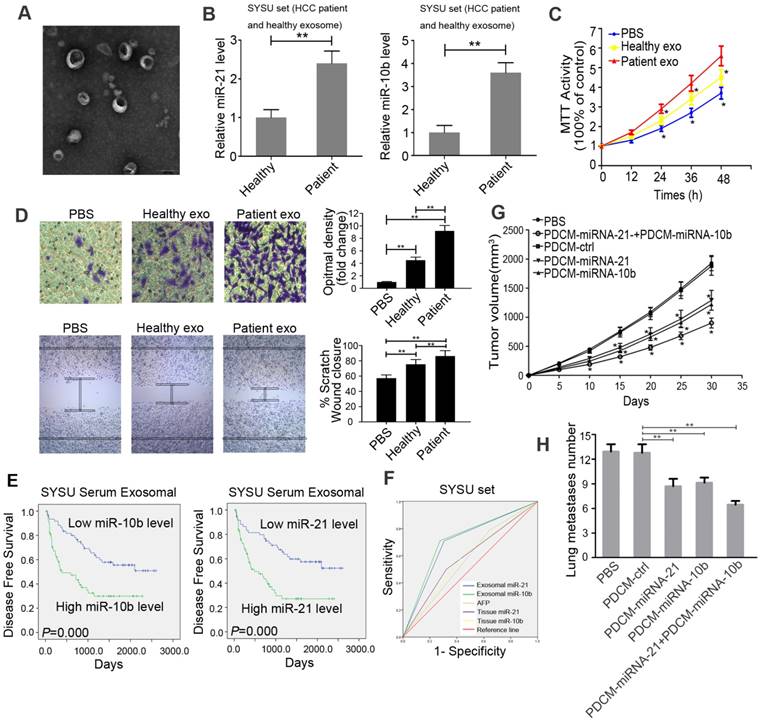
In 124 E-HCC patients who underwent surgery, serum exosomal miR-21 and miR-10b levels were closely associated with tumor size, cell differentiation, and recurrence. (Table 1). HIF-1α and HIF-2α expression were used to determine the correlation between exosomal miR-21 and miR-10b and acidic microenvironment (Supplementary Figure S17). As shown in Table 2, serum exosomal miR-21 and miR- 10b significantly correlated with HIF-1α and HIF-2α staining. Kaplan-Meier analysis revealed that high expression of serum exosomal miR-21 and miR-10b was associated with poor disease-free survival (Figure 7E). Multivariate Cox regression analysis also indicated that high levels of serum exosomal miR-21 and miR-10b were independent prognostic factors for the poor survival of E-HCC patients (Table 2). ROC analysis demonstrated that as prognostic factors, sensitivity and specificity of exosomal miR-21 and miR-10b were significantly higher than tissue miR-21/miR10b and AFP (Figure 7F). We further developed a nano-drug targeting exosomal miR-21 and miR-10b based on the PDCM system. In an in vivo assay, both PDCM-miR-21 and PDCM-miR-10b could significantly decrease tumor growth and numbers of metastatic lung nodules, indicating that exosomal miR-21 and miR-10b could serve as potential therapeutic targets (Figure 7G and 7H).
Univariate and multivariable analyses of clinical characteristics with disease-free survival of patients with E-HCC.
| Clinical variable | Disease-free survival | |
|---|---|---|
| HR(95% Cl) | p value | |
| Univariate analysis | ||
| Age (≥48.4 years vs. <48.4 years) | 0.902(0.547-1.486) | 0.685 |
| Sex (Male vs. Female) | 0.797(0.289-2.196) | 0.661 |
| Hepatitis history (No vs. Yes) | 1.381(0.595-3.206) | 0.453 |
| AFP (≤20 vs. >20) | 1.881(1.019-3.472) | 0.043 |
| Liver cirrhosis (No vs. Yes) | 1.056(0.572-1.946) | 0.863 |
| Tumor size (≤3cm vs. >3cm) | 2.621(1.462-4.699) | 0.001 |
| Tumor multiplicity (Signal vs. Multiple) | 2.032(1.171-3.524) | 0.012 |
| Differentiation (Well-moderate vs. Poor-undifferentiated) | 1.165(0.707-1.920) | 0.548 |
| Exosome miR-21(Low expression vs. High expression) | 4.327(2.475-7.564) | 0.000 |
| Exosome miR-10b (Low expression vs. High expression) | 4.535(2.589-7.944) | 0.000 |
| Multivariable analysis | ||
| AFP (≤20 vs. >20) | 1.475(0.771-2.823) | 0.240 |
| Tumor size (≤3cm vs. >3cm) | 1.841(0.988-3.430) | 0.055 |
| Tumor multiplicity (Signal vs. Multiple) | 1.620(0.898-2.921) | 0.109 |
| Exosome miR-21(Low expression vs. High expression) | 2.446(1.252-4.777) | 0.009 |
| Exosome miR-10b (Low expression vs. High expression) | 2.550(1.301-4.997) | 0.006 |
Discussion
In this study, we found that the acidic tumor microenvironment correlated with poor survival of HCC patients. Our results showed that acidic microenvironment increased HCC cell-derived exosomal miR-21 and miR-10b levels, which could promote migration and invasion of recipient HCC cells cultured under normal conditions both in vivo and in vitro. Also, we found that the levels of miR-21 and miR-10b in exosomes isolated from the sera of patients correlated with advanced tumor stage. Furthermore, exosomal miR-21 and miR-10b were independent prognostic factors for poor disease-free survival of patients with E-HCC and could serve as potential therapeutic targets. To the best of our knowledge, this is the first study illustrating that acidic tumor microenvironment can promote HCC cell proliferation and/or metastasis by delivering exosomal miR-21 and miR-10b.
Exosomes have been shown to participate in tumor progression, but the association between exosomes and acidic microenvironment, especially in E-HCC, had not been described. Here, by using the TCGA dataset, we showed that expression of pH-related genes was associated with HCC patient survival. We further found that low pH culture conditions could dramatically induce the release of exosomes in HCC cell lines. These acidic pH-derived exosomes substantially promoted proliferation, migration, and invasion of recipient tumor cells cultured in the normal non-acidic medium. Our results suggested that HCC cells can easily be induced to a pro-metastatic phenotype by acidic pH-derived exosomes.
It is known that exosomes function as a bridge between cells and the microenvironment by transferring biological molecules, especially mRNAs and miRNAs [39]. In the present study, we focused on exosomal miRNAs and their effects on tumor proliferation and metastasis. Using miRNA microarrays and a 3-fold cutoff, we identified 31 upregulated and 50 downregulated miRNAs in exosomes cultured in acidic-condition compared with those cultured in normal condition. The results also indicated that the packaging of exosomal miRNAs may be affected by different culture conditions, especially by acidic pH. We further compared the differential miRNAs in patient's tissues by exploring TCGA dataset. LASSO logistic regression analysis was used to shrink miRNA candidates [40]. Based upon the results from the LASSO logistic model combined with our microarray analysis, we hypothesized that the acidic exosomal miRNA-21 and miR-10b are remarkably correlated with tumor progression.
miR-21 and miR-10b are associated with cell proliferation, apoptosis, and invasion by targeting tumor suppressor genes in many types of human cancers [41, 42]. It has been reported that both miR-21 and miR-10b contained an HRE region and could be induced by HIF-1α and HIF-2α and by pH-dependent nucleolar sequestration of von Hippel-Lindau (VHL) protein [13, 43, 44]. We, therefore, further investigated the functions of exosomal miR-21 and miR-10b in modulating HCC progression and found that acidic microenvironment could clearly induce the expression of exosomal miR-21 and miR-10b in HCC in a HIF-1α and HIF-2α-dependent manner. Knockdown of miR-21 and miR-10b resulting in reduced levels in exosomes substantially decreased HCC cell proliferation, migration, and/or invasion, whereas restoration of miR-21 and miR-10b by knocking-down HIF-1α and HIF-2α- largely rescued the proliferation, migration, and invasion of HCC cells both in vivo and in vitro. Furthermore, we found that acidic pH- derived exosomes containing miR-21 and miR-10b strongly induced a pool of genes in recipient cells. Taken together, these results suggested that exosomal miR-21 and miR-10b induced by tumor acidic microenvironment play critical roles in promoting HCC cell growth and metastasis.
Given the importance of exosomal miR-21 and miR-10b in HCC development and/or aggressiveness, we investigated whether exosomal miR-21 and miR- 10b from the serum samples of E-HCC patients are biologically active. Our results demonstrated that the exosomes derived from E-HCC patients' sera had elevated levels of miR-21 and/or miR-10b, which could substantially promote HCC cell proliferation and migration. We then further investigated whether serum exosomal miR-21 and miR-10b could serve as prognostic biomarkers and useful therapeutic targets for metastasis prevention in E-HCC patients. We found that the levels of serum exosomal miR-21 and miR-10b in E-HCC patients were positively associated with E-HCC advanced clinical stage as well as HIF-1α and HIF-2α expression in the tumor. More importantly, we demonstrated that a nano-drug designed to target miR-21 and miR-10b had therapeutic effects in inhibiting HCC in a xenograft mouse model. Thus, these results provide evidence that serum exosomal miR-21 and/or miR-10b may serve as prognostic biomarkers and potential therapeutic targets for E-HCC.
In summary, our results clearly showed that upregulation of exosomal miR-21 and/or miR-10b by the acidic microenvironment is a frequent oncogenic event in HCC critical for the acquisition of an aggressive/poor prognostic phenotype of the disease. Functional and mechanistic results demonstrated that exosomal miR-21 and miR-10b can drive the recipient cells towards a pro-metastatic phenotype. More importantly, our findings indicated that exosomal miR-21 and miR-10b may be novel therapeutic targets for E-HCC and potentially broaden the treatment options for this highly prevalent disease.
Abbreviations
E-HCC: early-stage hepatocellular carcinoma; pHi: intracellular pH; pHe: extracellular pH; GLUT-1: glucose transporter 1; NHE-1: sodium-hydrogen exchanger-1; CA-IX: carbonic anhydrase IX; VHL: von Hippel-Lindau; HIFs: hypoxia-inducible factors; MMP-3: matrix mellapproteinase-3; MMP-9: matrix mellapproteinase-9; POPC: 1-palmitoyl-2-oleoyl-snglycero-3-phosphocholine; DOPE: 1,2-dioleoyl-snglycero-3-phosphoethanolamine; CHEMS: cholesterylhemisuccinate; MOCHOL: cholesteryl-4-((2-(4-morpholinyl)ethyl)amino)-4-oxoburanoate; TACE: transarterial chemoembolization; LASSO: least absolute shrinkage and selection operator; DFS: Disease-free survival; OS: Overall survival. SYSUCC, Sun Yat-Sen University Cancer Center; PDCM: the nano-liposomes consisted by 1-palmitoyl-2-oleoyl-snglycero-3-phosphocholine; PDCM-GLUT-1: GLUT-1 siRNA encapsulated in PDCM nano-particles; PDCM-CA9: CA9 siRNA is encapsulated in PDCM nano-particles.
Supplementary Material
Supplementary figures and tables.
Acknowledgements
We thank the TCGA research network for providing the data analyzed in this manuscript. We are grateful to Wenjun He (Department of Medical Statistics and Epidemiology, School of Public Health, Sun Yat-sen University, Guangzhou, China) for supporting part of the data extraction and processing.
This work was supported by grants from the National Key R&D Program of China (2017YFC1309001, 2016YFC1302305), the National Natural Science Foundation of China (81672686, 81730072); Natural Science Foundation of Guangdong Province, China (2015A030313020); Sister Institution Network Fund of the MD Anderson Cancer Center.
Ethics Committee Approval and Patient Consent
Approval was obtained for all animal studies under the guidelines of Sun Yat-sen University Cancer Center. The clinical study was approved by the Ethics Review Board of Sun Yat-sen University Cancer Center, and written informed consent was obtained from all subjects.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Maluccio M, Covey A. Recent progress in understanding, diagnosing, and treating hepatocellular carcinoma. CA Cancer J Clin. 2012;62:394-9
2. Ueno M, Hayami S, Shigekawa Y, Kawai M, Hirono S, Okada K. et al. Prognostic impact of surgery and radiofrequency ablation on single nodular HCC ≤ 5 cm: Cohort study based on serum HCC markers. J Hepatol. 2015;63:1352-9
3. Tabrizian P, Jibara G, Shrager B, Schwartz M, Roayaie S. Recurrence of hepatocellular cancer after resection: patterns, treatments, and prognosis. Ann Surg. 2015;261:947-55
4. Forner A, Reig M, Bruix J. Hepatocellular carcinoma. Lancet. 2018;391:1301-14
5. Wang H, Chen L. Tumor microenviroment and hepatocellular carcinoma metastasis. J Gastroenterol Hepatol. 2013;28(Suppl 1):43-8
6. Webb BA, Chimenti M, Jacobson MP, Barber DL. Dysregulated pH: a perfect storm for cancer progression. Nat Rev Cancer. 2011;11:671-7
7. Amann T, Hellerbrand C. GLUT1 as a therapeutic target in hepatocellular carcinoma. Expert Opin Ther Targets. 2009;13:1411-27
8. Stock C, Schwab A. Protons make tumor cells move like clockwork. Pflugers Arch. 2009;458:981-92
9. Gatenby RA, Gawlinski ET, Gmitro AF, Kaylor B, Gillies RJ. Acid-mediated tumor invasion: a multidisciplinary study. Cancer Res. 2006;66:5216-23
10. Robey IF, Baggett BK, Kirkpatrick ND, Roe DJ, Dosescu J, Sloane BF. et al. Bicarbonate increases tumor pH and inhibits spontaneous metastases. Cancer Res. 2009;69:2260-8
11. Fais S, Venturi G, Gatenby B. Microenvironmental acidosis in carcinogenesis and metastases: new strategies in prevention and therapy. Cancer Metastasis Rev. 2014;33:1095-108
12. Barar J, Omidi Y. Dysregulated pH in Tumor Microenvironment Checkmates Cancer Therapy. Bioimpacts. 2013;3:149-62
13. Mekhail K, Gunaratnam L, Bonicalzi ME, Lee S. HIF activation by pH-dependent nucleolar sequestration of VHL. Nat Cell Biol. 2004;6:642-7
14. Zhao J, Du F, Shen G, Zheng F, Xu B. The role of hypoxia-inducible factor-2 in digestive system cancers. Cell Death Dis. 2015;6:e1600
15. Yang Y, Sun M, Wang L, Jiao B. HIFs, angiogenesis, and cancer. J Cell Biochem. 2013;114:967-74
16. Panagiotou N, Neytchev O, Selman C, Shiels PG. Extracellular Vesicles, Ageing, and Therapeutic Interventions. Cells. 2018:7
17. Shao C, Yang F, Miao S, Liu W, Wang C, Shu Y. et al. Role of hypoxia-induced exosomes in tumor biology. Mol Cancer. 2018;17:120
18. Greening DW, Gopal SK, Xu R, Simpson RJ, Chen W. Exosomes and their roles in immune regulation and cancer. Semin Cell Dev Biol. 2015;40:72-81
19. Garcia-Murillas I, Schiavon G, Weigelt B, Ng C, Hrebien S, Cutts RJ. et al. Mutation tracking in circulating tumor DNA predicts relapse in early breast cancer. Sci Transl Med. 2015;7:302ra133
20. Ban JJ, Lee M, Im W, Kim M. Low pH increases the yield of exosome isolation. Biochem Biophys Res Commun. 2015;461:76-9
21. Wojtkowiak JW, Rothberg JM, Kumar V, Schramm KJ, Haller E, Proemsey JB. et al. Chronic autophagy is a cellular adaptation to tumor acidic pH microenvironments. Cancer Res. 2012;72:3938-47
22. Li L, Li C, Wang S, Wang Z, Jiang J, Wang W. et al. Exosomes Derived from Hypoxic Oral Squamous Cell Carcinoma Cells Deliver miR-21 to Normoxic Cells to Elicit a Prometastatic Phenotype. Cancer Res. 2016;76:1770-80
23. Qiu J, Peng B, Tang Y, Qian Y, Guo P, Li M. et al. CpG Methylation Signature Predicts Recurrence in Early-Stage Hepatocellular Carcinoma: Results From a Multicenter Study. J Clin Oncol. 2017;35:734-42
24. Wei JX, Lv LH, Wan YL, Cao Y, Li GL, Lin HM. et al. Vps4A functions as a tumor suppressor by regulating the secretion and uptake of exosomal microRNAs in human hepatoma cells. Hepatology. 2015;61:1284-94
25. Tian XP, Jin XH, Li M, Huang WJ, Xie D, Zhang JX. The depletion of PinX1 involved in the tumorigenesis of non-small cell lung cancer promotes cell proliferation via p15/cyclin D1 pathway. Mol Cancer. 2017;16:74
26. Daige CL, Wiggins JF, Priddy L, Nelligan-Davis T, Zhao J, Brown D. Systemic delivery of a miR34a mimic as a potential therapeutic for liver cancer. Mol Cancer Ther. 2014;13:2352-60
27. Tolcher AW, Rodrigueza WV, Rasco DW, Patnaik A, Papadopoulos KP, Amaya A. et al. A phase 1 study of the BCL2-targeted deoxyribonucleic acid inhibitor (DNAi) PNT2258 in patients with advanced solid tumors. Cancer Chemother Pharmacol. 2014;73:363-71
28. Beg MS, Brenner AJ, Sachdev J, Borad M, Kang YK, Stoudemire J. et al. Phase I study of MRX34, a liposomal miR-34a mimic, administered twice weekly in patients with advanced solid tumors. Invest New Drugs. 2017;35:180-8
29. Zheng F, Liao YJ, Cai MY, Liu YH, Liu TH, Chen SP. et al. The putative tumour suppressor microRNA-124 modulates hepatocellular carcinoma cell aggressiveness by repressing ROCK2 and EZH2. Gut. 2012;61:278-89
30. Gui J, Li H. Penalized Cox regression analysis in the high-dimensional and low-sample size settings, with applications to microarray gene expression data. Bioinformatics. 2005;21:3001-8
31. Goeman JJ. L1 penalized estimation in the Cox proportional hazards model. Biom J. 2010;52:70-84
32. Johnson LL, Pavlovsky AG, Johnson AR, Janowicz JA, Man CF, Ortwine DF. et al. A rationalization of the acidic pH dependence for stromelysin-1 (Matrix metalloproteinase-3) catalysis and inhibition. J Biol Chem. 2000;275:11026-33
33. Wilhelm SM, Shao ZH, Housley TJ, Seperack PK, Baumann AP, Gunja-Smith Z. et al. Matrix metalloproteinase-3 (stromelysin-1). Identification as the cartilage acid metalloprotease and effect of pH on catalytic properties and calcium affinity. J Biol Chem. 1993;268:21906-13
34. Putney LK, Barber DL. Expression profile of genes regulated by activity of the Na-H exchanger NHE1. BMC Genomics. 2004;5:46
35. Bourguignon LY, Singleton PA, Diedrich F, Stern R, Gilad E. CD44 interaction with Na+-H+ exchanger (NHE1) creates acidic microenvironments leading to hyaluronidase-2 and cathepsin B activation and breast tumor cell invasion. J Biol Chem. 2004;279:26991-7007
36. Chen LQ, Howison CM, Spier C, Stopeck AT, Malm SW, Pagel MD. et al. Assessment of carbonic anhydrase IX expression and extracellular pH in B-cell lymphoma cell line models. Leuk Lymphoma. 2015;56:1432-9
37. Estrella V, Chen T, Lloyd M, Wojtkowiak J, Cornnell HH, Ibrahim-Hashim A. et al. Acidity generated by the tumor microenvironment drives local invasion. Cancer Res. 2013;73:1524-35
38. Hoshino A, Costa-Silva B, Shen TL, Rodrigues G, Hashimoto A, Tesic Mark M. et al. Tumour exosome integrins determine organotropic metastasis. Nature. 2015;527:329-35
39. Brinton LT, Sloane HS, Kester M, Kelly KA. Formation and role of exosomes in cancer. Cell Mol Life Sci. 2015;72:659-71
40. Sun H, Wang S. Penalized logistic regression for high-dimensional DNA methylation data with case-control studies. Bioinformatics. 2012;28:1368-75
41. Pfeffer SR, Yang CH, Pfeffer LM. The Role of miR-21 in Cancer. Drug Dev Res. 2015;76:270-7
42. Heidary MF, Mahmoodzadeh Hosseini H, Mehdizadeh Aghdam E, Nourani MR, Ranjbar R, Mirnejad R. et al. Overexpression of Metastatic Related MicroRNAs, Mir-335 and Mir-10b, by Staphylococcal Enterotoxin B in the Metastatic Breast Cancer Cell Line. Adv Pharm Bull. 2015;5:255-9
43. Jiang S, Wang R, Yan H, Jin L, Dou X, Chen D. MicroRNA-21 modulates radiation resistance through upregulation of hypoxia-inducible factor-1alpha-promoted glycolysis in non-small cell lung cancer cells. Mol Med Rep. 2016;13:4101-7
44. Haque I, Banerjee S, Mehta S, De A, Majumder M, Mayo MS. et al. Cysteine-rich 61-connective tissue growth factor-nephroblastoma- overexpressed 5 (CCN5)/Wnt-1-induced signaling protein-2 (WISP-2) regulates microRNA-10b via hypoxia-inducible factor-1alpha-TWIST signaling networks in human breast cancer cells. J Biol Chem. 2011;286:43475-85
Author contact
Corresponding authors: xiedanorg.cn (Dan Xie), caiqqorg.cn (Qing-Qing Cai) and xurhorg.cn (Rui-Hua Xu).
 Global reach, higher impact
Global reach, higher impact