Impact Factor
Theranostics 2019; 9(7):2056-2070. doi:10.7150/thno.28119 This issue Cite
Research Paper
Non-invasive diagnosis of early-stage lung cancer using high-throughput targeted DNA methylation sequencing of circulating tumor DNA (ctDNA)
1. First Affiliated Hospital of Guangzhou Department of Thoracic Surgery/Oncology, the First Affiliated Hospital of Guangzhou Medical University; Guangzhou Institute of Respiratory Disease & Health; China State Key Laboratory and National Clinical Research Center for Respiratory Disease, Guangzhou, China;
2. AnchorDx Medical Co., Ltd., Guangzhou, China;
3. Tongji Hospital, Tongji Medical College Huazhong University of Science & Technology, Wuhan, China;
4. Van Andel Research Institute (VARI), Grand Rapids, MI, USA;
5. Department of Pathology, School of Basic Medical Science, Southern Medical University, Guangzhou, China.
*These authors contributed equally to the paper.
Abstract
Rational: LDCT screening can identify early-stage lung cancers yet introduces excessive false positives and it remains a great challenge to differentiate malignant tumors from benign solitary pulmonary nodules, which calls for better non-invasive diagnostic tools.
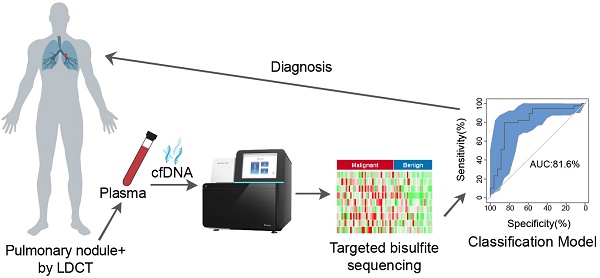
Methods: We performed DNA methylation profiling by high throughput DNA bisulfite sequencing in tissue samples (nodule size < 3 cm in diameter) to learn methylation patterns that differentiate cancerous tumors from benign lesions. Then we filtered out methylation patterns exhibiting high background in circulating tumor DNA (ctDNA) and built an assay for plasma sample classification.
Results: We first performed methylation profiling of 230 tissue samples to learn cancer-specific methylation patterns which achieved a sensitivity of 92.7% (88.3% - 97.1%) and a specificity of 92.8% (89.3% - 96.3%). These tissue-derived DNA methylation markers were further filtered using a training set of 66 plasma samples and 9 markers were selected to build a diagnostic prediction model. From an independent validation set of additional 66 plasma samples, this model obtained a sensitivity of 79.5% (63.5% - 90.7%) and a specificity of 85.2% (66.3% - 95.8%) for differentiating patients with malignant tumor (n = 39) from patients with benign lesions (n = 27). Additionally, when tested on gender and age matched asymptomatic normal individuals (n = 118), our model achieved a specificity of 93.2% (89.0% - 98.3%). Specifically, our assay is highly sensitive towards early‐stage lung cancer, with a sensitivity of 75.0% (55.0%-90.0%) in 20 stage Ia lung cancer patients and 85.7% (57.1%-100.0%) in 7 stage Ib lung cancer patients.
Conclusions: We have developed a novel sensitive blood based non‐invasive diagnostic assay for detecting early stage lung cancer as well as differentiating lung cancers from benign pulmonary nodules.
Keywords: Early-stage lung cancer, circulating tumor DNA, high-throughput targeted DNA methylation sequencing
 Global reach, higher impact
Global reach, higher impact