13.3
Impact Factor
Theranostics 2019; 9(12):3485-3500. doi:10.7150/thno.32033 This issue Cite
Research Paper
Molecularly annotation of mouse avatar models derived from patients with colorectal cancer liver metastasis
1. Key laboratory of Carcinogenesis and Translational Research (Ministry of Education/Beijing), Department of Gastrointestinal Oncology, Peking University Cancer Hospital and Institute, 52 Fucheng Road, Haidian District, Beijing 100142, China.
2. Key Laboratory of Carcinogenesis and Translational Research (Ministry of Education), Hepatopancreatobiliary Surgery Department I, Peking University Cancer Hospital and Institute, 52 Fucheng Road, Haidian District, Beijing 100142, China.
3. Department of Pathology, Key Laboratory of Carcinogenesis and Translational Research (Ministry of Education/Beijing), Peking University Cancer Hospital and Institute, 52 Fucheng Road, Haidian District, Beijing 100142, China.
*These authors contributed equally to this work.
Abstract
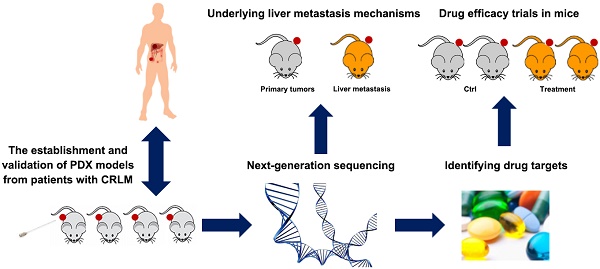
Background: Liver is the most common metastatic site in advanced colorectal cancer. Most patients with colorectal cancer liver metastasis (CRLM) do not benefit from current treatment. Patient-derived xenografts (PDXs) with defined molecular signatures are attractive models for preclinical studies.
Methods: Successfully established PDXs were evaluated to elucidate their fidelity of patients' biologic characteristics (pathologic, genetic and protein properties, together with chemosensitivity). The genomic variations of PDXs were analyzed by next-generation sequencing to explore the underlying molecular mechanism of metastasis and potential therapeutic targets.
Results: CRLM (N=73) showed a significantly higher successful PDX establishment rate than primary specimens (N=26; 76.7% vs. 57.7%). CRLM PDXs recapitulated the pathologic, genetic and protein properties of parental tumors, as well as chemosensitivity. Frequent altered genes in PDXs showed high consistency compared to patients' genomic alterations and were enriched in MAPK, ErbB, cell cycle, focal adhesion pathways for CRLM PDXs, whereas primary tumor-derived PDXs only exhibited genomic variations involving ErbB and cell cycle. The genetic alterations showed high concordance between paired PDXs from primary and metastatic tissues, except for recurrent gene mutations (ARID1A, CDK8, ETV1, STAT5B and WNK3) and common copy number gains in chromosomes 20q (e.g., SRC/AURKA). Several potential drug targets such as KRAS, HER2, and FGFR2 were validated using corresponding inhibitors. Additionally, PDX models could also be used in screening efficient regimens for patients with no druggable alterations.
Conclusion: This study has successfully established and validated a large panel of molecularly annotated platforms from patients with CRLM for preclinical studies.
Keywords: Patient-derived xenograft, colorectal cancer liver metastasis, molecular signature, therapeutic target
 Global reach, higher impact
Global reach, higher impact