Impact Factor
Theranostics 2019; 9(21):6175-6190. doi:10.7150/thno.35572 This issue Cite
Research Paper
DNA methylation-regulated QPCT promotes sunitinib resistance by increasing HRAS stability in renal cell carcinoma
1. Department of Urology, Changzheng Hospital, Second Military Medical University, Shanghai 200003, China.
2. Department of Medical Genetics, Second Military Medical University, Shanghai 200433, China.
3. Shanghai Key Laboratory of Cell Engineering, Second Military Medical University, Shanghai 200433, China.
* These authors contributed equally to this work.
Abstract
Rationale: Although sunitinib has been shown to improve the survival rate of advanced renal cell carcinoma (RCC) patients, poor drug response is a major challenge that reduces patient benefit. It is important to elucidate the underlying mechanism so that the therapeutic response to sunitinib can be restored.
Methods: We used an Illumina HumanMethylation 850K microarray to find methylation-differentiated CpG sites between sunitinib-nonresponsive and -responsive RCC tissues and Sequenom MassARRAY methylation analysis to verify the methylation chip results. We verified glutaminyl peptide cyclotransferase (QPCT) expression in sunitinib-nonresponsive and -responsive RCC tissues via qRT-PCR, western blot and immunohistochemical assays. Then, cell counting kit 8 (CCK-8), plate colony formation and flow cytometric assays were used to verify the function of QPCT in RCC sunitinib resistance after QPCT intervention or overexpression. Chromatin immunoprecipitation (ChIP) was performed to clarify the upstream regulatory mechanism of QPCT. A human proteome microarray assay was used to identify downstream proteins that interact with QPCT, and co-immunoprecipitation (co-IP) and confocal laser microscopy were used to verify the protein chip results.
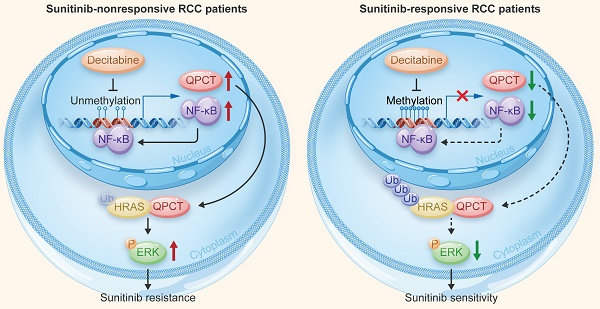
Results: We found that the degree of methylation in the QPCT promoter region was significantly different between sunitinib-nonresponsive and -responsive RCC tissues. In the sunitinib-nonresponsive tissues, the degree of methylation in the QPCT promoter region was significantly reduced, and the expression of QPCT was upregulated, which correlated with a clinically poor response to sunitinib. A knockdown of QPCT conferred sunitinib sensitivity traits to RCC cells, whereas an overexpression of QPCT restored sunitinib resistance in RCC cells. Mechanistically, reducing the methylation degree of the QPCT promoter region by 5-aza-2'-deoxycytidine (decitabine) in RCC cells could increase the expression of QPCT and NF-κB (p65) bound to the QPCT promoter region, positively regulating its expression, while the hypermethylation in the QPCT promoter region could inhibit the binding of NF-κB (p65). QPCT could bind to HRAS and attenuate the ubiquitination of HRAS, thus increasing its stability and leading to the activation of the ERK pathway in RCC cells.
Conclusion: QPCT may be a novel predictor of the response to sunitinib therapy in RCC patients and a potential therapeutic target.
Keywords: Renal cell carcinoma, Sunitinib, Glutaminyl peptide cyclotransferase, DNA methylation, HRAS
 Global reach, higher impact
Global reach, higher impact