Impact Factor
Theranostics 2020; 10(2):484-497. doi:10.7150/thno.38060 This issue Cite
Research Paper
A tumor suppressor enhancing module orchestrated by GATA4 denotes a therapeutic opportunity for GATA4 deficient HCC patients
1. Key Laboratory of Functional Protein Research of Guangdong Higher Education, Institute of Life and Health Engineering, College of Life Science and Technology, Jinan University, Guangzhou 510632, China.
2. Beijing Institute of Genomics, Chinese Academy of Sciences, Beijing 100101, China
3. Beijing Ditan Hospital, Beijing 100015, China
4. Department of Hepatobiliary Surgery, Beijing Tsinghua Changgung Hospital, Beijing 100084, China
5. Zhuhai Precision Medical Center, Zhuhai People's Hospital (Zhuhai Hospital Affiliated with Jinan University), Jinan University, Zhuhai 519000, Guangdong, China
6. The Biomedical Translational Research Institute, Jinan University Faculty of Medical Science, Jinan University, Guangzhou 510632, Guangdong, China
7. Beijing Tongren Hospital, Capital Medical University, Beijing 100730, China
* These authors contributed equally to this work
Abstract
Rationale: Effective targeting therapies are limited in Hepatocellular carcinoma (HCC) clinic. Characterization of tumor suppressor genes (TSGs) and elucidation their signaling cascades could shed light on new strategies for developing targeting therapies for HCC.
Methods: We checked genome-wide DNA copy number variation (CNV) of HCC samples, focusing on deleted genes for TSG candidates. Clinical data, in vitro and in vivo data were collected to validate the tumor suppressor functions.
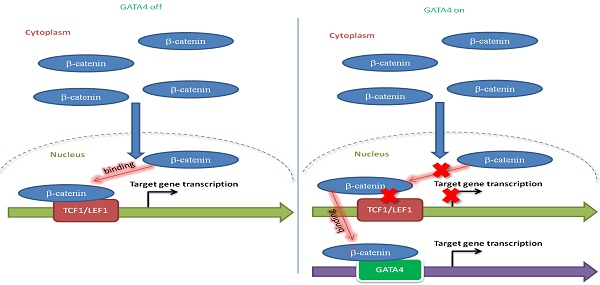
Results: Focal deletion of GATA4 gene locus was the most prominent feature across all liver cancer samples. Ectopic expression of GATA4 resulted in senescence of HCC cell lines. Mechanistically, GATA4 exerted tumor suppressive role by orchestrating the assembly of a tumor suppressor enhancing module: GATA4 directly bound and potently inhibited the mRNA transcription activity of β-catenin; meanwhile, β-catenin was recruited by GATA4 to promoter regions and facilitated transcription of GATA4 target genes, which were TSGs per se. Expression of GATA4 was effective to shrink GATA4-deficient HCC tumors in vivo. We also showed that β-catenin inhibitor was capable of shrinking GATA4-deficient tumors.
Conclusions: Our study unveiled a previously unnoticed tumor suppressor enhancing module assembled by ectopically expressed GATA4 in HCC cells and denoted a therapeutic opportunity for GATA4 deficient HCC patients. Our study also presented an interesting case that an oncogenic transcription factor conditionally functioned as a tumor suppressor when recruited by a TSG transcription factor.
Keywords: GATA4, Hepatocellular carcinoma, Tumor suppressor gene, targeting therapy
 Global reach, higher impact
Global reach, higher impact