Impact Factor
Theranostics 2020; 10(9):4168-4182. doi:10.7150/thno.43834 This issue Cite
Review
Targeting REV-ERBα for therapeutic purposes: promises and challenges
1. College of Pharmacy, Jinan University, Guangzhou, 510632, China
2. Integrated Chinese and Western Medicine Postdoctoral research station, Jinan University, Guangzhou, 510632, China
3. Guangzhou Jinan Biomedicine Research and Development Center, Jinan University, Guangzhou, 510632, China
4. International Cooperative Laboratory of Traditional Chinese Medicine Modernization and Innovative Drug Development of Chinese Ministry of Education (MOE), College of Pharmacy, Jinan University, Guangzhou, 510632, China
Received 2020-1-10; Accepted 2020-2-8; Published 2020-3-4
Abstract
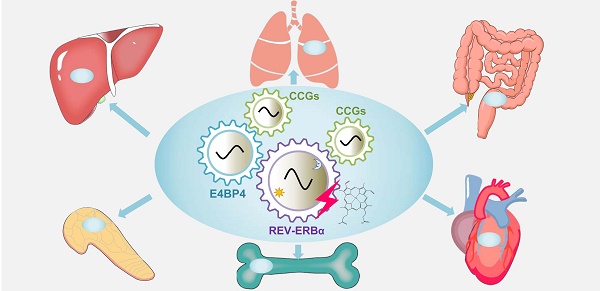
REV-ERBα (NR1D1) is a circadian clock component that functions as a transcriptional repressor. Due to its role in direct modulation of metabolic genes, REV-ERBα is regarded as an integrator of cell metabolism with circadian clock. Accordingly, REV-ERBα is first proposed as a drug target for treating sleep disorders and metabolic syndromes (e.g., dyslipidaemia, hyperglycaemia and obesity). Recent years of studies uncover a rather broad role of REV-ERBα in pathological conditions including local inflammatory diseases, heart failure and cancers. Moreover, REV-ERBα is involved in regulation of circadian drug metabolism that has implications in chronopharmacology. In the meantime, recent years have witnessed discovery of an array of new REV-ERBα ligands most of which have pharmacological activities in vivo. In this article, we review the regulatory role of REV-ERBα in various types of diseases and discuss the underlying mechanisms. We also describe the newly discovered ligands and the old ones together with their targeting potential. Despite well-established pharmacological effects of REV-ERBα ligands in animals (preclinical studies), no progress has been made regarding their translation to clinical trials. This implies certain challenges associated with drug development of REV-ERBα ligands. In particular, we discuss the potential challenges related to drug safety (or adverse effects) and bioavailability. For new drug development, it is advocated that REV-ERBα should be targeted to treat local diseases and a targeting drug should be locally distributed, avoiding the adverse effects on other tissues.
Introduction
REV-ERBα [also known as NR1D1 (nuclear receptor subfamily 1 group D member 1)] is a nuclear receptor and a core component of the molecular clock system. REV-ERBα was discovered in 1989 and its name was derived from genomic location on the reverse DNA strand of v-erbA oncogene (also called “thyroid hormone receptor α”) found in the avian erythroblastosis virus [1,2]. About five years later, REV-ERBβ (NR1D2), the other member of NR1D subfamily, was identified [3]. Due to the lack of an activation function 2 (AF2, a motif for recognition of co-activators) in ligand binding domain, REV-ERBα/β cannot activate gene transcription [4]. Instead, REV-ERBα/β function as transcriptional repressors, and inhibit gene transcription by recruiting co-repressors nuclear receptor co‑repressor 1 (NCOR1) and histone deacetylase 3 (HDAC3) [5]. REV-ERBα may play a more important role in regulating circadian rhythms as compared to its paralog REV-ERBβ. REV-ERBα-deficient mice show disrupted circadian rhythms characterized by a shortened period. However, impact of REV-ERBβ ablation on circadian rhythms is negligible [6].
Due to its role in direct modulation of clock and metabolic genes, REV-ERBα is first proposed as a drug target for treating sleep disorders and metabolic syndromes (e.g., dyslipidaemia, hyperglycaemia and obesity) in 2012 [7]. Recent years of studies uncover a rather broad role of REV-ERBα in pathological conditions including local inflammatory diseases, heart failure and cancers. Moreover, REV-ERBα is involved in regulation of circadian drug metabolism that has implications in chronopharmacology. In the meantime, recent years have witnessed discovery of an array of new REV-ERBα ligands most of which have pharmacological activities in vivo. In this article, we review the regulatory role of REV-ERBα in various types of diseases and discuss the underlying mechanisms. We also describe the newly discovered ligands and the old ones together with their targeting potential. In addition, the potential challenges associated with drug development of REV-ERBα ligands are discussed.
REV-ERBα in molecular clock system
REV-ERBα is a core component of circadian clock system in mammals. Mammalian molecular clock consists of three interlocked auto-regulatory feedback loops (Figure 1A) [8,9]. The main loop is driven by BMAL1 (brain and muscle ARNT-like 1)/CLOCK (circadian locomotor output cycles kaput) heterodimer that induces the expression of E-box-controlled genes (ECGs) including periods (PERs) and cryptochromes (CRYs). Once reaching a high level, PER and CRY proteins move from the cytoplasm to the nucleus, and inhibit BMAL1/CLOCK activity. When the levels of PER and CRY proteins are reduced due to protein degradation, PER and CRY are dissociated from the BMAL1 /CLOCK complex and a new cycle of transcription is started. Degradation of PER and CRY proteins are controlled by casein kinases (CKIδ and CKIɛ) and adenosine monophosphate kinase (AMPK), respectively. These kinases tag the proteins via phosphorylation for ubiquitination and proteasome degradation [10-12]. Alternatively, ubiquitination of CRYs can be mediated by FBXL3. However, FBXL21 forms an SCF E3 ligase complex to retain CRYs in the cytoplasm and protects CRYs from FBXL3-mediated degradation (Figure 1A) [13].
REV‑ERBα in circadian clock system. (A) Schematic diagram for molecular clock machinery. Mammalian molecular clock consists of three interlocked auto-regulatory feedback loops. The three loops are attained through PERs/CRYs (black lines), REV-ERBs/RORs (green lines) and DBP/E4BP4 (blue lines), respectively. (B) Circadian mRNA expression patterns of clock genes in mice. (C) General patterns for regulation of target genes by REV‑ERBα. REV‑ERBα directly regulates transcription of target genes via single RORE (i), RevDR2 (ii) or two adjacent ROREs (iii), and indirectly regulates gene transcription via other transcription factors (TFs) (iv).
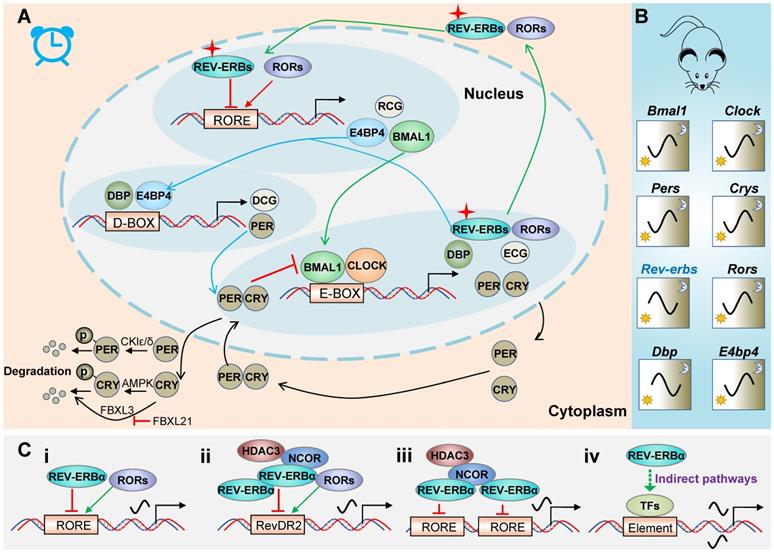
In the second loop (Figure 1A), BMAL1/CLOCK drives expression of REV-ERBs and RORs, which in turn respectively repress and activate BMAL1 transcription and RORE/RevRE-controlled genes (RCGs) (Table 1). RCGs include genes involved in immune responses, metabolic homeostasis, cancers, nervous and cardiovascular systems. The third loop (Figure 1A) involves DBP and E4BP4 that regulate PER2 (an output gene from the main loop) and D-box controlled genes (DCGs). All clock genes are cyclically expressed although the patterns differ (Figure 1B). Of note, REV-ERBα (in mice) oscillates with a maximum level (zenith) at ZT6-10 and a minimum level (nadir) at ZT18-22 (Figure 1B). A large portion of clock controlled genes (CCGs, including Bmal1 and E4bp4) are under the control of REV-ERBα (Table 1), and show characteristic patterns antiphase to REV-ERBα (Figure 1B).
Target genes of REV-ERBα
| Target genes | Type | Species | Refs |
|---|---|---|---|
| NPAS2 | RCGs | Human | [125] |
| CLOCK | RCGs | Human | [126] |
| E4bp4/Shp | RCGs | Mouse | [74] |
| IL-6 | RCGs | Mouse | [48] |
| IL-10 | RCGs | Human | [127] |
| IL-17a | RCGs | Mouse | [33,39] |
| Nlrp3/p65 | RCGs | Mouse | [25,27] |
| IL-1β | RCGs | Mouse | [27] |
| Ccl2 | RCGs | Mouse | [47] |
| TLR-4 | RCGs | Human | [45] |
| Mmp9/Cx3cr1 | RCGs | Mouse | [16] |
| PAI-1 | RCGs | Human | [128] |
| Pck1 | RCGs | Mouse | [58] |
| ApoC-III | RCGs | Human | [69] |
| Elovl3 | RCGs | Mouse | [72] |
| LRH-1 | RCGs | Mouse/Human | [76] |
| Fabp7 | RCGs | Mouse | [129] |
| βKlotho | RCGs | Mouse | [130] |
| Cyp2b10 | RCGs | Mouse | [101] |
| Ces2 | RCGs | Mouse | [98] |
| Cyp4a | RCGs | Mouse | [100] |
| Ugt2b | RCGs | Mouse | [99] |
| Cyclin A | RCGs | Mouse/Human | [88] |
| PFKFB3/G6PD | RCGs | Human | [89] |
| PGC1α | RCGs | Human | [131] |
| Bhmt/Cbs/Cth | RCGs | Mouse | [80] |
| Ucp1 | RCGs | Mouse | [132] |
| Fmo5 | DCGs | Mouse | [17] |
REV-ERBα generally functions as a monomer and binds to a consensus half-site motif (A/G)GGTCA preceded by an A/T rich 5' sequence (named RORE or RevRE) on target gene promoters (Figure 1C) [3,14]. In some cases, REV-ERBα can bind to direct repeats of RORE separated by 2 bp (RevDR2) as a dimer (Figure 1C). Moreover, two REV-ERBα molecules can separately bind to two adjacent ROREs and recruit co-repressors (i.e., NCOR1 and HDAC3) to regulate gene transcription (Figure 1C). Transcriptional repression mechanism of REV-ERBα may involve dynamic modulation of chromatin looping [15]. REV-ERBα also acts to suppress gene expression at a distance by repressing the transcription of enhancer-derived RNAs (eRNAs) [16]. In addition to direct regulation, REV-ERBα indirectly regulates gene transcription via repressing E4bp4 (Figure 1C) [17-19]. This is supported by the fact that REV-ERBα and E4bp4 share a large number of target genes [20]. REV-ERBα also indirectly regulates gene transcription by physically interacting with other transcription factors (e.g., HNF6, GR and NF-Y) (Figure 1C) [21-23].
REV-ERBα and diseases
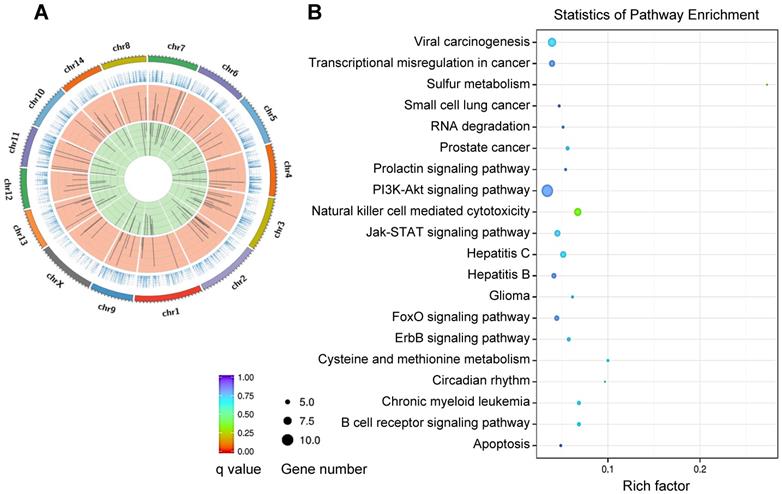
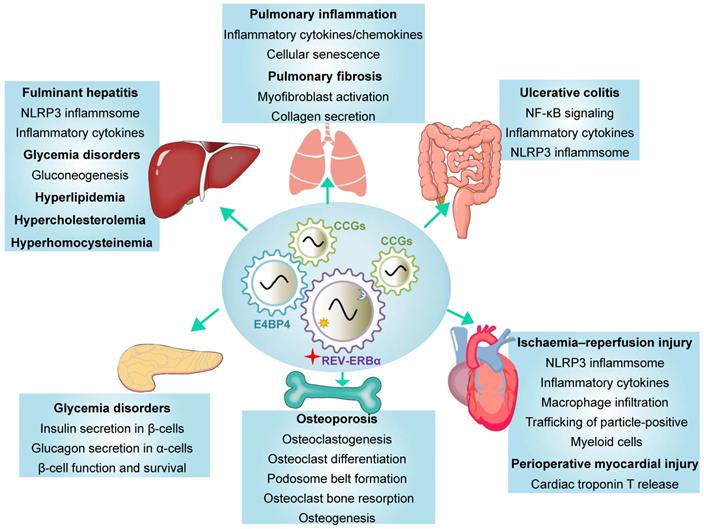
REV-ERBα has been implicated in regulation of a variety of diseases including inflammatory diseases, metabolic disorders and cancers (Table 2) [3,4]. REV-ERBα expression is often times altered (expression changed and rhythm disrupted) during disease development [24,25]. Reciprocally, dysregulation of REV-ERBα in humans and mice impacts the organism susceptibility to diseases [25-27]. REV-ERBα knockout elicits disturbance in genome-wide gene expression (Figure 2A). The differentially expressed genes are associated with pathways involved in various pathological processes and diseases (Figure 2B). There is accumulating experimental evidence supporting REV-ERBα as a therapeutic target for diseases in the liver, lung, colon, pancreas, heart and bone (Figure 3 & Table 2).
Phenotypes of REV-ERBα ablation and the underlying mechanisms
| Tissues | Phenotypes | Mechanisms | Refs |
|---|---|---|---|
| Liver | Fulminant hepatitis, IL-1β and IL-18 secretion | NLRP3 inflammasome, Transcription of Nlrp3 and IL-1β | [27] |
| Liver | Diabetes | Transcription of PCK and G6Pase | [6, 57-61] |
| Liver | Hyperlipidemia | Transcription of ApoC-III and Elovl3 | [6,69,70,72] |
| Liver | Hypercholesterolemia | Expression of HMGCR and CYP7A1 | [71,74-76] |
| Liver | Hyperhomocysteinemia | Transcription of Bhmt, Cbs, Cth and C/EBPα | [80] |
| Lung | Pulmonary inflammation | Expression of chemokines and inflammatory cytokines | [40] |
| Lung | Pulmonary fibrosis | TBPL1-integrinβ1 pathway | [56] |
| Colon | Ulcerative colitis | NF-κB signaling pathway, NLRP3 inflammasome, Transcription of Nlrp3 | [25] |
| Heart | Ischaemia-reperfusion injury, Immunocyte recruitment | NLRP3 inflammasome | [36] |
| Heart | Perioperative myocardial injury | Expression of CDKN1a/p21. | [30] |
| Bone | Osteoporosis, Osteoclastogenesis | Expression of FABP4 | [84] |
| Bone | Osteogenesis | Expression of bone sialoprotein | [85] |
| Pancreas | Glucose-induced insulin secretion in β-cells, Glucagon secretion in α-cells | Exocytotic process, AMPK/Nampt/Sirt1 pathway | [62-64] |
REV‑ERBα controlled genes and pathways. (A) Circos plot of differentially expressed genes between Rev-erbα-/- and wild-type mice, showing a disturbance in genome-wide gene expression. In the Circos plot, the outermost circle depicts the ideograms of each chromosome; the second circle represents gene expression levels; the third circle shows the distribution of the up-regulated genes; and the fourth circle shows the distribution of the down-regulated genes. (B) KEGG pathway analysis of Rev-erbα-induced differentially expressed genes in mouse liver (top 20 pathways are shown).
Functions of REV‑ERBα in various tissues. REV‑ERBα regulates clock-controlled genes (CCGs) to affect disease development in a tissue-specific manner. REV‑ERBα directly regulates target genes or indirectly regulates gene transcription via other transcription factors (e.g., E4BP4).
Role of REV-ERBα in inflammatory diseases
Inflammatory diseases often times exhibit time-varying severity or symptoms. For instance, patients with rheumatoid arthritis show diurnal variations in symptoms, as manifested by great joint pain, stiffness and functional disability in the morning [28,29]. Patients received aortic valve replacement in the afternoon show alleviated perioperative myocardial injury compared to individuals received aortic valve replacement at other times of the day [30]. Asthma is more severe in the early hours of the morning [31]. Diurnal rhythmicity in the severity of inflammatory diseases may be associated with circadian REV-ERBα, a negative regulator of rhythmic inflammatory factors. Mice display dramatic daily differences in their susceptibility to LPS/D-GalN-induced fulminant hepatitis, with a lowest survival time at ZT16 that corresponds to a low REV-ERBα expression [27]. Moreover, chronic colitis displays a diurnal rhythmicity in disease severity and its diurnal pattern is in an opposite phase to that of REV-ERBα [32]. REV-ERBα ablation abrogates the diurnal rhythms of REV-ERBα-related inflammatory factors [25,32].
Accumulating evidence supports that targeting REV-ERBα is a promising approach for management of inflammations. REV-ERBα activation is shown to ameliorate ulcerative colitis [25,32,33], fulminant hepatitis [27], neuroinflammation [34,35], heart failure [36,37], myocardial infarction [38], experimental autoimmune encephalomyelitis [33,39], and pulmonary inflammation [29,40]. Consistently, Rev-erbα-/- mice exhibit aggravated inflammations [25,27,33-40]. Contrasting with a general anti-inflammatory role of REV-ERBα, Montaigne et al uncover a detrimental role of REV-ERBα in ischaemia-reperfusion injury, an inflammation-related disease [30]. The authors show that REV-ERBα ablation or antagonism ameliorates ischaemia-reperfusion injury through promoting CDKN1a/p21 [30]. However, this study may not deny the anti-inflammatory effects of REV-ERBα because ischaemia-reperfusion injury is also determined by many other factors such as calcium overload, oxidative/nitrosative stress and endoplasmic reticulum stress in addition to inflammatory reactions [41].
The role of REV-ERBα in regulation of innate immune responses has been well established. REV-ERBα is involved in immune cell development, macrophage polarization, NF-κB signaling, transcription of inflammation-related genes (e.g., cytokine genes, chemokine genes and receptor genes) and activation of NLRP3 inflammasome. REV-ERBα impacts development of group 3 innate lymphoid cells (ILC3s) and secretion of related cytokines (i.e., IL-17 and IL-22) by controlling mitochondria [42]. Activation of REV-ERBα impairs pro-inflammatory M1 phenotype and enhances anti-inflammatory M2 phenotype [43]. REV-ERBα suppresses NF-κB signaling in human endometrial stroma cells and mouse macrophages/microglia cells, and down-regulates expressions of related genes, such as Nlrp3, IL-6, IL-1β, IL-18, Tnfα and Ccl2 [25,34,35,44]. In addition to an indirect regulation mechanism via NF-κB signaling, REV-ERBα directly regulates immune genes (e.g., Nlrp3, IL-1β, TLR4, IL-6, Ccl2, Mmp9 and Cx3cr1) [16,25,27,45-48] (Figure 4A). Furthermore, REV-ERBα down-regulates Nlrp3 inflammasome activity to prevent ulcerative colitis, peritoneal inflammation, fulminant hepatitis and heart failure in mice [25,27,32,36].
REV‑ERBα regulates immune genes in macrophages and TH17 cells. (A) In macrophages, a variety of immune genes (i.e., P65, Nlrp3, IL-1β, TLR4, IL-6, Ccl2, Mmp9 and Cx3cr1) are controlled by REV-ERBα. (B) REV-ERBα negatively regulates TH17 cell development by competing with RORgt at the RORE of Il-17a. An additional mechanism involves REV-ERBα-mediated repression of RORgt.
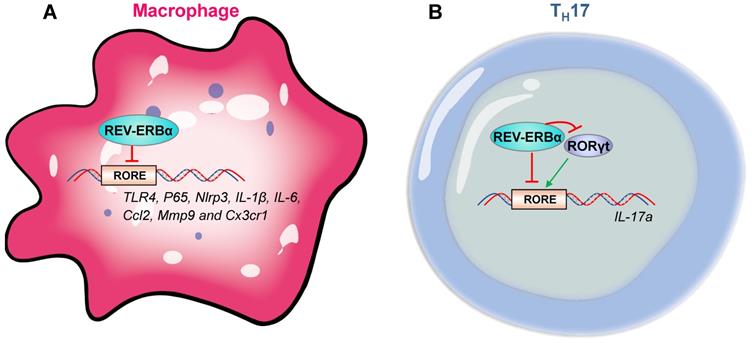
The adaptive immune responses is also under the control of REV-ERBα. Similar to innate immune cells, T and B cells exhibit strong circadian oscillations in the blood, peaking in the rest phase [49]. CD4+ and CD8+ T cells from murine lymph nodes exhibit a circadian rhythmicity in proliferation with a peak value in the evening [50]. Pro-inflammatory CD4+ T helper 17 (TH17) are the adaptive correlates of ILC3s based on shared developmental requirement for the master transcription factor RORγt and secretion of IL-17 and IL-22 [42,51]. TH17 drives inflammatory responses in many autoimmune diseases, and is a well-established cell model for studying regulation of immunity by circadian clock [19,52,53]. Effects of REV-ERBα on TH17 appear to be controversial. An early study reports that REV-ERBα drives TH17 cell differentiation and IL-17 production by repressing Nfil3 transcription [52]. Supporting this, Farez et al report that melatonin inhibits ROR-γt and ROR-α expression in TH17 cells by regulating REV-ERBα-Nfil3 axis [54]. However, later studies believe that REV-ERBα acts as a negative regulator of TH17 cell development by directly suppressing expression of IL-17 [33,39]. Chang et al proposed that the conflicting role of REV-ERBα in TH17 cells may be associated with its expression levels [39]. When expressed at a low level, REV-ERBα promotes RORγt expression via suppression of the negative regulator Nfil3 [52]. At a high level, REV-ERBα competes with RORγt for binding to the promoter of IL-17, inhibiting gene transcription (Figure 4B) [39]. Contrasting with an important role of REV-ERBα in TH17 cells, whether and how REV-ERBα regulates adaptive immunity in γδ T cells and regulatory T cells (with high REV-ERBα expressions) remain poorly explored [39].
Inflammation may lead to necrosis of parenchymal cells and promote the development of fibrosis. REV-ERBα agonist SR9009 alleviates CCl4-induced fibrosis in mice, as evidenced by reduced collagen deposition and decreased fibrotic gene expression [55]. Consistently, REV-ERBα suppresses the development of pulmonary fibrosis in mice in a recent study [56]. Lungs from Rev-erbα-/- mice reveal increased αSMA and collagen-1, two markers of myofibroblast activation. REV-ERBα agonist suppresses myofibroblast differentiation and collagen secretion in tissues from pulmonary fibrotic patients [56].
Role of REV-ERBα in metabolic disorders
Many metabolic genes exhibit significant circadian oscillations. Chronic disruption of circadian rhythms (e.g., by shift-work and sleep deprivation) have detrimental effects on cell metabolism, resulting in metabolic disorders such as diabetes, hyperlipidemia and obesity. There is accumulating evidence supporting a critical role of REV-ERBα in regulation of cell metabolism and metabolic diseases.
Glucose metabolism
REV-ERBα is implicated in glucose homeostasis and diabetes development due to its critical roles in regulation of glucose de novo synthesis and of pancreatic α/β-cell function. Activation of REV-ERBα reduces the levels of cellular and plasma glucose [7,57,58]. Consistently, REV-ERBα-deficient mice show an increased level of plasma glucose [6,59]. Yin et al demonstrate that REV-ERBα modulates glucose metabolism through regulating gluconeogenic rate-limiting enzymes phosphoenolpyruvate carboxykinase (PCK) and glucose‑6‑phosphatase (G6Pase) in human hepatoma cells and in primary mouse hepatocytes [57]. Accordingly, REV-ERBα can be targeted to alleviate glycemia disorders and diabetes [59-61]. In addition to the gluconeogenesis, REV-ERBα has a regulatory role in functions of pancreatic α and β-cells. At high glucose concentrations, REV-ERBα regulates glucose-induced insulin secretion in β-cells probably via modulation of the exocytotic process [62,63]. At low glucose levels, REV-ERBα promotes glucagon secretion in pancreatic α-cells through AMPK/Nampt/Sirt1 pathway [63,64]. Moreover, REV-ERBα enhances the survival and activity of β-cells under diabetogenic conditions [65].
Intracellular glucose levels oscillated in a circadian manner [66]. REV-ERBα has been implicated in regulation of glucose rhythm. Up-regulation of REV-ERBα by MYC leads to reduced level of Bmal1 and loss of circadian glucose metabolism [66]. CDK1-FBXW7 promotes REV-ERBα degradation in mouse liver, disrupting the circadian rhythmicity in glucose homeostasis [67]. Dietary iron modulates heme synthesis and REV-ERBα activity, thereby altering the circadian rhythm of hepatic gluconeogenesis [68].
Lipid metabolism
REV-ERBα-deficient mice exhibit a defect in lipid metabolism, causing increases in liver triglyceride and free fatty acids [6,69,70]. Activation of REV-ERBα results in reduced triglyceride and free fatty acids in mice [7,71]. The lipid-lowering effect is associated with transcriptional repression of ApoC-III (playing a key role in triglyceride metabolism by preventing catabolism of triglyceride-rich particles) and Elovl3 (elongation of fatty acids to produce very long-chain fatty acids) [69,72]. Regulation of lipogenic genes by REV-ERBα may require tethering factors such as HNF6 [73].
Cholesterol level mainly depends on the biosynthesis and elimination process. HMGCR (3-hydroxy-3-methylglutaryl-CoA reductase) and Cyp7A1 (cholesterol 7α-hydroxylase) are the rate limiting enzymes for cholesterol biosynthesis and catabolism, respectively. REV-ERBα was initially shown to regulate cholesterol catabolism (or biosynthesis of bile acids) and hypercholesteremia via a positive control of Cyp7a1 [74,75]. However, consensus was not reached regarding the mechanisms of action. REV-ERBα may regulate Cyp7a1 through E4bp4/Shp or Insig2/Srebp. By contrast, a recent study demonstrates that REV-ERBα inhibits Cyp7a1 expression via repressing Lrh-1 (an activator of Cyp7a1) that is supported by an early study [7,76]. Additionally, the effects of REV-ERBα on cholesterologenesis may also involve modification of cholesterol biosynthesis-related genes such as Hmgcr [71].
The relationships between REV-ERBα polymorphisms and predisposition to obesity have been also recognized. The REV-ERBα rs2071427 polymorphism modulates body fat mass in both adult and adolescent people [26]. Another polymorphism rs2314339 (in the intron of REV-ERBα) was associated with obesity in two cohorts from Mediterranean and North American population [77]. Recently, REV-ERBα polymorphism rs939347 is shown to modulate body fat mass in men, suggesting a gender-specific role of REV-ERBα in the development of obesity [78].
Amino acid metabolism
Homocysteine (a sulfur-containing amino acid) metabolism proceeds through two major pathways: remethylation to methionine and a two-step transsulfuration to cysteine [79]. REV-ERBα plays a crucial role in homocysteine metabolism and ammonia clearance [80]. Mechanistically, REV-ERBα regulates homocysteine catabolism through direct trans-repression of Bhmt, Cbs, and Cth, and ammonia clearance through inhibition of C/EBPα (CCAAT/enhancer-binding protein α) transactivation of Arg1, Cps1, and Otc [80]. It was proposed that targeting REV-ERBα represents a new approach in management of homocysteine- and ammonia-related diseases [80].
Bone metabolism
Disruption of circadian rhythms is associated with osteoporosis and abnormal bone metabolism, suggesting a close association between circadian clock and bone metabolism [81,82]. REV-ERBα is periodically expressed in murine calvarial bones [83]. Activation of REV-ERBα suppresses RANKL-induced podosome belt formation and inhibits osteoclast bone resorption, thereby ameliorating ovariotomy-induced bone loss [84]. Further, REV-ERBα regulates osteoclastogenesis via inducing FABP4 [84]. In addition, REV-ERBα inhibits osteogenesis by repressing the expression of bone sialoprotein in bone mesenchymal stem cells [85]. Therefore, REV-ERBα plays a pivotal role in maintaining metabolic homeostasis of bone by regulating osteoclastogenesis and osteogenesis.
Role of REV-ERBα in cancers
REV‑ERBα has been implicated in development and progression of gastric cancer [86]. It is associated with clinicopathological factors including poor differentiation, T stage, TMN stage and lymph node metastasis in human gastric cancer [86]. Patients with low REV‑ERBα expression exhibit poor prognosis compared with patients with high REV‑ERBα expression, indicating REV‑ERBα as a prognosis factor for gastric cancer [86]. REV‑ERBα activation induces apoptosis in human gastric cancer cells and in 3T3-L1 preadipocytes [86,87].
Anti-proliferative effects of REV-ERBα have been observed in human breast and gastric cancer cells [88,89]. Activation of REV-ERBα suppresses proliferation of breast cancer cells regardless of ER or HER2 status. REV-ERBα appears to pause the cell cycle of the breast cancer cells prior to M phase through direct targeting of cyclin A2 [88]. By contrast, anti-proliferative effects of REV-ERBα are attained through inhibiting glycolytic flux and pentose phosphate pathway in another study [89]. To be specific, REV-ERBα inhibits the expression of PFKFB3 and G6PD (two genes involved in glycolysis and pentose phosphate pathway), thereby interfering with glycolytic flux and pentose phosphate pathway [89].
Sulli et al proposed that pharmacological targeting of REV-ERBα is a promising strategy for cancer treatment [90]. The anticancer activity of SR9009 and SR9011 (two REV-ERBα agonists) affects a number of oncogenic drivers (such as HRAS, BRAF and PIK3CA) and persists in the absence of p53 and under hypoxic conditions [90]. Activation of REV-ERBα causes cancer cell death but does not affect the viability of nontransformed cells. Mechanistically, REV-ERBα suppresses de novo lipid biosynthesis through repression of two key rate-limiting enzymes (i.e., fatty acid synthase and stearoyl-CoA desaturase 1), resulting in a deficiency of oleic acid [90]. Moreover, REV-ERBα activation inhibits autophagy as evidenced by accumulation of p62 and lysosomes and a reduction in autophagosomes [90]. Additionally, SR9009 impairs viability of NRAS-driven naevi and glioblastoma growth and improves animal survival [90]. Taken together, REV-ERBα regulates cancer development via suppressing proliferation, de novo lipogenesis and autophagy, and inducing apoptosis in cancer cells. REV-ERBβ is also shown to be overexpressed in breast cancer cells in the study of De Mei et al [91]. Unlike REV-ERBα, REV-ERBβ displays a cytoprotective action [91]. The cytoprotective function of REV-ERBβ appears to operate downstream of autophagy blockade [91]. The authors demonstrated that dual inhibition of both REV-ERBβ and autophagy may be an effective strategy for eliciting cytotoxicity in cancer cells [91].
Role of REV-ERBα in drug metabolism
Metabolism (biotransformation catalyzed by drug-metabolizing enzymes) is a main defense mechanism of the body against xenobiotic threats, and regarded as a key determinant of pharmacokinetics (and drug exposure) and therefore of pharmacological effects. On the other hand, toxic metabolites may be generated from metabolism reactions, causing adverse effects and disfavoring new drug development. Many drug-metabolizing enzymes (DMEs) are expressed in a circadian time-dependent manner [18]. Circadian expressions of DMEs most likely result in dosing time-dependent pharmacokinetics and therefore in time-varying drug effects (toxicity and efficacy) [18]. Indeed, over 300 medications showed time-varying effects (up to a 10-fold magnitude) [92,93]. Oxaliplatin, a drug for treating colorectal cancer, is a well-documented case that was initially halted in phase I clinical trial due to safety problem (extensive toxicity) [94]. However, the safety of oxaliplatin was latter established in phase I and phase II clinical trials by using the knowledge of chronopharmacology [95,96]. Therefore, integrating chronopharmacology with drug development processes would help to reduce adverse effects and maximize efficacy via dosing time optimization [97].
Cyp7a1 is a REV-ERBα-controlled enzyme that catalyzes the first and rate-limiting step of bile acid biosynthesis (or cholesterol catabolism). REV-ERBα regulates expression of Cyp7a1 and its activity is a determinant of the efficiency of bile acid biosynthesis [74-76]. The mechanism by which REV-ERBα regulates Cyp7a1 is controversial. Indirect mechanisms involving one or two transcriptional factors such as Shp, E4bp4, Srebp and Lrh-1 have been proposed by multiple groups of investigators [74-76]. Moreover, REV-ERBα is a negative regulator of Ces2, a family of phase I enzymes that play an important role in xenobiotic clearance and lipid metabolism [98]. E4bp4 regulates Ces2 enzymes through inhibition of the repressor activity of REV-ERBα, thereby impacting the metabolism and pharmacokinetics of the Ces2 substrate CPT-11 (or irinotecan, a first-line drug for treating colorectal cancer).
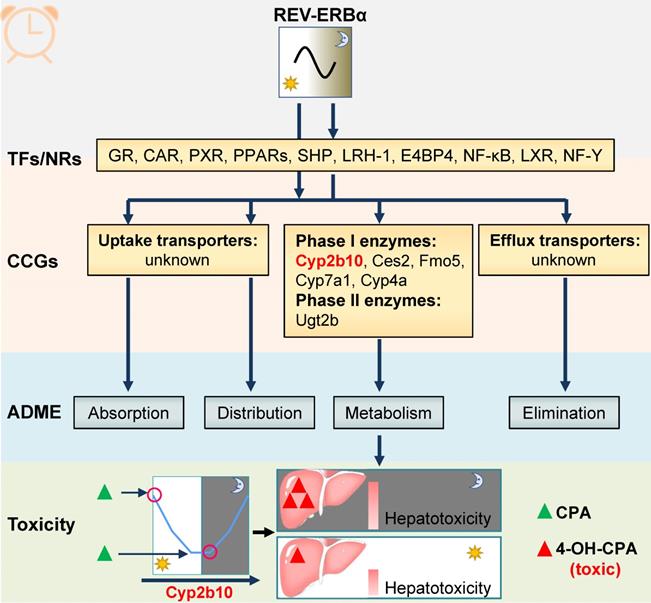
REV-ERBα transcriptionally regulates cycling enzymes such as Ugt2b, Cyp2b10 and Cyp4a10/14 (Figure 5) [99-101]. Regulation by REV-ERBα contributes to circadian expressions of these enzymes and to circadian metabolism and pharmacokinetics of drug substrates such as morphine [99]. Additionally, circadian enzymes and metabolism usually leads to chronotoxicity. For instance, Cyp2b10 metabolizes cyclophosphamide (CPA) to its toxic metabolite 4-OH CPA [101]. CPA hepatotoxicity is dosing time-dependent in mice with high levels of toxicity at ZT2/22 and low levels at ZT10/14 [101]. The CPA chronotoxicity is associated with time-varying formation of 4-OH CPA caused by diurnal Cyp2b10 expression (Figure 5) [101].
REV‑ERBα is involved in chronopharmacokinetics and chronotoxicity via regulating drug-metabolizing enzymes (DMEs). REV-ERBα directly or indirectly regulates clock-controlled genes (CCGs) involved in drug metabolism. The DMEs consists of “phase I enzymes” and “phase II enzymes”. Circadian expressions of these genes result in dosing time-dependent pharmacokinetics and therefore in time-varying drug effects (toxicity and efficacy). For instance, Cyp2b10 (a Rev-erbα target) metabolizes cyclophosphamide (CPA) to its toxic (4-hydroxylated) metabolite 4-OH-CPA. The CPA chronotoxicity is associated with time-varying generation of 4-OH-CPA caused by diurnal Cyp2b10 expression.
Overview of ligands for REV-ERBα
REV-ERBα is a nuclear receptor that can be targeted by small-molecule ligands. Burris and co-workers performed an excellent review of REV-ERB ligands in 2014 [4]. Recent years have witnessed discovery of an array of new REV-ERBα ligands most of which have pharmacological activities in vivo (Figure 6). In the following sections, we describe the newly discovered ligands and the old ones together with their targeting potential (Figure 6). It is noteworthy that all these REV-ERBα ligands most likely also act on its paralog REV-ERBβ.
Historical timeline for discovery of REV-ERBs and development of representative ligands from 1989 to 2019. Chemical structures of all ligands except SR12418 (unavailable) are shown in the figure.
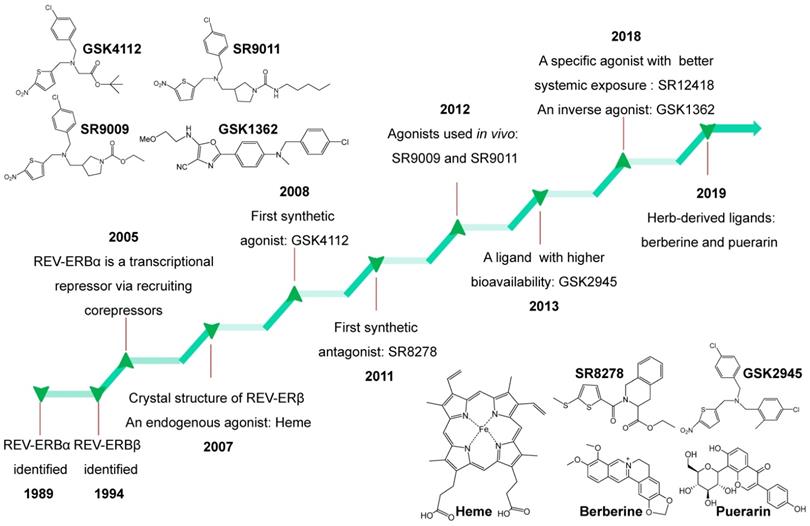
Heme
Heme was identified as an endogenous ligand (agonist) for REV-ERBs In 2007 [102,103]. Heme binds to ligand-binding pocket of REV-ERBs via interactions with two residues, a histidine on helix 11 and a cysteine on helix 3 [104]. Additionally, bulky hydrophobic residues in the ligand-binding pocket form hydrophobic interactions with the porphyrin ring of heme molecule [4]. As a prototypical agonist, heme has been used to verify the effects of REV-ERB activation on gene expressions in in vitro studies [105]. Manipulation of heme homeostasis is shown to alter circadian gene expression and glucose metabolism, highlighting the role of heme and REV-ERBs in circadian biology [68,106].
GSK4112
Discovery of heme as a REV-ERB ligand opens the door for the development of more potent and effective synthetic ligands. GSK4112 (also known as SR6472) is the first synthetic ligand for REV-ERBs, identified based on a fluorescence resonance energy transfer (FRET) assay [107]. GSK4112 has been used as an in vitro probe of REV-ERBα functions. Note that GSK4112 is not suitable to probe the functions of REV-ERBα in vivo due to a low system exposure (poor pharmacokinetic property). Activation of REV-ERBα by GSK4112 inhibits NF-κB signaling and NLRP3 inflammasome activity thus prevents production of cytokines and chemokine, pointing to an anti-inflammatory role of REV-ERBα [25,35,108,109]. GSK4112 decreases the viability of 3T3-L1 preadipocytes and reduces the expressions of cyclin D (a proliferation-related gene) and β-catenin, revealing a role of REV-ERBα in cell proliferation and apoptosis [87]. Also, GSK4112 reduces glycolysis in human gastric carcinoma cells by inhibiting expressions of the genes encoding rate-limiting enzymes [87].
SR8278
SR8278 is the first synthetic antagonist of REV-ERBs, identified based on Gal4 co-transfection and luciferase reporter assays [110]. SR8278 dose-dependently inhibits the transcriptional repressor activity of REV-ERBs [110]. However, it has a poor pharmacokinetic property with a very short elimination half-life of around 0.17 h [110,111]. SR8278 has been widely used in vitro to probe REV-ERBs functions. SR8278 induces expressions of myogenesis genes (Myod, Myog, and Mhc3) in both proliferating and differentiating myoblasts, indicating a regulatory role of REV-ERBs in myogenesis [23]. Antagonism of REV-ERBs by SR8278 increases the intracellular level of lactate (reduces glycolysis) in both SGC-7901 and BGC-823 cells [87]. There are also several attempts with SR8278 for in vivo studies. SR8278 treatment decreases the levels of plasma and liver homocysteine in mice, indicating alleviation of hyperhomocysteinemia by REV-ERBα antagonism [80]. SR8278 increases lean mass and improves muscle function in dystrophic mice through activation of Wnt signaling [89].
SR9009 and SR9011
SR9009 and SR9011 are two potent REV-ERBs agonists designed based on the chemical structure of GSK4112 [7,112,113]. These two compounds are about threefold more potent and efficacious than GSK4112, and they show better pharmacokinetic properties (may be suitable for in vivo studies). In addition, their exclusive actions on REV-ERBs (no effects on other 46 nuclear receptors) have been confirmed by Gal4-chimeric assays. Accordingly, SR9009 and SR9011 have been widely used to test the effects of REV-ERBs on circadian behaviors and diseases both in vitro and in vivo.
The REV-ERB-specific actions of SR9009 and SR9011 are also supported by loss-of-function studies with REV-ERBα-deficient mice [25,36]. SR9009 alleviates DSS-induced colitis and myocardial ischemia-reperfusion in wild-type mice, but fails to do so in REV-ERBα-deficient mice, indicating that the effects of SR9009 are REV-ERBα-dependent [25,36]. However, two groups of investigators report potential off-target effects of SR9009 and SR9011. These two agonists show certain LXR activity in the study of Trump et al [114]. SR9009 and SR9011 displace a radioligand from the LXRα binding site, and SR9011 increases ABCA1 (a LXR target gene) mRNA in THP-1 cells. In the study of Dierickx et al, SR9009 shows REV-ERBs-independent effects on proliferation, metabolism, and gene transcription in REV-ERBs-deficient mESCs and hepatocytes, although the exact mechanisms for the REV-ERBs-independent effects of SR9009 remain unknown [115].
GSK2945
GSK2945 was also designed based on GSK4112 scaffold, but shows a superior pharmacokinetic profile with a longer half-life of 2.0 h and an oral bioavailability of 23% [114]. This compound dose-dependently represses BMAL1 promoter-driven luciferase reporter activity in U2OS cells, suggesting an agonistic effect on REV-ERBs [114]. However, Zhang et al report that GSK2945 dose-dependently antagonizes the repressor activity of REV-ERBα in a Gal4-chimeric assay [76]. GSK2945 also represses the Bmal1 reporter activity and blocks the agonistic activity of GSK4112 [76]. Additionally, the authors demonstrate that GSK2945 increases the mRNA expressions of BMAL1 and PEPCK (i.e., two known target genes of REV-ERBs) in HepG2 cells and hepatocytes as well as in mice [76]. Whether GSK2945 is an agonist or antagonist is not conclusive so far. There is a possibility that the action of GSK2945 may be cell/tissue specific as the activities of REV-ERBs can be affected by the cellular microenvironments such as the redox state, small-molecule gasses and the types of cofactors [104,114,116,117]. Modifications of ligand-bound REV-ERBs by redox conditions and gasses may result in ligand switching [110].
ARN5187
ARN5187 directly interacts with the LBD of REV-ERBβ, and acts as an antagonist [91]. ARN5187 induces the activity of a RORE-driven luciferase reporter in a concentration-dependent manner, and this effect is lost when RORE is mutated. Additionally, ARN5187 is a dual inhibitor of REV-ERB and autophagy. Application of this dual inhibitor may be an effective strategy for eliciting cytotoxicity in cancer cells.
Chelidamic acid and bilirubin
Hering et al identified chelidamic acid as a REV-ERBα agonist using a cell-based two-hybrid assay system [118]. Chelidamic acid binds specifically to the LBD site of REV-ERBα, leading to enhanced binding of REV-ERBα to the co-repressor NcoR1. Wang et al identified bilirubin, a catabolic product of heme, as an antagonist of REV-ERBα based on Gal4 co-transfection and Bmal1 luciferase reporter assays [119]. Despite structurally related, bilirubin and heme display opposite effects on REV-ERBs (i.e., antagonist vs. agonist). Similar findings are also noted for other structurally related compounds (e.g., SR8278 vs. GSK4112; cobalt protoporphyrin IX vs. heme) [110].
GSK1362
Pariollaud et al developed a novel selective oxazole-based inverse agonist for REV-ERBs, named GSK1362 [40]. GSK1362 inhibits interactions of REV-ERBα with NCoR1 and SMRT2 peptides according to FRET assays. It also dose-dependently increases Bmal1 promoter-driven luciferase reporter activity in HEK293 cells. Furthermore, the authors established a model for binding of GSK1362 to REV-ERBα with a cellular thermal shift assay, and demonstrated that the O-methyl ethanolamine side chain of the oxazole (forming a key hydrogen bond with Lys473) is crucial for the compound's activity. Of note, GSK1362 does not induce expressions of Abca1 and Abcg1 (two known LXR target genes), suggesting no effects on LXR receptor. Surprisingly, GSK1362 represses LPS-induced Il-6 in bone marrow-derived macrophages as an REV-ERB agonist does, raising a possibility of an off-target effect.
SR12418
Amir et al synthesized a REV-ERB-specific synthetic ligand (named SR12418) by modifying the chemical structure of SR9009 [33]. SR12418 binds to REV-ERBs according to the time-resolved fluorescence resonance energy transfer assay, and shows an exclusive action based on Gal4-chimeric assays. It potently suppresses Bmal1-luciferase reporter activity with an IC50 less than one tenth of that of SR9009 (68 nM for SR12418 and 710 nM for SR9009). SR12418 is more effective than SR9009 in inhibiting REV-ERB target genes such as IL-17. Moreover, SR12418 displays a better pharmacokinetic property than SR9009. It has been used as an in vivo probe to examine the pharmacological effects of REV-ERBs on experimental autoimmune encephalomyelitis and colitis [33].
Berberine and puerarin
Berberine (initially isolated from Rhizoma Coptidis) is reported to be an agonist of REV-ERBα based on three lines of evidence [32]. First, berberine inhibits Bmal1-luciferase and Nlrp3-luciferase reporter activities. Second, berberine enhances the REV-ERBα repressor activity in a Gal4 co-transfection assay. Third, treatment of bone marrow-derived macrophages with berberine results in decreased expressions of REV-ERBα target genes. Puerarin is isolated from Puerariae Radix, a traditional Chinese medicine widely used to treat fever, emesis, diarrhea, cardiac dysfunctions, and liver injury [120]. Chen et al found that puerarin acts as an antagonist of REV-ERBα based on luciferase reporter, Gal4 co-transfection and target gene expression assays [121]. Berberine and puerarin differ greatly from other synthetic ligands in chemical structure, indicating discovery of novel chemical scaffolds for REV-ERBα ligands. However, the selectivity of berberine and puerarin toward REV-ERBs has not been validated.
Other ligands
GSK0999, GSK5072 and GSK2667 were identified together with GSK2945 in the same study [114]. These three compounds show similar pharmacokinetic profiles to that of SR9009. Additionally, they have no effects on LXRα. ENA_T5382514, ENA_T5445822 and ENA_T5603164 were identified as REV-ERB ligands in a large-scale screening with 29568 diverse compounds from the Enamine compound library [113]. ENA_T5382514 and ENA_T5445822 are agonists, whereas ENA_T5603164 is an antagonist. However, all these three compounds are concerned with off-target effects (e.g., effects on xenobiotic nuclear receptors such as CAR and PXR).
Promises and challenges
Extensive studies uncover a rather broad role of REV-ERBα in pathological conditions including local inflammatory diseases, metabolic disorders, heart failure and cancers. REV-ERBα ligands have been shown to ameliorate the pathologic conditions in animals (preclinical studies), defining pharmacological activities of these ligands. One prominent advantage of targeting REV-ERBα refers to its pleiotropic effects on multiple cellular and molecular pathways [122]. For instance, treatment of diet-induced obese mice with a REV-ERB agonist decreases obesity by reducing fat mass, increasing energy expenditure, and improving dyslipidaemia and hyperglycaemia [7]. Despite well-established pharmacological effects of REV-ERBα ligands in animals, no progress has been made regarding their translation to clinical trials. This implies certain challenges associated with drug development of REV-ERBα ligands. Such challenges may include drug safety problem (or adverse effects), suboptimal pharmacokinetics, and potential gap of circadian mechanisms between humans and rodents.
The broad role of REV-ERBα in pathophysiology is a double-edged sword. The broad actions may ensure effectiveness of drugs in treatment of diseases involving multiple cellular and molecular pathways as noted above. However, in terms of drug development, broad actions also mean possible unwanted effects (adverse effects or toxicity). Severe toxicity is one of main cases of drug attrition [123]. This is particularly the case for REV-ERBα targeting because activation of REV-ERBα is therapeutically beneficial for certain pathologic conditions (e.g., obesity and inflammations), but is detrimental under other circumstances such as Alzheimer's disease and hyperhomocysteinamia [80,124]. Therefore, adverse effects might be a limiting factor to dug development of REV-ERBα ligands.
Suboptimal pharmacokinetic property with poor bioavailability is perhaps another limiting factor for drug development of synthetic REV-ERBα ligands. Current synthetic ligands are cleared rapidly in the body with short half-lives of < 3 h. Because of this, ligands are repeatedly injected daily for more than one week (an unsatisfactory dosing regimen for humans) in efficacy studies with animals. There is a high possibility that these synthetic ligands are rapidly cleared in humans (and even more rapid compared with rodents) and their effectiveness against diseases is impossible to maintain.
Due to an inversed activity-rest cycle between humans and mice (a nocturnal species), serious concerns are raised regarding whether the defined roles of the circadian clock component REV-ERBα in various diseases (and disease-regulatory mechanisms) with mice can be translated to humans. This means that REV-ERBα ligands may be not efficacious at all in humans although they are in animals. Elucidating circadian mechanisms in diseases in humans remains a major task in the field of chronobiology due to the lack of appropriate approaches for extrapolating animal circadian data to humans. It is postulated that advanced models such as humanized animals and primates may address the current gap in circadian studies between humans and mice.
Another issue worthy of attention is that several REV-ERB ligands such as SR9009 have been shown to be “impure”, displaying REV-ERB-independent biological effects (off-target effects). There is a need to determine the selectivity of other REV-ERB ligands (particularly, potential REV-ERB-targeting drugs) and to understand the off-target effects.
Conclusion
Extensive studies uncover a rather broad role of REV-ERBα in pathological conditions including local inflammatory diseases, metabolic disorders, heart failure and cancers. An array of REV-ERBα ligands have been shown to target REV-ERBα to elicit pharmacological effects. Despite well-established pharmacological effects of REV-ERBα ligands in animals (preclinical studies), no progress has been made regarding their translation to clinical trials. The challenges associated with drug development of REV-ERBα ligands may include safety problem (or adverse effects), suboptimal pharmacokinetics, and potential gap of circadian mechanisms between humans and rodents. For successful drug development, it is suggested that REV-ERBα should be targeted to treat local diseases and a targeting drug should be locally distributed, avoiding the adverse effects on other tissues.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 81722049) and China Postdoctoral Science Foundation (No. 2019M663401).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Miyajima N, Horiuchi R, Shibuya Y, Fukushige S, Matsubara K, Toyoshima K. et al. Two erbA homologs encoding proteins with different T3 binding capacities are transcribed from opposite DNA strands of the same genetic locus. Cell. 1989;57:31-9
2. Yin L, Wu N, Lazar MA. Nuclear receptor Rev-erbalpha: a heme receptor that coordinates circadian rhythm and metabolism. Nucl Recept Signal. 2010;8:e001
3. Everett LJ, Lazar MA. Nuclear receptor Rev-erbα: up, down, and all around. Trends Endocrinol Metab. 2014;25:586-92
4. Kojetin DJ, Burris TP. REV-ERB and ROR nuclear receptors as drug targets. Nat Rev Drug Discov. 2014;13:197-216
5. Alenghat T, Meyers K, Mullican SE, Leitner K, Adeniji-Adele A, Avila J, Bućan M. et al. Nuclear receptor corepressor and histone deacetylase 3 govern circadian metabolic physiology. Nature. 2008;456:997-1000
6. Cho H, Zhao X, Hatori M, Yu RT, Barish GD, Lam MT. et al. Regulation of circadian behaviour and metabolism by REV-ERB-α and REV-ERB-β. Nature. 2012;485:123-7
7. Solt LA, Wang Y, Banerjee S, Hughes T, Kojetin DJ, Lundasen T. et al. Regulation of circadian behaviour and metabolism by synthetic REV-ERB agonists. Nature. 2012;485:62-8
8. Curtis A M, Bellet M M, Sassone-Corsi P, O'Neill LA. Circadian clock proteins and immunity. Immunity. 2014;40:178-86
9. Takahashi JS. Transcriptional architecture of the mammalian circadian clock. Nat Rev Genet. 2017;18(3):164-179
10. Eide EJ, Woolf MF, Kang H, Woolf P, Hurst W, Camacho F. et al. Control of mammalian circadian rhythm by CKIɛ-regulated proteasome-mediated PER2 degradation. Mol. Cell. Biol. 2005;25:2795-807
11. Camacho F, Cilio M, Guo Y, Virshup DM, Patel K, Khorkova O. et al. Human casein kinase Iδ phosphorylation of human circadian clock proteins period 1 and 2. FEBS Lett. 2001;489:159-65
12. Hirano A, Fu YH, Ptáček LJ. The intricate dance of post-translational modifications in the rhythm of life. Nat Struct Mol Biol. 2016;23:1053-60
13. Yoo SH, Mohawk JA, Siepka SM, Shan Y, Huh SK, Hong HK. Competing E3 ubiquitin ligases govern circadian periodicity by degradation of CRY in nucleus and cytoplasm. Cell. 2013;152:1091-105
14. Papazyan R, Zhang Y, Lazar MA. Genetic and epigenomic mechanisms of mammalian circadian transcription. Nat Struct Mol Biol. 2016;23:1045-1052
15. Kim YH, Marhon SA, Zhang Y, Steger DJ, Won KJ, Lazar MA. Rev-erbα dynamically modulates chromatin looping to control circadian gene transcription. Science. 2018;359:1274-7
16. Lam MT, Cho H, Lesch HP, Gosselin D, Heinz S, Tanaka-Oishi Y. et al. Rev-Erbs repress macrophage gene expression by inhibiting enhancer- directed transcription. Nature. 2013;498:511-515
17. Chen M, Guan B, Xu H, Yu F, Zhang T, Wu B. The Molecular Mechanism Regulating Diurnal Rhythm of Flavin-Containing Monooxygenase 5 in Mouse Liver. Drug Metab Dispos. 2019;47:1333-42
18. Zhao M, Xing H, Chen M, Dong D, Wu B. Circadian clock-controlled drug metabolism and transport. Xenobiotica. 2019:1-11
19. X. Yu, D. Rollins, K. A. Ruhn, J. J. Stubblefield, C. B. Green, M. Kashiwada et al. TH17 cell differentiation is regulated by the circadian clock. Science. 2013;342:727-30
20. Fang B, Everett LJ, Jager J, Briggs E, Armour SM, Feng D. et al. Circadian enhancers coordinate multiple phases of rhythmic gene transcription in vivo. Cell. 2014;159:1140-52
21. Zhang Y, Fang B, Emmett MJ, Damle M, Sun Z, Feng D. et al. Discrete functions of nuclear receptor Rev-erbα couple metabolism to the clock. Science. 2015;348:1488-92
22. Caratti G, Iqbal M, Hunter L, Kim D, Wang P, Vonslow RM. et al. REVERBa couples the circadian clock to hepatic glucocorticoid action. J Clin Invest. 2018;128:4454-71
23. Welch RD, Guo C, Sengupta M, Carpenter KJ, Stephens NA, Arnett SA. Rev-Erb co-regulates muscle regeneration via tethered interaction with the NF-Y cistrome. Mol Metab. 2017;6:703-14
24. Pivovarova O, Gögebakan Ö, Sucher S, Groth J, Murahovschi V, Kessler K. et al. Regulation of the clock gene expression in human adipose tissue by weight loss. Int J Obes. 2016;40:899-906
25. Wang S, Lin Y, Yuan X, Li F, Guo L, Wu B. REV-ERBα integrates colon clock with experimental colitis through regulation of NF-κB/NLRP3 axis. Nat Commun. 2018;9:4246
26. Goumidi L, Grechez A, Dumont J, Cottel D, Kafatos A, Moreno LA. et al. Impact of REV-ERB alpha gene polymorphisms on obesity phenotypes in adult and adolescent samples. Int J Obes. 2013;37:666-72
27. Pourcet B, Zecchin M, Ferri L, Beauchamp J, Sitaula S, Billon C. et al. Nuclear Receptor Subfamily 1 Group D Member 1 Regulates Circadian Activity of NLRP3 Inflammasome to Reduce the Severity of Fulminant Hepatitis in Mice. Gastroenterology. 2018;154:1449-64.e20
28. Straub R.H. and Cutolo, M.Circadian rhythms in rheumatoid arthritis: implications for pathophysiology and therapeutic management. Arthritis Rheum. 2007;56:399-408
29. Bechtold DA, Gibbs JE, Loudon AS. Circadian dysfunction in disease. Trends Pharmacol Sci. 2010;31:191-8
30. Montaigne D, Marechal X, Modine T, Coisne A, Mouton S, Fayad G. et al. Daytime variation of perioperative myocardial injury in cardiac surgery and its prevention by Rev-Erbα antagonism: a single-centre propensity-matched cohort study and a randomised study. Lancet. 2018;391:59-69
31. Durrington HJ, Farrow SN, Loudon AS, Ray DW. The circadian clock and asthma. Thorax. 2014;69:90-2
32. Zhou Z, Lin Y, Gao L, Yang Z, Wang S, Wu B. Circadian pharmacological effects of berberine on chronic colitis in mice: role of the clock component Rev-erbα. Biochem Pharmacol. 2019:113773
33. Amir M, Chaudhari S, Wang R, Campbell S, Mosure SA, Chopp LB. et al. REV-ERBα Regulates TH17 Cell Development and Autoimmunity. Cell Rep. 2018;25:3733-3749.e8
34. Griffin P, Dimitry JM, Sheehan PW, Lananna BV, Guo C, Robinette ML. Circadian clock protein Rev-erbα regulates neuroinflammation. Proc Natl Acad Sci U S A. 2019;116:5102-7
35. Guo DK, Zhu Y, Sun HY, Xu XY, Zhang S, Hao ZB. Pharmacological activation of REV-ERBα represses LPS-induced microglial activation through the NF-κB pathway. Acta Pharmacol Sin. 2019;40:26-34
36. Reitz CJ, Alibhai FJ, Khatua TN, Rasouli M, Bridle BW, Burris TP. et al. SR9009 administered for one day after myocardial ischemia-reperfusion prevents heart failure in mice by targeting the cardiac inflammasome. Commun Biol. 2019;2:353
37. Zhang L, Zhang R, Tien CL, Chan RE, Sugi K, Fu C. REV-ERBα ameliorates heart failure through transcription repression. JCI Insight. 2017;2:pii 95177
38. Stujanna EN, Murakoshi N, Tajiri K, Xu D, Kimura T, Qin R. Rev-erb agonist improves adverse cardiac remodeling and survival in myocardial infarction through an anti-inflammatory mechanism. PLoS One. 2017;12:e0189330
39. Chang C, Loo CS, Zhao X, Solt LA, Liang Y, Bapat SP. The nuclear receptor REV-ERBα modulates Th17 cell-mediated autoimmune disease. Proc Natl Acad Sci U S A. 2019;116:18528-36
40. Pariollaud M, Gibbs JE, Hopwood TW, Brown S, Begley N, Vonslow R. Circadian clock component REV-ERBα controls homeostatic regulation of pulmonary inflammation. J Clin Invest. 2018;128:2281-96
41. Kalogeris T, Baines CP, Krenz M, Korthuis RJ. Cell biology of ischemia/reperfusion injury. Int Rev Cell Mol Biol. 2012;298:229-317
42. Wang Q, Robinette ML, Billon C, Collins PL, Bando JK, Fachi JL. et al. Circadian rhythm-dependent and circadian rhythm-independent impacts of the molecular clock on type 3 innate lymphoid cells. Sci Immunol. 2019:4 (40). pii: eaay7501
43. Sitaula S, Billon C, Kamenecka TM, Solt LA, Burris TP. Suppression of atherosclerosis by synthetic REV-ERB agonist. Biochem Biophys Res Commun. 2015;460:566-71
44. Zhao W, Cui L, Huang X, Wang S, Li D, Li L. et al. Activation of Rev-erbα attenuates lipopolysaccharide-induced inflammatory reactions in human endometrial stroma cells via suppressing TLR4-regulated NF-κB activation. Acta Biochim Biophys Sin. 2019;51:908-14
45. Fontaine C, Rigamonti E, Pourcet B, Duez H, Duhem C, Fruchart JC. et al. The nuclear receptor Rev-erbalpha is a liver X receptor (LXR) target gene driving a negative feedback loop on select LXR-induced pathways in human macrophages. Mol. Endocrinol. 2008;22:1797-811
46. Gibbs JE, Blaikley J, Beesley S, Matthews L, Simpson KD, Boyce SH. et al. The nuclear receptor REV-ERBα mediates circadian regulation of innate immunity through selective regulation of inflammatory cytokines. Proc Natl Acad Sci U S A. 2012;109:582-7
47. Sato S, Sakurai T, Ogasawara J, Takahashi M, Izawa T, Imaizumi K. et al. A circadian clock gene, Rev- erbα, modulates the inflammatory function of macrophages through the negative regulation of Ccl2 expression. J. Immunol. 2014;192:407-17
48. Sato S, Sakurai T, Ogasawara J, Shirato K, Ishibashi Y, Oh-ishi S. et al. Direct and indirect suppression of interleukin-6 gene expression in murine macrophages by nuclear orphan receptor REV-ERBα. ScientificWorldJournal. 2014;2014:685854
49. Scheiermann C, Gibbs J, Ince L, Loudon A. Clocking in to immunity. Nat Rev Immunol. 2018;18:423-37
50. Fortier EE, Rooney J, Dardente H, Hardy MP, Labrecque N, Cermakian N. Circadian variation of the response of T cells to antigen. J. Immunol. 2011;187:6291-300
51. E. Vivier, D. Artis, M. Colonna, A. Diefenbach, J. P. Di Santo, G. Eberl, et al. Innate lymphoid cells: 10 years on. 2018;174:1054-66
52. Yu X, Rollins D, Ruhn KA, Stubblefield JJ, Green CB, Kashiwada M. et al. TH17 cell differentiation is regulated by the circadian clock. Science. 2013;342:727-30
53. Cho J.H. The genetics and immunopathogenesis of inflammatory bowel disease. Nat. Rev. Immunol. Nat Rev Immunol. 2008;8:458-66
54. Farez MF, Mascanfroni ID, Méndez-Huergo SP, Yeste A, Murugaiyan G, Garo LP. et al. Melatonin Contributes to the Seasonality of Multiple Sclerosis Relapses. Cell. 2015;162:1338-52
55. Li T, Eheim AL, Klein S, Uschner FE, Smith AC, Brandon-Warner E, Ghosh S. et al. Novel role of nuclear receptor Rev-erbα in hepatic stellate cell activation: potential therapeutic target for liver injury. Hepatology. 2014;59:2383-96
56. Cunningham PS, Meijer P, Nazgiewicz A, Anderson SG, Borthwick LA, Bagnall J. et al. The circadian clock protein REVERBα inhibits pulmonary fibrosis development. Proc Natl Acad Sci U S A. 2019: pii. 2019 12109
57. Yin L, Wu N, Curtin JC, Qatanani M, Szwergold NR, Reid RA. et al. Rev-erb alpha, a heme sensor that coordinates metabolic and circadian pathways. Science. 2007;318:1786-9
58. Yuan X, Dong D, Li Z, Wu B. Rev-erbα activation down-regulates hepatic Pck1 enzyme to lower plasma glucose in mice. Pharmacol Res. 2019;141:310-8
59. Delezie J, Dumont S, Dardente H, Oudart H, Gréchez-Cassiau A, Klosen P. et al. The nuclear receptor REV-ERBα is required for the daily balance of carbohydrate and lipid metabolism. FASEB J. 2012;26:3321-35
60. Li X, Xu M, Wang F, Kohan AB, Haas MK, Yang Q. et al. Apolipoprotein A-IV reduces hepatic gluconeogenesis through nuclear receptor NR1D1. J Biol Chem. 2014;289:2396-404
61. Grant D, Yin L, Collins JL, Parks DJ, Orband-Miller LA, Wisely GB. et al. GSK4112, a small molecule chemical probe for the cell biology of the nuclear heme receptor Rev-erbα. ACS Chem Biol. 2010;5:925-32
62. Vieira E, Marroquí L, Batista TM, Caballero-Garrido E, Carneiro EM, Boschero AC. et al. The clock gene Rev-erbα regulates pancreatic β cell function: modulation by leptin and high-fat diet. Endocrinology. 2012;153:592-601
63. Vieira E, Merino B, Quesada I. Role of the clock gene Rev-erbα in metabolism and in the endocrine pancreas. Diabetes Obes Metab. 2015;17(Suppl 1):106-14
64. Vieira E, Marroquí L, Figueroa AL, Merino B, Fernandez-Ruiz R, Nadal A. et al. Involvement of the clock gene Rev-erb alpha in the regulation of glucagon secretion in pancreatic alpha-cells. PLoS One. 2013;8:e69939
65. Costes S, Laouteouet D, Ravier M, Delobel M. et al. 325-LB: Circadian clock nuclear receptor REV-ERBa Is a novel regulator of beta-cell function, survival, and autophagy under diabetogenic conditions. Diabetes. 2019:68 (Supplement 1)
66. Altman BJ, Hsieh AL, Sengupta A, Krishnanaiah SY, Stine ZE, Walton ZE. MYC Disrupts the Circadian Clock and Metabolism in Cancer Cells. Cell Metab. 2015;22:1009-19
67. Zhao X, Hirota T, Han X, Cho H, Chong LW, Lamia K. Circadian Amplitude Regulation via FBXW7-Targeted REV-ERBα Degradation. Cell. 2016;165:1644-57
68. Simcox JA, Mitchell TC, Gao Y, Just SF, Cooksey R, Cox J. et al. Dietary iron controls circadian hepatic glucose metabolism through heme synthesis. Diabetes. 2015;64:1108-19
69. Raspe E, Duez H, Mansen A, Fontaine C, Fievet C, Fruchart JC. et al. Identification of Rev-erb alpha as a physiological repressor of apoC-III gene transcription. J Lipid Res. 2002;43:2172-9
70. Bugge A, Feng D, Everett LJ, Briggs ER, Mullican SE, Wang F. et al. Rev-erbalpha and Rev-erbbeta coordinately protect the circadian clock and normal metabolic function. Genes Dev. 2012;26:657-67
71. Sitaula S, Zhang J, Ruiz F, Burris TP. Rev-erb regulation of cholesterologenesis. Biochem Pharmacol. 2017;131:68-77
72. Anzulovich A, Mir A, Brewer M, Ferreyra G, Vinson C, Baler R. Elovl3: a model gene to dissect homeostatic links between the circadian clock and nutritional status. J Lipid Res. 2006;47:2690-700
73. Zhang Y, Fang B, Damle M, Guan D, Li Z, Kim YH. et al. HNF6 and Rev-erbα integrate hepatic lipid metabolism by overlapping and distinct transcriptional mechanisms. Genes Dev. 2016;30:1636-44
74. Duez H, van der Veen JN, Duhem C, Pourcet B, Touvier T, Fontaine C. et al. Regulation of bile acid synthesis by the nuclear receptor Reverbalpha. Gastroenterology. 2008;135:689-98
75. Le Martelot G, Claudel T, Gatfield D, Schaad O, Kornmann B, Lo Sasso G. et al. REV-ERBalpha participates in circadian SREBP signaling and bile acid homeostasis. PLoS Biol. 2009;7:e1000181
76. Zhang T, Zhao M, Lu D, Wang S, Yu F, Guo L. et al. REV-ERBα Regulates CYP7A1 Through Repression of Liver Receptor Homolog-1. Drug Metab Dispos. 2018;46:248-58
77. Garaulet M, Smith CE, Gomez-Abellán P, Ordovás-Montañés M, Lee YC, Parnell LD. et al. REV-ERB-ALPHA circadian gene variant associates with obesity in two independent populations: Mediterranean and North American. Mol Nutr Food Res. 2014;58:821-9
78. Ruano EG, Canivell S, Vieira E. REV-ERB ALPHA polymorphism is associated with obesity in the Spanish obese male population. PLoS One. 2014;9:e104065
79. Selhub J. Homocysteine metabolism. Annu Rev Nutr. 1999;19:217-46
80. Zhang T, Chen M, Guo L, Yu F, Zhou C, Xu H. et al. Reverse Erythroblastosis Virus α Antagonism Promotes Homocysteine Catabolism and Ammonia Clearance. Hepatology. 2019;70:1770-84
81. Quevedo I, Zuniga AM. Low bone mineral density in rotating-shift workers. J Clin Densitom. 2010;13:467-9
82. Feskanich D, Hankinson SE, Schernhammer ES. Nightshift work and fracture risk: the Nurses' Health Study. Osteoporos Int. 2009;20:537-42
83. Zvonic S, Ptitsyn AA, Kilroy G, Wu X, Conrad SA, Scott LK. et al. Circadian oscillation of gene expression in murine calvarial bone. J Bone Miner Res. 2007;22(3):357-65
84. Song C, Tan P, Zhang Z, Wu W, Dong Y, Zhao L. et al. REV-ERB agonism suppresses osteoclastogenesis and prevents ovariectomy-induced bone loss partially via FABP4 upregulation. FASEB J. 2018;32:3215-28
85. He Y, Lin F, Chen Y, Tan Z, Bai D, Zhao Q. Overexpression of the Circadian Clock Gene Rev-erbα Affects Murine Bone Mesenchymal Stem Cell Proliferation and Osteogenesis. Stem Cells Dev. 2015;24:1194-204
86. Wang X, Wang N, Wei X, Yu H, Wang Z. REV-ERBα reduction is associated with clinicopathological features and prognosis in human gastric cancer. Oncol Lett. 2018;16:1499-506
87. Chu G, Zhou X, Hu Y, Shi S, Yang G. Rev-erbα Inhibits Proliferation and Promotes Apoptosis of Preadipocytes through the Agonist GSK4112. Int J Mol Sci. 2019:20 pii: E4524
88. Wang Y, Kojetin D, Burris TP. Anti-proliferative actions of a synthetic REV-ERBα/β agonist in breast cancer cells. Biochem Pharmacol. 2015;96:315-22
89. Tao L, Yu H, Liang R, Jia R, Wang J, Jiang K. et al. Rev-erbα inhibits proliferation by reducing glycolytic flux and pentose phosphate pathway in human gastric cancer cells. Oncogenesis. 2019;8:57
90. Sulli G, Rommel A, Wang X, Kolar MJ, Puca F, Saghatelian A. et al. Pharmacological activation of REV-ERBs is lethal in cancer and oncogene-induced senescence. Nature. 2018;553:351-5
91. De Mei C, Ercolani L, Parodi C, Veronesi M, Lo Vecchio C, Bottegoni G. et al. Dual inhibition of REV-ERBβ and autophagy as a novel pharmacological approach to induce cytotoxicity in cancer cells. Oncogene. 2015;34:2597-608
92. Levi F, Schibler U. Circadian rhythms: mechanisms and therapeutic implications. Annu Rev Pharmacol Toxicol. 2007;47:593-628
93. Dallmann R, Brown SA, Gachon F. Chronopharmacology: new insights and therapeutic implications. Annu Rev Pharmacol Toxicol. 2014;54:339-61
94. Extra JM, Espie M, Calvo F, Ferme C, Mignot L, Marty M. Phase I study of oxaliplatin in patients with advanced cancer. Cancer Chemother Pharmacol. 1990;25:299-303
95. Caussanel JP, Lévi F, Brienza S, Misset JL, Itzhaki M, Adam R. et al. Phase I trial of 5-day continuous venous infusion of oxaliplatin at circadian rhythm-modulated rate compared with constant rate. J Natl Cancer Inst. 1990;82:1046-50
96. Levi F, Perpoint B, Garufi C, Focan C, Chollet P, Depres-Brummer P. et al. Oxaliplatin activity against metastatic colorectal cancer. A phase II study of 5-day continuous venous infusion at circadian rhythm modulated rate. Eur J Cancer. 1993;29A:1280-4
97. Cederroth CR, Albrecht U, Bass J, Brown SA, Dyhrfjeld-Johnsen J, Gachon F. et al. Medicine in the Fourth Dimension. Cell Metab. 2019;30:238-50
98. Zhao M, Zhang T, Yu F, Guo L, Wu B. E4bp4 regulates carboxylesterase 2 enzymes through repression of the nuclear receptor Rev-erbα in mice. Biochem Pharmacol. 2018;152:293-301
99. Zhang T, Guo L, Yu F, Chen M, Wu B. The nuclear receptor Rev-erbα participates in circadian regulation of Ugt2b enzymes in mice. Biochem Pharmacol. 2019;161:89-97
100. Zhang T, Yu F, Guo L, Chen M, Yuan X, Wu B. Small Heterodimer Partner Regulates Circadian Cytochromes p450 and Drug-Induced Hepatotoxicity. Theranostics. 2018;8:5246-58
101. Zhao M, Zhao H, Deng J, Guo L, Wu B. Role of the CLOCK protein in liver detoxification. Br J Pharmacol. 2019;176:4639-52
102. Raghuram S, Stayrook KR, Huang P, Rogers PM, Nosie AK, McClure DB. et al. Identification of heme as the ligand for the orphan nuclear receptors REV-ERBalpha and REV-ERBbeta. Nat Struct Mol Biol. 2007;14:1207-13
103. Yin L, Wu N, Curtin JC, Qatanani M, Szwergold NR, Reid RA. et al. R Rev-erbalpha, a heme sensor that coordinates metabolic and circadian pathways. Science. 2007;318:1786-9
104. Pardee KI, Xu X, Reinking J, Schuetz A, Dong A, Liu S. et al. The structural basis of gas-responsive transcription by the human nuclear hormone receptor REV-ERBbeta. PLoS Biol. 2009;7:e43
105. Lee J, Lee S, Chung S, Park N, Son GH, An H. et al. Identification of a novel circadian clock modulator controlling BMAL1 expression through a ROR/REV-ERB-response element-dependent mechanism. Biochem Biophys Res Commun. 2016;469:580-6
106. Adachi Y, Umeda M, Kawazoe A, Sato T, Ohkawa Y, Kitajima S. et al. The novel heme-dependent inducible protein, SRRD regulates heme biosynthesis and circadian rhythms. Arch Biochem Biophys. 2017;631:19-29
107. Meng QJ, McMaster A, Beesley S, Lu WQ, Gibbs J, Parks D. et al. Ligand modulation of REV-ERBα function resets the peripheral circadian clock in a phasic manner. J Cell Sci. 2008;121:3629-35
108. Morioka N, Tomori M, Zhang FF, Saeki M, Hisaoka-Nakashima K, Nakata Y. Stimulation of nuclear receptor REV-ERBs regulates tumor necrosis factor-induced expression of proinflammatory molecules in C6 astroglial cells. Biochem Biophys Res Commun. 2016;469:151-7
109. Sundar IK, Rashid K, Sellix MT, Rahman I. The nuclear receptor and clock gene REV-ERBα regulates cigarette smoke-induced lung inflammation. Biochem Biophys Res Commun. 2017;493(4):1390-5
110. Kojetin D, Wang Y, Kamenecka TM, Burris TP. Identification of SR8278, a synthetic antagonist of the nuclear heme receptor REV-ERB. ACS Chem Biol. 2011;6:131-4
111. Dong D, Sun H, Wu Z, Wu B, Xue Y, Li Z. A validated ultra-performance liquid chromatography-tandem mass spectrometry method to identify the pharmacokinetics of SR8278 in normal and streptozotocin-induced diabetic rats. J Chromatogr B. 2016;1020:142-7
112. Noel R, Song X, Shin Y, Banerjee S, Kojetin D, Lin L. et al. Synthesis and SAR of tetrahydroisoquinolines as Rev-erbα agonists. Bioorg Med Chem Lett. 2012;22:3739-42
113. Shin Y, Noel R, Banerjee S, Kojetin D, Song X, He Y. et al. Small molecule tertiary amines as agonists of the nuclear hormone receptor Rev-erbα. Bioorg Med Chem Lett. 2012;22:4413-7
114. Trump RP, Bresciani S, Cooper AW, Tellam JP, Wojno J, Blaikley J. et al. Optimized chemical probes for REV-ERBα. J Med Chem. 2013;56:4729-37
115. Dierickx P, Emmett MJ, Jiang C, Uehara K, Liu M, Adlanmerini M. et al. SR9009 has REV-ERB-independent effects on cell proliferation and metabolism. Proc Natl Acad Sci U S A. 2019;116:12147-52
116. Marvin KA, Reinking JL, Lee AJ, Pardee K, Krause HM, Burstyn JN. Nuclear receptors Homo sapiens Rev-erbβ and Drosophila melanogaster E75 are thiolate-ligated heme proteins which undergo redox-mediated ligand switching and bind CO and NO. Biochemistry. 2009;48:7056-71
117. Matta-Camacho E, Banerjee S, Hughes TS, Solt LA, Wang Y, Burris TP. et al. Structure of REV-ERBβ ligand-binding domain bound to a porphyrin antagonist. J Biol Chem. 2014;289:20054-66
118. Y. Hering, A. Berthier, H. Duez, P. Lefebvre, B. Deprez, P. Gribbon, et al. Development and implementation of a cell-based assay to discover agonists of the nuclear receptor REV-ERBα. J Biol Methods. 2018;5:e94
119. Wang S, Lin Y, Zhou Z, Gao L, Yang Z, Li F, Wu B. Circadian Clock Gene Bmal1 Regulates Bilirubin Detoxification: A Potential Mechanism of Feedback Control of Hyperbilirubinemia. Theranostics. 2019;9:5122-33
120. Zhou YX, Zhang H, Peng C. Puerarin: a review of pharmacological effects. Phytother Res. 2014;28:961-75
121. Chen M, Zhou C, Xu H, Zhang T, Wu B. Chronopharmacological targeting of Rev-erbα by puerarin alleviates hyperhomocysteinemia in mice. Biomed Pharmacother. 2020;125:109936
122. Sulli G, Manoogian ENC, Taub PR, Panda S. Training the Circadian Clock, Clocking the Drugs, and Drugging the Clock to Prevent, Manage, and Treat Chronic Diseases. Trends Pharmacol Sci. 2018;39:812-27
123. Guengerich FP. Mechanisms of drug toxicity and relevance to pharmaceutical development. Drug Metab Pharmacokinet. 2011;26:3-14
124. Lee J, Kim DE, Griffin P, Sheehan PW, Kim DH, Musiek ES. et al. Inhibition of REV-ERBs stimulates microglial amyloid-beta clearance and reduces amyloid plaque deposition in the 5XFAD mouse model of Alzheimer's disease. Aging Cell. 2019 e13078
125. Crumbley C, Wang Y, Kojetin DJ, Burris TP. Characterization of the core mammalian clock component, NPAS2, as a REV-ERBalpha/RORalpha target gene. J Biol Chem. 2010;285:35386-92
126. Crumbley C, Burris TP. Direct regulation of CLOCK expression by REV-ERB. PLoS One. 2011;6(3):e17290
127. Chandra V, Mahajan S, Saini A, Dkhar HK, Nanduri R, Raj EB. et al. Human IL10 gene repression by Rev-erbα ameliorates Mycobacterium tuberculosis clearance. J Biol Chem. 2013;288:10692-702
128. Wang J, Yin L, Lazar MA. The orphan nuclear receptor Rev-erb alpha regulates circadian expression of plasminogen activator inhibitor type 1. J Biol Chem. 2006;281:33842-8
129. Schnell A, Chappuis S, Schmutz I, Brai E, Ripperger JA, Schaad O. et al. The nuclear receptor REV-ERBα regulates Fabp7 and modulates adult hippocampal neurogenesis. PLoS One. 2014;9:e99883
130. Jager J, Wang F, Fang B, Lim HW, Peed LC, Steger DJ. et al. The Nuclear Receptor Rev-erbα Regulates Adipose Tissue-specific FGF21 Signaling. J Biol Chem. 2016;291:10867-75
131. Wu N, Yin L, Hanniman EA, Joshi S, Lazar MA. Negative feedback maintenance of heme homeostasis by its receptor, Rev-erbalpha. Genes Dev. 2009;23:2201-9
132. Gerhart-Hines Z, Feng D, Emmett MJ, Everett LJ, Loro E, Briggs ER. et al. The nuclear receptor Rev-erbα controls circadian thermogenic plasticity. Nature. 2013;503:410-3
Author contact
Corresponding author: Baojian Wu Ph.D. Email: bj.wucom
 Global reach, higher impact
Global reach, higher impact