13.3
Impact Factor
Theranostics 2020; 10(15):6661-6673. doi:10.7150/thno.44880 This issue Cite
Research Paper
RNAi-mediated control of CRISPR functions
1. National and Local Joint Engineering Laboratory of Medical Synthetic Biology, Shenzhen, Second People's Hospital, The First Affiliated Hospital of Shenzhen University Health Science Center, Shenzhen518035, China
2. Department of Urology, Shenzhen Second People's Hospital, The First Affiliated Hospital of Shenzhen University Health Science Center, Shenzhen518035, China
3. Guangdong Key Laboratory of Systems Biology and Synthetic Biology for Urogenital Tumors, Shenzhen Second People's Hospital, The First Affiliated Hospital of Shenzhen University Health, Science Center, Shenzhen 518035, China
Abstract
CRISPR-Cas9 has become a versatile tool for genome editing and regulation, and strategies to effectively control its activity have attracted much attention. RNAi, also a gene-regulating tool, is used as another mechanism by which eukaryotes resist the invasion of foreign genetic material.
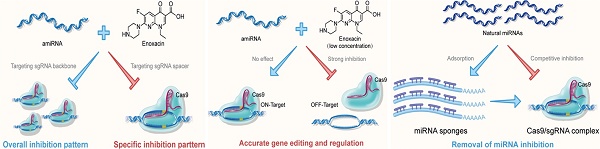
Methods: In this study, we analyzed the quantitative inhibition of the CRISPR system by using artificial miRNAs (amiRNAs) combined with the RNAi enhancer enoxacin to improve the targeting specificity of the CRISPR system. Furthermore, we examined the feasibility of improving the efficiency of gene editing and regulation by blocking the effects of natural intracellular miRNAs on sgRNAs.
Results: amiRNAs targeting the sgRNA were used to control its expression, and the small molecule drug denoxacin was utilized to enhance this effect, especially in the presence of Cas9. amiRNA/enoxacin inhibited CRISPR-mediated gene editing and regulation both in vitro and in vivo and could tune sgRNA-targeting specificity. Furthermore, CRISPR efficiency was increased by blocking the effects of endogenous miRNAs.
Conclusion: Our study provides an efficient molecular switch for conditional regulation of CRISPR activities in mammalian cells and also presents potentially useful approaches for solving current key issues of off-target effects and low targeting efficiency.
Keywords: CRISPR switch, artificial miRNA, miRNA sponge, enoxacin
 Global reach, higher impact
Global reach, higher impact