Impact Factor
Theranostics 2020; 10(17):7581-7598. doi:10.7150/thno.44306 This issue Cite
Research Paper
Single-cell Transcriptome Profiling reveals Dermal and Epithelial cell fate decisions during Embryonic Hair Follicle Development
1. Key Laboratory of Animal Genetics, Breeding and Reproduction of Shaanxi Province, College of Animal Science and Technology, Northwest A&F University, Yangling, Shaanxi 712100, China.
2. College of Life Sciences, Qingdao Agricultural University, Qingdao 266109, China.
Abstract
It is estimated that 50% of men and 25% of women worldwide suffer from hair loss, and therefore it is of great significance to investigate the molecular pathways driving hair follicle de novo morphogenesis. However, due to high cellular heterogeneity and the asynchronous development of hair follicles, our current understanding of the molecular mechanisms involved in follicle development remains limited.
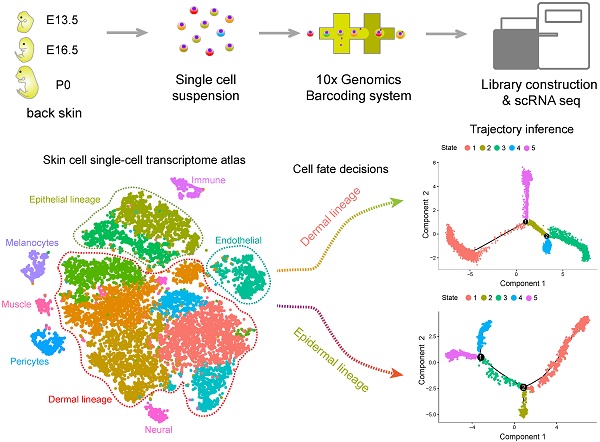
Methods: Single-cell suspensions from the dorsal skin of E13.5 (induction stage), E16.5 (organogenesis) fetal mice, and newborn mice (cytodifferentiation stage, postnatal day 0, P0) were prepared for unbiased single-cell RNA sequencing. To delineate the single-cell transcriptional landscape during hair follicle de novo morphogenesis, we performed t-distributed Stochastic Neighbor Embedding (tSNE), pseudotime cell trajectory inference, and regulon enrichment analysis to dissect cellular heterogeneity and reveal the molecular pathways underlying major cell type cell fate decisions. To validate our analysis, we further performed immunohistochemistry analysis of the key molecules involved during hair follicle morphogenesis. Meanwhile, intercellular communication between different cell populations was inferred based on a priori knowledge of ligand-receptor pairs.
Results: Based on tSNE analysis, we identified 14 cell clusters from skin tissue and delineated their cellular identity from specific gene expression profiles. By using pseudotime ordering analysis, we successfully constructed the epithelium/dermal cell lineage differentiation trajectory. For dermal cell lineage, our analysis here recapitulated the dynamic gene expression profiles during dermal condensate (DC) cell fate commitment and delineated the heterogeneity of the different dermal papilla (DP) cell populations during in utero hair follicle development. For the epithelium cell lineage, our analysis revealed the dynamic gene expression profiles of the underappreciated matrix, interfollicular epidermis (IFE), hair shaft and inner root sheath (IRS) cell populations. Furthermore, single-cell regulatory network inference and clustering analysis revealed key regulons during cell fate decisions. Finally, intercellular communication analysis demonstrated that strong intercellular communication was involved during early hair follicle development.
Conclusions: Our findings here provide a molecular landscape during hair follicle epithelium/dermal cell lineage fate decisions, and recapitulate the sequential activation of core regulatory transcriptional factors (TFs) in different cell populations during hair follicle morphogenesis. More importantly, our study here represents a valuable resource for understanding the molecular pathways involved during hair follicle de novo morphogenesis, which will have implications for future hair loss treatments.
Keywords: Single-cell transcriptome, Cell fate decision, Hair follicle morphogenesis
 Global reach, higher impact
Global reach, higher impact