Impact Factor
Theranostics 2020; 10(17):7821-7835. doi:10.7150/thno.47987 This issue Cite
Review
COVID-19: Progress in diagnostics, therapy and vaccination
1. State Key Laboratory of Molecular Vaccinology and Molecular Diagnostics & National Institute of Diagnostics and Vaccine Development in Infectious Disease, School of Public Health, Xiamen University, 361102, China.
2. State Key Laboratory of Molecular Vaccinology and Molecular Diagnostics & Center for Molecular Imaging and Translational Medicine, School of Public Health, Xiamen University, Xiamen, 361102, China.
#These authors contributed equally to this work.
Received 2020-5-9; Accepted 2020-6-7; Published 2020-6-19
Abstract
Coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has recently become a pandemic. As the sudden emergence and rapid spread of SARS-CoV-2 is endangering global health and the economy, the development of strategies to contain the virus's spread are urgently needed. At present, various diagnostic kits to test for SARS-CoV-2 are available for use to initiate appropriate treatment faster and to limit further spread of the virus. Several drugs have demonstrated in vitro activity against SARS-CoV-2 or potential clinical benefits. In addition, institutions and companies worldwide are working tirelessly to develop treatments and vaccines against COVID-19. However, no drug or vaccine has yet been specifically approved for COVID-19. Given the urgency of the outbreak, we focus here on recent advances in the diagnostics, treatment, and vaccine development for SARS-CoV-2 infection, helping to guide strategies to address the current COVID-19 pandemic.
Keywords: COVID-19, SARS-CoV-2, diagnostics, antiviral therapy
Introduction
The rapid spread of SARS-CoV-2, a novel coronavirus that emerged in December 2019, and the resulting disease, namely, coronavirus disease 2019 (COVID-19), has posed a serious global public health emergency. Whole-genome sequencing results show that the causative agent is a novel coronavirus that was initially named 2019-nCoV by the World Health Organization (WHO) [1-3]. COVID-19 was subsequently recommended as the disease name. As the genomic sequence of the new virus is closely related to that of severe acute respiratory syndrome (SARS) coronavirus (SARS-CoV), the International Committee on Taxonomy of Viruses (ICTV) officially designated the virus SARS-CoV-2 [4]. Patients infected by SARS-CoV-2 manifest a range of symptoms, such as dry cough, fever, headache and dyspnea, and are generally diagnosed with viral pneumonia [5, 6], but asymptomatic infections have also been reported [7, 8].
Coronaviruses, a genus in the Coronaviridae family, are enveloped, positive-sense, single-stranded RNA viruses with the largest known genomes (from 25 to 32 kb) among RNA viruses [9, 10]. Coronavirinae is subdivided into four genera: alpha-, beta-, gamma-, and delta coronaviruses [11]. Phylogenetic analysis of coronavirus genomes has revealed that SARS-CoV-2 is a new member of the Beta coronavirus genus, which includes SARS-CoV and Middle East respiratory syndrome (MERS) coronavirus (MERS-CoV), as well as other viruses that infect humans and diverse animal species. Among the several coronaviruses that are pathogenic to humans, most are associated with mild clinical symptoms (e.g., HCoV-229E, HCoV-NL63, HCoV-OC43, and HCoV-HKU1) [12, 13], with three notable exceptions: SARS-CoV in November 2002 [14-16], MERS-CoV in April 2012 [17-19], and, more recently, SARS-CoV-2 causing COVID-19. SARS-CoV provoked a large-scale epidemic beginning in China and involving 37 countries with approximately 8,000 cases and 774 deaths [20]; MERS-CoV began in Saudi Arabia and caused a total of 2,519 MERS cases with 866 associated deaths as of the end of January 2020 [21].
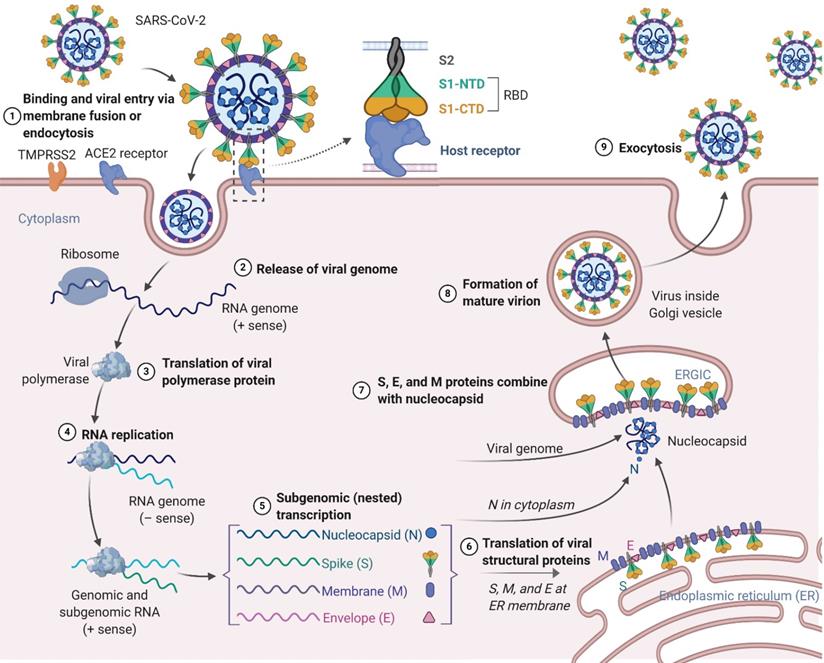
Similar to SARS-CoV and MERS-CoV, the SARS-CoV-2 genome encodes nonstructural proteins (NSPs, such as 3-chymotrypsin-like protease, papain-like protease, helicase, and RNA-dependent RNA polymerase), structural proteins and accessory proteins [22]. SARS-CoV-2 has four structural proteins: the spike (S) protein, envelope (E) protein, membrane (M) protein, and nucleocapsid (N) protein. Among these proteins, the trimeric S protein is indispensable for virus-cell receptor interactions during viral entry [23, 24]. S protein comprises an N-terminal S1 subunit responsible for virus-receptor binding and a C-terminal S2 subunit responsible for virus-cell membrane fusion. S1 is further divided into an N-terminal domain (NTD) and a receptor-binding domain (RBD) [25, 26]. SARS-CoV-2 targets cells through the S protein, which binds to the human angiotensin-converting enzyme 2 (ACE2) receptor and employs the cellular serine protease TMPRSS2 for S protein priming [27-31]. Such binding triggers a cascade of events leading to fusion between the cellular and viral membranes for cell entry. The viral RNA genome is released into the cytoplasm after membrane fusion (see the SARS-CoV-2 lifecycle in Figure 1). Polyproteins are subsequently synthesized to encode the viral replicase-transcriptase complex. The viral RNA is then synthesized by RNA-dependent RNA polymerase. Synthesis of structural proteins is followed by viral particle assembly and release [32, 33]. These viral lifecycle steps provide potential targets for vaccines and therapeutics to prevent and treat SARS-CoV-2 infection. Promising targets include NSPs, which are involved in transcription and replication of the virus, as the key enzymes in the viral lifecycle. Additional targets include viral entry and immune regulation pathways. For example, the S protein plays key roles in the induction of T cell responses and neutralizing antibodies (NAbs), as well as protective immunity, during infection with SARS-CoV-2.
Schematic presentation of the SARS-CoV-2 viral lifecycle. SARS-CoV-2 enters host cells by first binding to angiotensin-converting enzyme 2 (ACE2) via the surface spike (S) protein. Following entry of the virus into the host cell, viral genomic RNA is released and translated into viral polymerase proteins. In this process, subgenomic (-) RNAs are synthesized and used as a template to form subgenomic (+) messenger RNAs (mRNAs). The nucleocapsid (N) structural protein and viral RNA are replicated, transcribed, and synthesized in the cytoplasm, whereas other viral structural proteins, including the S protein, membrane (M) protein and envelope (E) protein, are transcribed and then translated in the endoplasmic reticulum (ER). The resulting structural proteins are further assembled into the nucleocapsid and viral envelope at the ER-Golgi intermediate compartment (ERGIC) to form a mature virion, followed by release of the nascent virion from the host cell.
To elucidate the specific mechanisms of SARS-CoV-2 infection, the crystal structure of the SARS-CoV-2 spike RBD bound to the cell receptor ACE2 at 2.45 Å resolution was recently determined [34]. The overall binding mode of the SARS-CoV-2 RBD with ACE2 is nearly identical to that of the SARS-CoV RBD, which also utilizes ACE2 on the surface of host cells as a receptor. HeLa cells expressing ACE2 are susceptible to SARS-CoV-2 infection, but cells without ACE2 expression are not infected [1]. In vitro binding experiments also show that the SARS-CoV-2 RBD has affinity for ACE2 in the low nM range, indicating that the RBD is the key functional component of the S1 subunit responsible for the binding of ACE2 to SARS-CoV-2 [34]. More structural information at the atomic level would greatly enhance our understanding of the interaction between SARS-CoV-2 and host cells, providing precise targets for NAbs, and assist us in urgently needed structure-based vaccine design in the ongoing fight against SARS-CoV-2. In addition, research on cell modeling tools might promote insight into our understanding of infection mechanisms and speed up the development of therapeutic agents and vaccines for COVID-19 [35].
In this review, we aim to discuss the current status of diagnostics, therapeutics, and vaccines against COVID-19. As rapid diagnostics and development of drugs and vaccines for the virus are important interventions to control the outbreak, a review article that encompasses the current research on this topic may provide help for guiding strategies to address the current COVID-19 pandemic.
Diagnostics
Rapid and accurate detection of COVID-19 is essential to initiate the appropriate treatment rapidly, to limit further spread of the virus and to ultimately eliminate the virus from circulation. COVID-19 patients present a wide range of clinical symptoms (e.g., cough, fever, and dyspnea) that are similar to influenza or other respiratory infections and thus cannot be used for accurate diagnosis [36, 37]. At present, accurate and rapid diagnosis has evolved as a tool for detecting this novel coronavirus (Table 1). Molecular-based approaches are the first-line methods to confirm suspected cases. Nucleic acid testing is the main technique for laboratory diagnosis. As with other emerging viruses, the development of methods to detect antibodies and viral antigens began after identification of the viral genome. In fact, the genomic sequence of SARS-CoV-2 was released to public databases on January 10, 2020 (GenBank accession No. MN908947), which facilitated the development of standardized laboratory PCR protocols for COVID-19 in January of 2020 [38]. This was followed by the launch of a range of commercial SARS-CoV-2 PCR-based assays in the last 3 months [39-45].
Diagnostics methods for COVID-19
| Method | Characteristics | Limitations | Refs |
|---|---|---|---|
| CT Scan | Available earlier; check severity of condition; check possible infection | Expensive; unable to distinguish from other viral pneumonias; hysteresis of abnormal CT imaging | [52-60, 70-72] |
| RT-qPCR | Specific, sensitive and simple quantitative assay, which greatly helps in the diagnosis of early infection | Costly and time consuming to perform; more prone to false negatives or low value | [38-42, 70, 73-75] |
| CRISPR-based detection | High sensitivity and specificity with efficiency and no requirement for elaborate instrumentation | Certain biological safety hazards brought by the retention and operation of patient samples | [50, 51, 76-78] |
| Antibody-detection | Fast, robust and easy to perform; requiring only a small amount of sample | Unable to detect the presence of infection during the early stage of disease; cross-reactivity | [61-64,79-82] |
| Antigen-detection | Low complexity; rapid; easy to perform | Best used to identify acute or early infection; more prone to false negatives | [68, 69,83-86] |
Reverse transcription quantitative PCR (RT-qPCR) is the most common and straightforward method for the detection of SARS-CoV-2 owing to its advantages as a specific, sensitive and simple quantitative assay, which greatly helps in the diagnosis of early infection. As of May 25, a total of 12 RT-qPCR tests to detect SARS-CoV-2 have been approved in China by the National Medical Products Administration (NMPA) [46]. Although RT-qPCR assays using respiratory samples provided sensitive and specific diagnostic tests at the initial phase of the outbreak, these RT-qPCR test kits have many limitations [47]. For example, this method is not efficient in rapidly screening a large number of individuals in places where thousands of people transit per hour. In addition, the performance of RT-qPCR depends on many factors, such as the sample type, stage of infection, skill of sample collection, and quality and consistency of the PCR assay being used [48, 49]. These problems lead to a noteworthy delay in early diagnosis and subsequent management and pose a serious challenge to providing timely life-support treatment and preventive quarantine. Notably, another powerful and promising tool involves clustered regularly interspaced short palindromic repeats (CRISPR) technology, which is being quickly deployed in the molecular diagnostics landscape. On May 6, Sherlock Biosciences received Emergency Use Authorization (EUA) from the US Food and Drug Administration for its Sherlock™ CRISPR SARS-CoV-2 kit for detection of the virus that causes COVID-19. Based on the SHERLOCK (Specific High-sensitivity Enzymatic Reporter unLOCKing) method, the kit works by programming a CRISPR molecule to detect the presence of a specific genetic signature for SARS-CoV-2 [50]. If the signature is found, the CRISPR enzyme generates a fluorescent glow. Furthermore, Mammoth Biosciences has developed a rapid (<40 min), easy-to-implement and accurate CRISPR-based assay for the detection of SARS-CoV-2 from respiratory swab RNA extracts called the SARS-CoV-2 DNA Endonuclease-Targeted CRISPR Trans Reporter (DETECTR) [51]. CRISPR-based diagnosis methods benefit from high sensitivity and specificity with efficiency and no requirement for elaborate instrumentation. As the assay uses similar specimen collection and nucleic acid extraction methods as the CDC and other RT-qPCR assays, it is subjected to the same potential limitations with regard to the availability of personal protective equipment, extraction kits and reagents. Therefore, many clinicians have proposed that computed tomography (CT) scans should be a necessary auxiliary diagnostic method [52-56]. To identify and quarantine patients at an early date, patients clinically diagnosed by chest CT with typical ground-glass lung opacities were also counted as confirmed cases beginning in early February [57-60]. However, CT scans also have some shortcomings, such as the hysteresis of abnormal CT imaging and indistinguishability from other viral pneumonia presentations. Accordingly, a combination of chest CT scans and repeated RT-qPCR tests may be helpful for individuals with a high clinical suspicion of SARS-CoV-2 infection but who are negative by RT-qPCR screening. Compared to the currently used nucleic acid detection and CT scans, other methods, such as virus antigen or serological antibody testing, are advantageous with fast turn-around times, high throughput and reduced workload for the detection of novel coronavirus infection. Currently, immunological identification technologies (point-of-care testing (POCT) of IgM/IgG and enzyme-linked immunosorbent assay (ELISA)) kits for SARS-CoV-2 have become available, with detection rates higher than those of nucleic acid detection approaches.
One type of rapid diagnostic test based on host antibody detection has become available more recently [61-64]. Antibodies are produced over days to weeks after infection with the virus. Antibody testing is easy to administer and only requires minimal training. It may be helpful as a complement to nucleic acid testing for the diagnosis of suspected COVID-19 cases with negative RT-qPCR results as well as for the identification of asymptomatic infections [65]. Confirming suspected cases as early as possible with the benefit of antibody testing might reduce exposure to risk factors during repeated sampling and save valuable RT-qPCR tests. Due to urgency and demand, many antibody test reagents are rapidly being developed and made available on the market to assist in the diagnosis of SARS-CoV-2 infection, including kits for testing total Abs, IgM antibody or IgG antibody, by chemiluminescence, ELISA or colloidal gold methods. On March 6, an anti-SARS-CoV-2 antibody test kit based on chemiluminescent particle immunoassay jointly developed by our team and Beijing Wantai Biological Pharmacy Enterprise Co., Ltd. was approved for listing by the State Drug Administration for emergency approval and is the best performing commercial kit according to the latest WHO validation data [66]. The kit is based on the double-antigen sandwich principle that detects total antibodies (including IgM, IgG, IgA and other antibody types). Specific testing for SARS-CoV-2 has great significance for public health and clinical practice, and evidence from studies further demonstrate that RT-qPCR detection combined with serological testing enhances diagnosis sensitivity while maintaining high specificity [67].
Another type of rapid diagnostic test (RDT) that detects the presence of viral antigens expressed by SARS-CoV-2 virus in a respiratory tract sample is of low complexity and may provide results typically within 30 minutes [68,69]. The antigen(s) detected are expressed only when the virus is actively replicating; thus, such tests are best used to identify acute or early infection. Monoclonal antibodies (mAbs) against the nucleocapsid protein of SARS-CoV-2 have been developed and might constitute the basis of a future rapid antigen detection test. As of the time of this review, at least 5 antigen-detection RDTs were under evaluation (https://www.finddx.org/covid-19/sarscov2-eval-immuno/), though not all are CE Marking approved or widely available.
Antiviral therapy
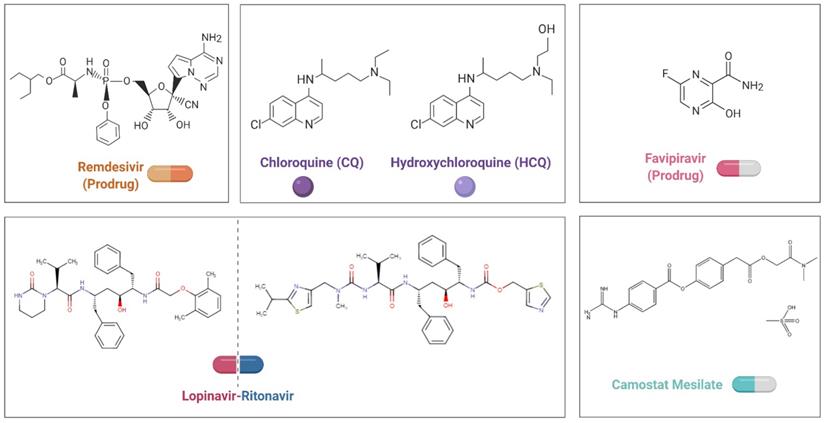
To date, there are no specific antiviral therapies or vaccines for COVID-19 available for human use. Given the urgency of treating these patients, rather than taking months to years to develop and test compounds from scratch, researchers are seeking to repurpose drugs that are already approved for other diseases and have acceptable safety profiles. Small-molecule agents approved for other human diseases can exert their antiviral effect through multiple mechanisms, including blocking viral entry, inhibiting a virally encoded enzyme, targeting a host factor required for replication, or blocking virus particle formation [87]. Notably, some antivirals and antimalarials have shown promising abilities to treat patients and reduce the danger of COVID-19 (Figure 2, Table 2).
Chemical structural formula of candidate therapeutic agents.
List of candidate therapeutic agents for COVID-19 treatment
| Candidate therapeutic | Manufacturer | Target | Mechanism of action | Infectious diseases | Status of clinical trials | Refs. |
|---|---|---|---|---|---|---|
| Remdesivir (GS-5734) | Gilead | RdRp | Terminates the nonobligate chain | Ebola, SARS-CoV-2 | • Phase 2 | phase 3 for Ebola (NCT03719586) • Phase 3 for SARS-CoV-2 (NCT04292899, NCT04292730, NCT04280705, NCT04321616, etc.) | [88-91, 107, 108] |
| Chloroquine (CQ) | Sanofi | Endosomal acidification | A lysosomotropic base that appears to disrupt intracellular trafficking and viral fusion events | SARS-CoV, MERS-CoV, SARS-CoV-2 | •Approved for malaria and certain amoeba infections • Under clinical trial for SARS-CoV-2 Phase 2 (NCT04323527, NCT04328493, NCT04344951, NCT04342650, etc.) Phase 3 (NCT04324463, NCT04341727, etc.) Phase 4 (NCT04316377 NCT04362332, etc.) | [92, 104, 109] |
| Hydroxychloroquine (HCQ) | Sanofi | Endosomal acidification | Hydroxychloroquine shares the same mechanism of action as chloroquine | SARS-CoV, MERS-CoV, SARS-CoV-2 | Under clinical trial for SARS-CoV-2 Phase 2 (NCT04329832, NCT04335552, etc.) Phase 3 (NCT04345692, NCT04342221, etc.) Phase 4 (NCT04362332, NCT04355026, etc.) | [110-112] |
| Lopinavir-ritonavir | Abbott | 3CL protease | Inhibits 3CLpro | HIV, SARS-CoV-2 | • Under clinical trials for SARS • Under clinical trial for SARS-CoV-2 Phase 2 (NCT04307693, NCT04328012, etc.) Phase 3 (NCT04328285, NCT04364022, etc.) Phase 4 (NCT04255017, NCT04350684, etc.) | [99, 103, 104] |
| Favipiravir (T-705) | Toyama | RdRp | Inhibits RdRp | Influenza, SARS-CoV-2 | • Approved for influenza in Japan • Under clinical trial for SARS-CoV-2 Phase 2 (NCT04358549, NCT04351295, etc.) Phase 3 (NCT04349241, NCT04303299, etc.) Phase 4 (NCT04359615) | [105, 113-115] |
| Camostat mesilate (FoipanTM) | Ono Pharmaceutical | TMPRSS2 | Inhibits serine protease | SARS-CoV-2 | Under clinical trial for SARS-CoV-2 Phase 2 (NCT04374019) Phase 3 (NCT04355052) | [106] |
Remdesivir (GS-5734) is a nucleotide analog prodrug that shuts down viral replication by inhibiting a key viral enzyme, RNA polymerase. A recent study reported that remdesivir potently blocks SARS-CoV-2 infection (EC50 = 0.77 μM) in Vero E6 cells [88], and the first patient with confirmed SARS-CoV-2 infection in the United States recovered after receiving intravenous remdesivir in January [89]. On April 10, a study reported clinical outcomes for compassionate use of remdesivir in a small cohort of patients with severe COVID-19. Specifically, clinical improvement was observed in 36 of 53 patients (68%) [90]. However, a trial (ClinicalTrials.gov: NCT04257656) found that compared with placebo, intravenous remdesivir did not provide significant clinical or antiviral effects in patients with serious COVID-19. Ongoing trials with larger sample sizes will provide information on the efficacy of remdesivir for COVID-19 [91]. On May 1, the US Food and Drug Administration granted emergency use authorization for remdesivir to treat COVID-19 in adults and children hospitalized with severe disease.
Chloroquine (CQ), a 9-aminoquinoline that is a widely used antimalarial and autoimmune disease drug, has recently been reported as a potential broad-spectrum antiviral drug [92]. CQ inhibits virus infection by increasing the endosomal pH required for virus/cell fusion, and a recent study reported its activity against COVID-19 (EC50=1.13 μM) in Vero E6 cells [88, 93]. Patients with COVID-19 are being recruited in randomized trials (e.g., ClinicalTrials.gov: NCT04316377, NCT04323527) to determine the safety and efficacy of CQ. Although a few clinical trials have suggested some beneficial effects of CQ for COVID-19, most of the reported data are preliminary [94, 95]. Furthermore, it is likely that the risk of adverse reactions to CQ will hamper the successful treatment of patients with COVID-19.
Hydroxychloroquine (HCQ), which exhibits an antiviral effect highly similar to that of chloroquine, might be a better therapeutic approach. Studies have indicated that CQ shows antagonism against SARS-CoV-2 in vitro. Chen et al. partially confirmed the potential of HCQ in patients with COVID-19 (ClinicalTrials.gov: ChiCTR2000029559) [96]. Although positive results from small studies have been reported for hydroxychloroquine, accumulating trial and observational evidence has prompted skepticism regarding whether there are any meaningful clinical benefits for patients with COVID-19. Indeed, a study from France did not support the use of HCQ in patients hospitalized with COVID-19 who require oxygen [97]. Additionally, a randomized clinical trial (ClinicalTrials.gov: ChiCTR2000029868) from China shows that hospitalized patients with mild to moderate persistent COVID-19 who received HCQ did not eliminate the virus more quickly than those receiving standard care. Moreover, adverse events were higher among HCQ recipients than nonrecipients [98]. Nonetheless, randomized clinical trials are needed to provide definitive causal evidence while offering an opportunity to continuously optimize the treatment plan.
The lopinavir-ritonavir combination, which targets an enzyme that cleaves a long protein chain during the assembly of new viruses, can inhibit HIV protease [99,100]. The combination has been evaluated in patients with SARS and MERS, though the results are ambiguous [101-104]. Clinical trials (e.g., ClinicalTrials.gov: NCT04255017, NCT04307693) with COVID-19 have been initiated to test the potency of the combination. Recently, a randomized, controlled, open-label trial (ClinicalTrials.gov: ChiCTR2000029308) involving 199 patients with COVID-19 was initiated, but no benefit beyond standard care was observed with lopinavir-ritonavir treatment. Future trials including patients with severe illness may help to confirm or exclude the possibility of a treatment benefit.
Favipiravir (T-705), a purine nucleic acid analog approved for the treatment of influenza, can effectively inhibit the RNA-dependent RNA polymerase (RdRp) of RNA viruses such as influenza, Ebola and norovirus [105]. Studies in Vero E6 cells have suggested that favipiravir can cripple the SARS-CoV-2 virus (EC50 = 61.88 μM) [88], and patients with COVID-19 are being recruited in randomized trials to evaluate the efficacy of favipiravir plus other antivirals (e.g., ClinicalTrials.gov: ChiCTR2000029600, ChiCTR2000029544).
Camostat mesylate is a potent serine protease inhibitor that has been used primarily to treat pancreatitis and some types of cancer [106]. SARS-CoV-2 uses ACE2 for entry and the serine protease TMPRSS2 for S protein priming. Utilizing research on SARS-CoV and the closely related SARS-CoV-2 cell entry mechanism, it has been demonstrated that camostat mesylate can block SARS-CoV-2 infection of lung cells [28]. Patients with COVID-19 are being recruited in randomized trials (ClinicalTrials.gov: NCT04374019, NCT04355052) to determine whether this drug reduces SARS-COV-2 viral load in early COVID-19 disease.
Antibody-based Interventions
The antibody-mediated humoral response plays crucial roles in controlling viral infection. NAbs, which can reduce viral infectivity by binding to the surface epitopes of viral particles and thereby blocking entry of the virus into host cells, are a promising approach to prevent and treat virus infection.
Passive antibody therapy, through transfusion of convalescent plasma from patients who have recovered from viral infections, can also be used as a treatment. Importantly, this therapy offers the only short-term strategy of conferring immediate immunity to infected patients. The antibodies present in convalescent plasma can specifically bind to a given pathogen (e.g., an infectious virus), directly neutralizing its infectivity. Other antibody-mediated pathways, such as antibody-dependent cellular cytotoxicity, complement activation, and/or phagocytosis, may also contribute to their therapeutic effect. There are numerous examples in which convalescent plasma therapy has been successfully applied to treat patients, as follows: SARS; MERS; H1N1 (pandemic 2009 influenza A); H5N1 (avian influenza A); several hemorrhagic fevers, such as Ebola; and other viral infections [116-120]. As no specific therapeutic agents or vaccines are available for COVID-19, this therapy is the only strategy that is immediately available for use to prevent and treat a novel, emerging infectious disease such as SARS-CoV-2 infection [121, 122]. Shen et al reported findings from a preliminary study of 5 severely ill patients with COVID-19 who were given plasma from recovered individuals [123]. In this case series, the clinical status of all patients improved by approximately 7 days after transfusion. In addition, the patients' specific NAb titers increased following the transfusion, and respiratory samples became negative within 12 days. Although the limited case size and study design preclude a definitive statement about the potential efficacy of this treatment, the results provide some evidence to support the possibility of evaluating this therapy in more rigorous investigations involving patients with COVID-19.
Moreover, sophisticated technologies are devoted to identifying strongly neutralizing mAbs that can be produced in large quantities. In coronaviruses, the viral component most frequently targeted by NAbs is the surface-located envelope S protein [124-126]. To this end, several potently neutralizing human mAbs against SARS-CoV-2 have been developed using various approaches, including single-cell culturing methods of memory B cells isolated from SARS-CoV-2 patients, fusing B cells with an appropriate partner to produce hybridomas, selection of antibodies from phage-display libraries of human antibody fragments, and hybridoma fusion of B cells from immunized transgenic mice encoding human variable immunoglobulin domains [127-131]. All of these antibodies were selected based on their in vitro neutralizing capacity, and most of them target the SARS-CoV-2 RBD.
Ju et al. isolated 206 mAbs specific for the SARS-CoV-2 RBD derived from single B cells of eight SARS-CoV-2-infected individuals [130]. Using bioinformatic and biologic characterization, several mAbs with potent binding and neutralizing activity against pseudovirus encoding the S protein and live SARS-CoV-2 have been identified. In particular, the most potent antibodies, P2B-2F6 and P2C-1F11, out-competed ACE2 with close to 100% efficiency, indicating that blocking the RBD and ACE2 interaction is a useful surrogate for antibody neutralization. The results indicate that the humoral immune response is viral species specific and diverse and can produce potent NAbs. Wu et al. isolated 4 mAbs specific to SARS-CoV-2 RBD from a convalescent patient [131], and these antibodies effectively neutralized the SARS-CoV-2, with two of them exhibiting additive inhibition effects. Moreover, a therapeutic study in a mouse model validated that these antibodies can reduce virus titers in infected lungs. These antibodies will serve as the most promising candidates for the development of prophylactic and therapeutic interventions against SARS-CoV-2.
CD147-spike protein, a receptor on host cells, is a novel route for SARS-CoV-2 invasion [132]. Meplazumab, an anti-CD147 antibody, effectively inhibited SARS-CoV-2 replication and virus-induced cytopathic effects in Vero E6 cells; it also blocks CD147, the receptor of the proinflammatory factor CyPA, thus attenuating inflammation [132,133]. Based on this evidence, the team designed an open-label, concurrent controlled trial (NCT04275245) to assess the efficacy and safety of meplazumab in patients with COVID-19 pneumonia. The data demonstrated that meplazumab efficiently improved the recovery of the patients, with a favorable safety profile. However, the results were limited by the absence of a parallel control and a small sample size. Therefore, confirmation from a larger scale and randomized, double-blind trial is required to fully assess the efficacy of meplazumab in patients with COVID-19 pneumonia.
The S proteins of SARS-CoV-2 and SARS-CoV, which are structurally very similar, have an amino acid sequence identity of approximately 77% and commonly bind ACE2 as a host receptor [134-136]. To reduce the time that is required to develop SARS-CoV-2-neutralizing antibodies de novo, researchers have assessed available SARS-CoV antibodies for cross-neutralization of SARS-CoV-2, given the high degree of sequence similarity. For example, a SARS-CoV RBD-specific human NAb (CR3022) previously isolated from a convalescent SARS patient is able to bind the SARS-CoV-2 RBD with high affinity (Kd of 6.3 nM) and recognize an epitope on the RBD that does not overlap with the ACE2-binding site [137]. However, CR3022 does not neutralize SARS-CoV-2 in vitro, and it is possible that this epitope can confer in vivo protection [138]. Further studies will require suitable animal models, which have yet to be established. Another anti-SARS-CoV antibody, 47D11, was isolated from transgenic H2L2 mice encoding the human immunoglobulin variable regions to generate antibodies that neutralize SARS-CoV-2 (and SARS-CoV). 47D11 binds a conserved epitope on the spike RBD, explaining its ability to cross-neutralize SARS-CoV and SARS-CoV-2, with diverse mechanisms of action that are independent of receptor binding inhibition [139]. This antibody offers potential for the prevention and treatment of COVID-19.
Although antibodies are generally beneficial and protective, in rare cases, pathogen-specific antibodies could promote pathology, resulting in a phenomenon known as antibody-dependent enhancement (ADE) [140, 141]. The ADE phenomenon is documented for dengue virus and other viruses [142, 143]. However, the neutralizing antibodies can be engineered of Fc region with molecular precision to avoid the potential ADE.
The current pipeline for SARS-CoV-2 vaccines
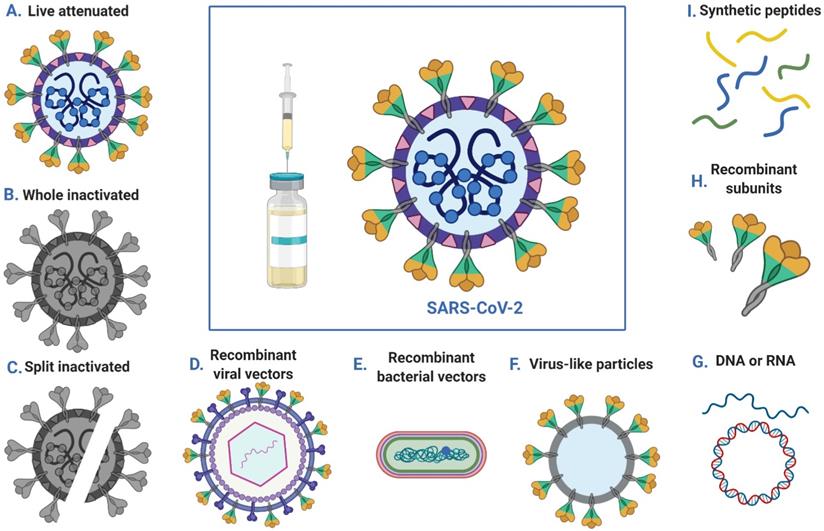
Vaccination can be used to prevent infection or to reduce disease severity, viral shedding and thereby transmission, thus helping to control SARS-COV-2 outbreaks. Multiple strategies have been employed to generate SARS-COV-2 vaccines, including DNA- and RNA-based vaccines, viral vector vaccines, inactivated virus vaccines, live-attenuated virus vaccines and recombinant protein vaccines (Figure 3). According to reports from the WHO, there are more than 100 vaccines against SARS-CoV-2 at various stages of development. Specifically, ten candidate vaccines are already in clinical evaluation (Table 3).
Approaches to SARS-CoV-2 vaccine development.
Clinical-phase vaccine candidates for COVID-19
| Vaccine strategy | Candidate | Lead developer | Vaccine characteristics | Status | Same platform for noncoronavirus candidates |
|---|---|---|---|---|---|
| RNA vaccines | mRNA-1273 | Moderna/NIAID | Lipid nanoparticles containing the mRNA vaccine encoding the SARS-CoV-2 S protein | Phase 2 (IND accepted) Phase 1 NCT04283461 | Multiple candidates |
| BNT162 | BioNTech/Fosun Pharma/Pfizer | Lipid nanoparticles containing the mRNA vaccine encoding the SARS-CoV-2 S protein | Phase 1/22020-001038-36 NCT04368728 | - | |
| Viral vector-based vaccines | Ad5-nCoV | CanSino Biological Inc./Beijing Institute of Biotechnology | Nonreplicating adenovirus type 5 vector carrying the gene for the SARS-CoV-2 S protein | Phase 2 ChiCTR2000031781, phase 1 ChiCTR2000030906 | Ebola |
| ChAdOx1 nCoV-19 | University of Oxford | ChAdOx1 construct (an adenovirus vaccine vector) carrying the gene for the SARS-CoV-2 S protein | Phase 1/2 NCT04324606 | MERS, influenza, TB, Chikungunya, Zika, MenB, plague | |
| DNA vaccines | INO-4800 | Inovio Pharmaceuticals | DNA plasmid encoding the SARS-CoV-2 S protein delivered by electroporation | Phase 1 NCT04336410 | Lassa, Nipah, HIV, filovirus, HPV, cancer indications, Zika, hepatitis B |
| Inactivated virus vaccines | Inactivated | Beijing Institute of Biological Products/Wuhan Institute of Biological Products | Virions inactivated with chemicals or radiation, with or without adjuvant, rendering the viral genome noninfectious while maintaining the virion structure, which is antigenically similar to the live virus | Phase 1/2 ChiCTR2000032459 | - |
| Inactivated | Wuhan Institute of Biological Products/Sinopharm | Phase 1/2 ChiCTR2000031809 | - | ||
| Inactivated + alum | Sinovac | Phase 1/2 NCT04383574 NCT04352608 | SARS | ||
| Inactivated | Institute of Medical Biology, Chinese Academy of Medical Sciences | Phase 1 | - | ||
| Protein subunit | NVX-CoV2373 | Novavax | Full length recombinant SARS-CoV-2 glycoprotein nanoparticle vaccine adjuvanted with Martrix MTM adjuvant | Phase 1/2 NCT04368988 | RSV, CCHF, HPV, VZV, EBOV |
Source: WHO.
mRNA vaccines are an emerging new vaccine modality in which patients are treated with mRNA oligonucleotides that encode either an antigen of interest or an antigen as well as its viral replication machinery [144]. The potential advantages of the mRNA approach to prophylactic vaccines include the ability to mimic natural infection to stimulate a more potent immune response, combining multiple mRNAs into a single vaccine, rapid discovery to respond to emerging pandemic threats and manufacturing agility derived from the platform nature of mRNA vaccine design and production [145]. mRNA vaccines have elicited potent immunity against infectious disease targets in animal models of infection with influenza virus, Zika virus, rabies virus and others, especially in recent years, using lipid-encapsulated or naked forms of sequence-optimized mRNA [146-148]. Moderna and the Vaccine Research Center, a unit of the National Institute of Allergy and Infectious Diseases (NIAID), have collaborated to develop mRNA vaccines (mRNA-1273) for the novel coronavirus. mRNA-1273 encodes a prefusion stabilized form of the SARS-CoV-2 S protein. The vaccine consists of a mRNA drug substance that is manufactured into lipid nanoparticles (LNPs) composed of the proprietary ionizable lipid SM-102 and 3 commercially available lipids: cholesterol, DSPC, and PEG2000 DMG. Extensive basic research into RNA and lipid and polymer biochemistry has enabled translating mRNA vaccines into clinical trials. On May 18, Moderna announced positive interim phase 1 results for mRNA-1273 against SARS-CoV-2 [149]. After two doses, mRNA-1273 elicited neutralizing antibody titer levels at or above the levels in convalescent sera in all initial participants. To continue progress on this potential vaccine during the ongoing global public health emergency, Moderna intends to work with the FDA and other government and nongovernment organizations to be ready for phase 2 and any subsequent trials, which are anticipated to include a large number of subjects and will seek to generate additional safety and immunogenicity data. Furthermore, BioNTech and Pfizer are jointly developing another mRNA vaccine (BNT162), which has been approved for a phase 1/2 clinical trial to determine the optimal dose for further studies as well as to evaluate the safety and immunogenicity of the vaccine. The trial is the first clinical trial of a COVID-19 vaccine candidate to start in Germany and is part of a global development program.
DNA vaccines are appropriate for emerging infectious diseases because they allow for the rapid design of multiple candidates for novel antigens [150]. In addition, they are directly injected or otherwise inoculated into recipients. Recently, various DNA vaccine platforms have been developed to improve the efficacy of vaccines by using electroporation (EP) to deliver plasmids and adding adjuvants to enhance immune responses. Inovio Pharmaceuticals was the first to advance its vaccine (INO-4700) against MERS-CoV [151], a related coronavirus, into evaluation in humans, and this company has developed a DNA vaccine (INO-4800) to prevent infection by SARS-CoV-2. INO-4800 induces T cell activation by delivering DNA plasmids that express the SARS-CoV-2 S protein. The effects based on measuring functional antibodies and antigen-specific T cell responses in multiple animal models are particularly encouraging [152]. Inovio has also initiated an open-label trial (NCT04336410) to evaluate the safety, tolerability and immunological profile of INO-4800 administered by intradermal (ID) injection followed by EP using a CELLECTRA® 2000 device in healthy adult volunteers.
Viral vector vaccines, which function as viral gene expression and delivery systems, rely on a host viral genome, and an unrelated viral genome lacking packaging elements is engineered to encode antigenic components of the virus of interest to elicit an immune response. Because viral vector vaccines persist in the host as genetic material, they directly infect antigen-presenting cells and have strong immunogenicity, efficiently inducing B cell- and T cell-mediated immune responses. Furthermore, viral vector vaccines can result in high-titer NAbs. Several different virus vectors have been developed as SARS-CoV-2 vaccines. CanSinoBIO and the Beijing Institute of Biotechnology (BIB) have collaborated to develop an adenovirus type 5 vector-based novel coronavirus vaccine (“Ad5-nCoV”) that encodes the full-length S protein of SARS-CoV-2. Ad5-nCoV protects against SARS-COV-2 infection by relying on a recombinant replication-defective human adenovirus type-5 vector to induce the immune response. The vaccine candidate is built upon CanSinoBIO's adenovirus-based viral vector vaccine technology platform, which has also been successfully applied to develop a globally innovative vaccine against Ebola virus infection. The results from preclinical studies of Ad5-nCoV show that the vaccine candidate can induce a strong immune response in animal models, and preclinical animal safety studies demonstrate a good safety profile. This is currently the first novel coronavirus vaccine for COVID-19 that has progressed to this stage in China. A first-in-human trial showed that the Ad5-nCoV vaccine is tolerable and immunogenic in healthy adults [153], with specific humoral responses against SARS-CoV-2 peaking at day 28 postvaccination and rapid specific T-cell responses noted at day 14. There is potential for further investigation of the Ad5-nCoV vaccine for the control of the COVID-19 outbreak. Furthermore, University of Oxford researchers have begun testing a new COVID-19 vaccine called ChAdOx1 nCoV-19; it is derived from a virus (ChAdOx1) that is an attenuated version of a common cold virus (adenovirus) causing infections in chimpanzees, but it has been genetically modified such that it is unable to proliferate in humans. The vaccine is based on an adenovirus vector (ChAdOx1) and the SARS-CoV-2 S protein. The results showed that a single vaccination with ChAdOx1 nCoV-19 is effective for preventing lung damage upon high-dose challenge with SARS-CoV-2; however, all of the monkeys vaccinated with ChAdOx1 nCoV-19 became infected when challenged, and there was no difference in the amount of viral RNA detected from nasal secretions in the vaccinated and unvaccinated monkeys [154]. As of May 13, more than 1,000 volunteers participated in the trial, which aims to assess whether healthy people can be protected from COVID-19. It will also provide valuable information on the safety aspects of the vaccine and its ability to generate good immune responses against the virus. In addition to adenovirus, our team and the University of Hong Kong have used influenza virus vectors for the development of candidate SARS-CoV-2 vaccines that are in preclinical phases.
Inactivated virus vaccines use chemicals (such as formalin, β-propiolactone and diethylpyrocarbonate) or heat to render the viral genome noninfectious while maintaining the virion structure, thus preserving antigenicity but eliminating the potential to cause productive infection. Thus, in theory, inactivated virus vaccines are easily prepared and antigenically similar to live viruses. Such vaccines have been found to be effective and safe for the prevention of diseases caused by viruses such as poliovirus and influenza virus [155, 156]. Several institutes are developing inactivated virus COVID-19 vaccines. Recently, the Beijing Institute of Biological Products and Wuhan Institute of Biological Products have developed an inactivated novel coronavirus pneumonia vaccine. Healthy people at different ages are being recruited in randomized, double-blind, placebo parallel-controlled phase I/II clinical trials (ClinicalTrials.gov: ChiCTR2000032459, ChiCTR2000031809) to explore the safety and immunogenicity of this COVID-19 vaccine. Drawing on previous experience in SARS vaccine development, Sinovac has developed another inactivated vaccine. On April 13, 2020, a randomized, double-blinded, placebo-controlled phase I clinical trial (ClinicalTrials.gov: NCT04352608) of an inactivated SARS-CoV-2 vaccine was approved by China's NMPA to evaluate the safety and preliminary immunogenicity in healthy adults, with two different dosages of the vaccine candidate [157].
Other vaccine approaches include live-attenuated vaccines that are produced by reducing or eliminating the virulence of a live virus, typically using chemical-driven or site-directed mutagenesis. Thus, the virus is capable of productive infection, but the resulting disease is either diminished or eliminated. Live-attenuated vaccines can elicit both innate and adaptive immune responses, and protection can be life-long. Additionally, their production is inexpensive. Codagenix and the Serum Institute of India are using viral deoptimization to synthesize "rationally designed", live-attenuated vaccines against the emergent coronavirus. Recombinant-protein-based vaccines are advantageous over other types of vaccines in that they are safe and have few side effects, inducing the immune system without the introduction of infectious viruses. One example is the NVX-CoV2373 vaccine developed by Novavax, which is in the clinical stage [158]. It was created using recombinant nanoparticle technology to generate antigen derived from the S protein and contains Matrix-M™ adjuvant to enhance the immune response and stimulate high levels of neutralizing antibodies. A trial of Clover's S-trimer vaccine developed by Clover Biopharmaceuticals will be recruiting healthy adults in Australia, and this vaccine will be widely available upon confirmation of its safety and efficacy. In addition, LV-SMENP-DC and pathogen-specific aAPC vaccines from Shenzhen Geno-Immune Medical Institute are in phase 1 clinical trials to evaluate safety and immune reactivity [159,160].
Conclusions and Prospects
Despite tremendous efforts to prevent the spread of SARS-CoV-2 worldwide, the high mortality and person-to-person transmission pose a significant threat to global public health. Diagnostics are important for dealing with virus outbreaks because they can enable healthcare workers to direct efforts to patients with COVID-19. Remarkably, various diagnostic kits to test for COVID-19 are available. Several drugs have demonstrated in vitro activity against the SARS-CoV-2 virus or potential clinical benefits in observational or small, nonrandomized studies. Additionally, researchers are evaluating several different technologies, and more than 100 vaccines are under development against SARS-CoV-2 worldwide. Specifically, ten candidate vaccines have already been administered to volunteers in safety trials; others have started testing in animals. However, as the development of vaccines for human use can take years, their development will likely come too late to impact the first wave of the pandemic. These challenges will remain in the long term. Clearly, there is an urgent need for effective antiviral therapy and vaccines for this emerging global threat.
There are a series of patient-associated and virologic factors that pose major clinical challenges to the development of anti-SARS-CoV-2 drugs. SARS-CoV-2 is a virus with diversity that becomes mutated, and novel SARS-CoV-2 mutants are likely to emerge repeatedly at unpredictable times [161,162]. Therefore, most drugs that specifically target the existing SARS-CoV-2 may not be effective against a mutant SARS-CoV-2. The lag time between mutant emergence and the development of new prophylactic treatments is of concern. Thus, there is an urgent need for the development of new, broad-spectrum drugs that target conserved sites to prepare for future outbreaks involving mutants. Recent studies have reported that lipopeptide EK1C4 shows exceptional promise to be developed as the first pan-CoV fusion inhibitor-based antiviral therapeutic or prophylactic for treatment or prevention of infection by the currently circulating SARS-CoV-2 and mutants or emerging SARS-related CoVs [163-165]. In addition, combining the advanced knowledge of bioinformatics and immunology to screen and confirm the conserved dominant epitopes of SARS-CoV-2, it will be helpful to develop therapeutic antibodies with broad-spectrum neutralization activity and vaccines that can induce a broad-spectrum antiviral immune response. Currently, there are a limited number of animal models available for infections caused by SARS-CoV-2. Normally, antiviral therapy or vaccines enter into human trials after tests for safety and effectiveness in appropriate animal models. However, the virus does not grow in wild-type mice and induces only mild disease in transgenic animals expressing human ACE2 [166,167]. The limited availability of mouse-adapted virus strains and ACE2 transgenic mice remains a major obstacle to testing anti-SARS-CoV-2 drugs or vaccines. Other potential animal models include ferrets and nonhuman primates (NHPs), for which pathogenicity studies are ongoing. In addition, understanding how the immune system interacts with the pathogen and the vaccine itself is crucial for avoiding pitfalls, such as ADE phenomenon, a process in which a virus leverages antibodies to promote infection.
The viral load curve of COVID-19 is similar to that of influenza, and the viral load is very high at the time of presentation [168,169]. Experience from the treatment of influenza patients, in this case of high-viral load patients, suggested that the combination of multiple antiviral drugs is more effective than single drug treatments [170-172]. Indeed, the combination of appropriate antiviral drugs might rapidly suppress the high initial viral load, improve the clinical parameters, and reduce the risk to medical staff by controlling the spread of the virus. A multicenter randomized open-label phase 2 trial (ChiCTR2000029308) showed that triple antiviral therapy with interferon beta-1b, lopinavir-ritonavir, and ribavirin was safe and superior to lopinavir-ritonavir alone in alleviating symptoms, suppressing shedding of SARS-CoV-2, and facilitating discharge of patients with mild to moderate COVID-19 [168]. There are also many other studies investigating combination therapy; these smaller clinical studies examining virologic endpoints may be sufficient to identify which combinations can be used for large studies in risk groups or hospitalized patients. For combinations involving immunomodulatory interventions, the challenges may be great because the goal is disease amelioration by regulating the host complex responses. Regardless, it is certain that as our continuous in-depth understanding of the immune response dynamics and the disease pathogenesis of SARS-CoV-2 deepens, combination therapy involving immunomodulation, vaccines or other immunotherapies will have great potential.
The outbreak has emphasized the urgent need for renewed efforts to develop specific antiviral therapies or vaccines to combat SARS-CoV-2. We need well-developed emergency plans that allow us to develop, test, produce, and distribute agents or vaccines as quickly as possible. Such endeavors would require tight coordination among governments, regulatory agencies, pharmaceutical companies, and the WHO, as well as novel approaches and clinical trial design.
Acknowledgements
This work was supported by the Xiamen Science and Technology Major Project (3502Z2020YJ02), the Major State Basic Research Development Program of China (2017YFA0205201), the National Natural Science Foundation of China (NSFC) (81925019, 81422023, and U1705281), the Fundamental Research Funds for the Central Universities (20720190088 and 20720200019), and the Program for New Century Excellent Talents in University, China (NCET-13-0502).
Competing Interests
The authors have declared that no competing interest exists.
References
1. Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;579:270-3
2. Wu F, Zhao S, Yu B, Chen YM, Wang W, Song ZG. et al. A new coronavirus associated with human respiratory disease in China. Nature. 2020;579:265-9
3. Zhu N, Zhang D, Wang W, Li X, Yang B, Song J. et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N Engl J Med. 2020;382:727-33
4. Coronaviridae Study Group of the International Committee on Taxonomy of V. The species Severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat Microbiol. 2020;5:536-44
5. Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y. et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet. 2020;395:497-506
6. Xu Z, Shi L, Wang Y, Zhang J, Huang L, Zhang C. et al. Pathological findings of COVID-19 associated with acute respiratory distress syndrome. Lancet Respir Med. 2020;8:420-2
7. Vetter P, Vu DL, L'Huillier AG, Schibler M, Kaiser L, Jacquerioz F. Clinical features of covid-19. BMJ. 2020;369:m1470
8. Kim GU, Kim MJ, Ra SH, Lee J, Bae S, Jung J. et al. Clinical characteristics of asymptomatic and symptomatic patients with mild COVID-19. Clin Microbiol Infect. 2020 S1198-743X (20) 30268-8
9. Weiss SR, Navas-Martin S. Coronavirus pathogenesis and the emerging pathogen severe acute respiratory syndrome coronavirus. Microbiol Mol Biol Rev. 2005;69:635-64
10. Woo PC, Lau SK, Huang Y, Yuen KY. Coronavirus diversity, phylogeny and interspecies jumping. Exp Biol Med (Maywood). 2009;234:1117-27
11. Malik YA. Properties of Coronavirus and SARS-CoV-2. Malays J Pathol. 2020;42:3-11
12. Su S, Wong G, Shi W, Liu J, Lai ACK, Zhou J. et al. Epidemiology, Genetic Recombination, and Pathogenesis of Coronaviruses. Trends Microbiol. 2016;24:490-502
13. Forni D, Cagliani R, Clerici M, Sironi M. Molecular Evolution of Human Coronavirus Genomes. Trends Microbiol. 2017;25:35-48
14. Drosten C, Gunther S, Preiser W, van der Werf S, Brodt HR, Becker S. et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N Engl J Med. 2003;348:1967-76
15. Ksiazek TG, Erdman D, Goldsmith CS, Zaki SR, Peret T, Emery S. et al. A novel coronavirus associated with severe acute respiratory syndrome. N Engl J Med. 2003;348:1953-66
16. Zhao Z, Zhang F, Xu M, Huang K, Zhong W, Cai W. et al. Description and clinical treatment of an early outbreak of severe acute respiratory syndrome (SARS) in Guangzhou, PR China. J Med Microbiol. 2003;52:715-20
17. Zaki AM, van Boheemen S, Bestebroer TM, Osterhaus AD, Fouchier RA. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med. 2012;367:1814-20
18. Sousou J. Middle East Respiratory Syndrome Coronavirus: What Do We Know? Jnp-J Nurse Pract. 2015;11:131-4
19. de Wit E, van Doremalen N, Falzarano D, Munster VJ. SARS and MERS: recent insights into emerging coronaviruses. Nat Rev Microbiol. 2016;14:523-34
20. WHO. Summary of probable SARS cases with onset of illness from 1 November 2002 to 31 July 2003. Revised 26 September 2003. http://www.who.int/
21. WHO. MERS situation update, January 2020. http://www.who.int/
22. Li G, De Clercq E. Therapeutic options for the 2019 novel coronavirus (2019-nCoV). Nat Rev Drug Discov. 2020;19:149-50
23. Walls AC, Park YJ, Tortorici MA, Wall A, McGuire AT, Veesler D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell. 2020;181:281-92 e6
24. Lu R, Zhao X, Li J, Niu P, Yang B, Wu H. et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395:565-74
25. Du L, He Y, Zhou Y, Liu S, Zheng BJ, Jiang S. The spike protein of SARS-CoV-a target for vaccine and therapeutic development. Nat Rev Microbiol. 2009;7:226-36
26. Du L, Yang Y, Zhou Y, Lu L, Li F, Jiang S. MERS-CoV spike protein: a key target for antivirals. Expert Opin Ther Targets. 2017;21:131-43
27. Datta PK, Liu F, Fischer T, Rappaport J, Qin X. SARS-CoV-2 pandemic and research gaps: Understanding SARS-CoV-2 interaction with the ACE2 receptor and implications for therapy. Theranostics. 2020 doi:10.7150/thno.48076; in press
28. Hoffmann M, Kleine-Weber H, Schroeder S, Kruger N, Herrler T, Erichsen S. et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell. 2020;181:281-80 e8
29. Ou X, Liu Y, Lei X, Li P, Mi D, Ren L. et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun. 2020;11:1620
30. Tai W, He L, Zhang X, Pu J, Voronin D, Jiang S. et al. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol Immunol. 2020;17(6):613-620
31. Letko M, Marzi A, Munster V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses. Nat Microbiol. 2020;5:562-9
32. Chen Y, Liu Q, Guo D. Emerging coronaviruses: Genome structure, replication, and pathogenesis. J Med Virol. 2020;92:418-23
33. Fung TS, Liu DX. Coronavirus infection, ER stress, apoptosis and innate immunity. Front Microbiol. 2014;5:296
34. Lan J, Ge J, Yu J, Shan S, Zhou H, Fan S. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature. 2020;581(7807):215-220
35. Raimondi MT, Donnaloja F, Barzaghini B, Bocconi A, Conci C, Parodi V. et al. Bioengineering tools to speed up the discovery and preclinical testing of vaccines for SARS-CoV-2 and therapeutic agents for COVID-19. Theranostics. 2020;10(16):7034-7052
36. Jiang C, Yao X, Zhao Y, Wu J, Huang P, Pan C. et al. Comparative review of respiratory diseases caused by coronaviruses and influenza A viruses during epidemic season. Microbes Infect. 2020 S1286-4579(20) 30083-6
37. Cifu A, Levinson W. Influenza. JAMA. 2000;284:2847-9
38. Corman VM, Landt O, Kaiser M, Molenkamp R, Meijer A, Chu DK. et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Euro Surveill. 2020;25(3):2000045
39. Reusken C, Broberg EK, Haagmans B, Meijer A, Corman VM, Papa A. et al. Laboratory readiness and response for novel coronavirus (2019-nCoV) in expert laboratories in 30 EU/EEA countries, January 2020. Euro Surveill. 2020;25(6):2000082
40. Chu DKW, Pan Y, Cheng SMS, Hui KPY, Krishnan P, Liu Y. et al. Molecular Diagnosis of a Novel Coronavirus (2019-nCoV) Causing an Outbreak of Pneumonia. Clin Chem. 2020;66:549-55
41. Chan JF, Yip CC, To KK, Tang TH, Wong SC, Leung KH. et al. Improved Molecular Diagnosis of COVID-19 by the Novel, Highly Sensitive and Specific COVID-19-RdRp/Hel Real-Time Reverse Transcription-PCR Assay Validated In vitro and with Clinical Specimens. J Clin Microbiol. 2020;58(5):e00310-20
42. Lan L, Xu D, Ye G, Xia C, Wang S, Li Y. et al. Positive RT-PCR Test Results in Patients Recovered From COVID-19. JAMA. 2020;323(15):1502-1503
43. Young BE, Ong SWX, Kalimuddin S, Low JG, Tan SY, Loh J. et al. Epidemiologic Features and Clinical Course of Patients Infected With SARS-CoV-2 in Singapore. JAMA. 2020;323(15):1488-1494
44. Sheridan C. Coronavirus and the race to distribute reliable diagnostics. Nat Biotechnol. 2020;38:382-4
45. Li D, Zhang J, Li J. Primer design for quantitative real-time PCR for the emerging Coronavirus SARS-CoV-2. Theranostics. 2020;10(16):7150-7162
46. Nayional Medicial Products Administration. NMPA Issued the Report for Medical Device Registration. http://www.nmpa.gov.cn
47. Ai T, Yang Z, Hou H, Zhan C, Chen C, Lv W. et al. Correlation of Chest CT and RT-PCR Testing in Coronavirus Disease 2019 (COVID-19) in China: A Report of 1014 Cases. Radiology. 2020: 200642.
48. Zou L, Ruan F, Huang M, Liang L, Huang H, Hong Z. et al. SARS-CoV-2 Viral Load in Upper Respiratory Specimens of Infected Patients. N Engl J Med. 2020;382:1177-9
49. Chan JF, Yuan S, Kok KH, To KK, Chu H, Yang J. et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet. 2020;395:514-23
50. Gootenberg JS, Abudayyeh OO, Kellner MJ, Joung J, Collins JJ, Zhang F. Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science. 2018;360:439-44
51. Broughton JP, Deng X, Yu G, Fasching CL, Servellita V, Singh J. et al. CRISPR-Cas12-based detection of SARS-CoV-2. Nat Biotechnol. 2020 10.1038/s41587-020-0513-4
52. Zhao W, Zhong Z, Xie X, Yu Q, Liu J. CT scans of patients with 2019 novel Coronavirus (COVID-19) pneumonia. Theranostics. 2020;10:4606-13
53. Liu F, Zhang Q, Huang C, Shi C, Wang L, Shi N. et al. CT quantification of pneumonia lesions in early days predicts progression to severe illness in a cohort of COVID-19 patients. Theranostics. 2020;10(12):5613-5622
54. Yu Q, Wang Y, Huang S, Liu S, Zhou Z, Zhang S. et al. Multicenter cohort study demonstrates more consolidation in upper lungs on initial CT increases the risk of adverse clinical outcome in COVID-19 patients. Theranostics. 2020;10:5641-8
55. Li L, Yang L, Gui S, Pan F, Ye T, Liang B. et al. Association of clinical and radiographic findings with the outcomes of 93 patients with COVID-19 in Wuhan, China. Theranostics. 2020;10:6372-83
56. Xu P, Tian R, Luo S, Zu Z, Fan B, Wang X. et al. Risk factors for adverse clinical outcomes with COVID-19 in China: a multicenter, retrospective, observational study. Theranostics. 2020;10:6113-21
57. Yang W, Yan F. Patients with RT-PCR-confirmed COVID-19 and Normal Chest CT. Radiology. 2020;295:E3
58. Bernheim A, Mei X, Huang M, Yang Y, Fayad ZA, Zhang N. et al. Chest CT Findings in Coronavirus Disease-19 (COVID-19): Relationship to Duration of Infection. Radiology. 2020;295(3):200463
59. Pan F, Ye T, Sun P, Gui S, Liang B, Li L. et al. Time Course of Lung Changes On Chest CT During Recovery From 2019 Novel Coronavirus (COVID-19) Pneumonia. Radiology. 2020;295(3):715-721
60. Wang D, Hu B, Hu C, Zhu F, Liu X, Zhang J. et al. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus-Infected Pneumonia in Wuhan, China. JAMA. 2020;323(11):1061-1069
61. Sheridan C. Fast, portable tests come online to curb coronavirus pandemic. Nat Biotechnol. 2020;38(5):515-518
62. Zhang W, Du RH, Li B, Zheng XS, Yang XL, Hu B. et al. Molecular and serological investigation of 2019-nCoV infected patients: implication of multiple shedding routes. Emerg Microbes Infect. 2020;9:386-9
63. Zhao J, Yuan Q, Wang H, Liu W, Liao X, Su Y. et al. Antibody responses to SARS-CoV-2 in patients of novel coronavirus disease 2019. Clin Infect Dis. 2020 ciaa344
64. Li Z, Yi Y, Luo X, Xiong N, Liu Y, Li S. et al. Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. J Med Virol. 2020 10.1002/jmv.25727
65. Petherick A. Developing antibody tests for SARS-CoV-2. Lancet. 2020;395:1101-2
66. WHO. Population-based age-stratified seroepidemiological investigation protocol for COVID-19 virus infection. http://www.who.int/
67. To KK, Tsang OT, Leung WS, Tam AR, Wu TC, Lung DC. et al. Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. Lancet Infect Dis. 2020;20:565-74
68. Sheridan C. Fast, portable tests come online to curb coronavirus pandemic. Nat Biotechnol. 2020;38:515-8
69. Venter M, Richter K. Towards effective diagnostic assays for COVID-19: a review. J Clin Pathol. 2020 jclinpath-2020-206685
70. Udugama B, Kadhiresan P, Kozlowski HN, Malekjahani A, Osborne M, Li VYC. et al. Diagnosing COVID-19: The Disease and Tools for Detection. ACS Nano. 2020;14:3822-35
71. Huang Y, Cheng W, Zhao N, Qu H, Tian J. CT screening for early diagnosis of SARS-CoV-2 infection. Lancet Infect Dis. 2020 S1473-3099(20) 30241-3
72. Cellina M, Orsi M, Toluian T, Valenti Pittino C, Oliva G. False negative chest X-Rays in patients affected by COVID-19 pneumonia and corresponding chest CT findings. Radiography (Lond). 2020 S1078-8174(20) 30069-9
73. Li X, Geng M, Peng Y, Meng L, Lu S. Molecular immune pathogenesis and diagnosis of COVID-19. J Pharm Anal. 2020;10:102-8
74. Sakai J, Tarumoto N, Orihara Y, Kawamura R, Kodana M, Matsuzaki N. et al. Evaluation of a high-speed but low-throughput RT-qPCR system for SARS-CoV-2 detection. J Hosp Infect. 2020 S0195-6701(20) 30256-5
75. Wang Y, Kang H, Liu X, Tong Z. Combination of RT-qPCR testing and clinical features for diagnosis of COVID-19 facilitates management of SARS-CoV-2 outbreak. J Med Virol. 2020 10.1002/jmv.25721
76. Adli M. The CRISPR tool kit for genome editing and beyond. Nat Commun. 2018;9:1911
77. Guo L, Sun X, Wang X, Liang C, Jiang H, Gao Q. et al. SARS-CoV-2 detection with CRISPR diagnostics. Cell Discov. 2020;6:34
78. Abbott TR, Dhamdhere G, Liu Y, Lin X, Goudy L, Zeng L. et al. Development of CRISPR as an Antiviral Strategy to Combat SARS-CoV-2 and Influenza. Cell. 2020;181:865-76 e12
79. Li Z, Yi Y, Luo X, Xiong N, Liu Y, Li S. et al. Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. J Med Virol. 2020 10.1002/jmv.25727
80. Long QX, Liu BZ, Deng HJ, Wu GC, Deng K, Chen YK. et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat Med. 2020 10.1038/s41591-020-0897-1
81. Uhteg K, Jarrett J, Richards M, Howard C, Morehead E, Geahr M. et al. Comparing the analytical performance of three SARS-CoV-2 molecular diagnostic assays. J Clin Virol. 2020;127:104384
82. Xiang F, Wang X, He X, Peng Z, Yang B, Zhang J. et al. Antibody Detection and Dynamic Characteristics in Patients with COVID-19. Clin Infect Dis. 2020 ciaa461
83. Bruning AHL, Aatola H, Toivola H, Ikonen N, Savolainen-Kopra C, Blomqvist S. et al. Rapid detection and monitoring of human coronavirus infections. New Microbes New Infect. 2018;24:52-5
84. Liu IJ, Chen PJ, Yeh SH, Chiang YP, Huang LM, Chang MF. et al. Immunofluorescence assay for detection of the nucleocapsid antigen of the severe acute respiratory syndrome (SARS)-associated coronavirus in cells derived from throat wash samples of patients with SARS. J Clin Microbiol. 2005;43:2444-8
85. Chen Y, Chan KH, Hong C, Kang Y, Ge S, Chen H. et al. A highly specific rapid antigen detection assay for on-site diagnosis of MERS. J Infect. 2016;73:82-4
86. Diao B, Wen K, Chen J, Liu Y, Yuan Z, Han C. et al. Diagnosis of Acute Respiratory Syndrome Coronavirus 2 Infection by Detection of Nucleocapsid Protein. medRxiv. 2020 doi: https://doi.org/10.1101/2020.03.07.20032524
87. Zumla A, Chan JF, Azhar EI, Hui DS, Yuen KY. Coronaviruses - drug discovery and therapeutic options. Nat Rev Drug Discov. 2016;15:327-47
88. Wang M, Cao R, Zhang L, Yang X, Liu J, Xu M. et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020;30:269-71
89. Holshue ML, DeBolt C, Lindquist S, Lofy KH, Wiesman J, Bruce H. et al. First Case of 2019 Novel Coronavirus in the United States. N Engl J Med. 2020;382:929-36
90. Grein J, Ohmagari N, Shin D, Diaz G, Asperges E, Castagna A. et al. Compassionate Use of Remdesivir for Patients with Severe Covid-19. N Engl J Med. 2020 NEJMoa2007016
91. Wang Y, Zhang D, Du G, Du R, Zhao J, Jin Y. et al. Remdesivir in adults with severe COVID-19: a randomised, double-blind, placebo-controlled, multicentre trial. Lancet. 2020;395:1569-78
92. Savarino A, Boelaert JR, Cassone A, Majori G, Cauda R. Effects of chloroquine on viral infections: an old drug against today's diseases? Lancet Infect Dis. 2003;3:722-7
93. Chen Z, Hu J, Zhang Z, Jiang S, Han S, Yan D. et al. Efficacy of hydroxychloroquine in patients with COVID-19: results of a randomized clinical trial. medRxiv. 2020 doi: https://doi.org/10.1101/2020.03.22.20040758
94. Gao J, Tian Z, Yang X. Breakthrough: Chloroquine phosphate has shown apparent efficacy in treatment of COVID-19 associated pneumonia in clinical studies. Biosci Trends. 2020;14:72-3
95. Huang M, Tang T, Pang P, Li M, Ma R, Lu J. et al. Treating COVID-19 with Chloroquine. J Mol Cell Biol. 2020;12:322-5
96. Gautret P, Lagier JC, Parola P, Hoang VT, Meddeb L, Mailhe M. et al. Hydroxychloroquine and azithromycin as a treatment of COVID-19: results of an open-label non-randomized clinical trial. Int J Antimicrob Agents. 2020: 105949.
97. Mahevas M, Tran VT, Roumier M, Chabrol A, Paule R, Guillaud C. et al. Clinical efficacy of hydroxychloroquine in patients with covid-19 pneumonia who require oxygen: observational comparative study using routine care data. BMJ. 2020;369:m1844
98. Tang W, Cao Z, Han M, Wang Z, Chen J, Sun W. et al. Hydroxychloroquine in patients with mainly mild to moderate coronavirus disease 2019: open label, randomised controlled trial. BMJ. 2020;369:m1849
99. Kanters S, Socias ME, Paton NI, Vitoria M, Doherty M, Ayers D. et al. Comparative efficacy and safety of second-line antiretroviral therapy for treatment of HIV/AIDS: a systematic review and network meta-analysis. Lancet HIV. 2017;4:e433-e41
100. Itani R, Tobaiqy M, Al Faraj A. Optimizing use of theranostic nanoparticles as a life-saving strategy for treating COVID-19 patients. Theranostics. 2020;10(13):5932-5942
101. Sheahan TP, Sims AC, Leist SR, Schafer A, Won J, Brown AJ. et al. Comparative therapeutic efficacy of remdesivir and combination lopinavir, ritonavir, and interferon beta against MERS-CoV. Nat Commun. 2020;11:222
102. Chu CM, Cheng VC, Hung IF, Wong MM, Chan KH, Chan KS. et al. Role of lopinavir/ritonavir in the treatment of SARS: initial virological and clinical findings. Thorax. 2004;59:252-6
103. Chen XP, Cao Y. Consideration of highly active antiretroviral therapy in the prevention and treatment of severe acute respiratory syndrome. Clin Infect Dis. 2004;38:1030-2
104. de Wilde AH, Jochmans D, Posthuma CC, Zevenhoven-Dobbe JC, van Nieuwkoop S, Bestebroer TM. et al. Screening of an FDA-approved compound library identifies four small-molecule inhibitors of Middle East respiratory syndrome coronavirus replication in cell culture. Antimicrob Agents Chemother. 2014;58:4875-84
105. Furuta Y, Komeno T, Nakamura T. Favipiravir (T-705), a broad spectrum inhibitor of viral RNA polymerase. Proc Jpn Acad Ser B Phys Biol Sci. 2017;93:449-63
106. Ramsey ML, Nuttall J, Hart PA, Team TI. A phase 1/2 trial to evaluate the pharmacokinetics, safety, and efficacy of NI-03 in patients with chronic pancreatitis: study protocol for a randomized controlled trial on the assessment of camostat treatment in chronic pancreatitis (TACTIC). Trials. 2019;20(1):501
107. Check Hayden E. Experimental drugs poised for use in Ebola outbreak. Nature. 2018;557:475-6
108. Tchesnokov EP, Feng JY, Porter DP, Gotte M. Mechanism of Inhibition of Ebola Virus RNA-Dependent RNA Polymerase by Remdesivir. Viruses. 2019;11(4):326
109. Vincent MJ, Bergeron E, Benjannet S, Erickson BR, Rollin PE, Ksiazek TG. et al. Chloroquine is a potent inhibitor of SARS coronavirus infection and spread. Virol J. 2005;2:69
110. Costanzo M, De Giglio MAR, Roviello GN. SARS-CoV-2: Recent Reports on Antiviral Therapies Based on Lopinavir/Ritonavir, Darunavir/Umifenovir, Hydroxychloroquine, Remdesivir, Favipiravir and Other Drugs for the Treatment of the New Coronavirus. Curr Med Chem. 2020 10.2174/0929867327666200416131117
111. Liu J, Cao R, Xu M, Wang X, Zhang H, Hu H. et al. Hydroxychloroquine, a less toxic derivative of chloroquine, is effective in inhibiting SARS-CoV-2 infection in vitro. Cell Discov. 2020;6:16
112. Sanders JM, Monogue ML, Jodlowski TZ, Cutrell JB. Pharmacologic Treatments for Coronavirus Disease 2019 (COVID-19): A Review. JAMA. 2020; 10.1001/jama. 2020 6019
113. Cai Q, Yang M, Liu D, Chen J, Shu D, Xia J. et al. Experimental Treatment with Favipiravir for COVID-19: An Open-Label Control Study. Engineering (Beijing). 2020; 10.1016/j.eng. 2020 03.007
114. Du YX, Chen XP. Favipiravir: Pharmacokinetics and Concerns About Clinical Trials for 2019-nCoV Infection. Clin Pharmacol Ther. 2020 10.1002/cpt.1844
115. Fang QQ, Huang WJ, Li XY, Cheng YH, Tan MJ, Liu J. et al. Effectiveness of favipiravir (T-705) against wild-type and oseltamivir-resistant influenza B virus in mice. Virology. 2020;545:1-9
116. Fletcher TE, Fischer WA 2nd, Jacob ST. Convalescent Plasma for Ebola Virus Disease. N Engl J Med. 2016;374:2499-500
117. Zhou B, Zhong N, Guan Y. Treatment with convalescent plasma for influenza A (H5N1) infection. N Engl J Med. 2007;357:1450-1
118. Stockman LJ, Bellamy R, Garner P. SARS: systematic review of treatment effects. PLoS Med. 2006;3:e343
119. Arabi YM, Hajeer AH, Luke T, Raviprakash K, Balkhy H, Johani S. et al. Feasibility of Using Convalescent Plasma Immunotherapy for MERS-CoV Infection, Saudi Arabia. Emerg Infect Dis. 2016;22:1554-61
120. Hung IF, To KK, Lee CK, Lee KL, Chan K, Yan WW. et al. Convalescent plasma treatment reduced mortality in patients with severe pandemic influenza A (H1N1) 2009 virus infection. Clin Infect Dis. 2011;52:447-56
121. Roback JD, Guarner J. Convalescent Plasma to Treat COVID-19: Possibilities and Challenges. JAMA. 2020 doi: 10.1001/jama.2020.4940
122. Zhang B, Liu S, Tan T, Huang W, Dong Y, Chen L. et al. Treatment with Convalescent Plasma for Critically Ill Patients With SARS-CoV-2 Infection. Chest. 2020 S0012-3692(20)30571-7
123. Shen C, Wang Z, Zhao F, Yang Y, Li J, Yuan J. et al. Treatment of 5 Critically Ill Patients With COVID-19 With Convalescent Plasma. JAMA. 2020;323(16):1582-1589
124. Zhou G, Zhao Q. Perspectives on therapeutic neutralizing antibodies against the Novel Coronavirus SARS-CoV-2. Int J Biol Sci. 2020;16:1718-23
125. Sui J, Li W, Murakami A, Tamin A, Matthews LJ, Wong SK. et al. Potent neutralization of severe acute respiratory syndrome (SARS) coronavirus by a human mAb to S1 protein that blocks receptor association. Proc Natl Acad Sci U S A. 2004;101:2536-41
126. Zhu Z, Chakraborti S, He Y, Roberts A, Sheahan T, Xiao X. et al. Potent cross-reactive neutralization of SARS coronavirus isolates by human monoclonal antibodies. Proc Natl Acad Sci U S A. 2007;104:12123-8
127. Wu Y, Li C, Xia S, Tian X, Kong Y, Wang Z. et al. Identification of Human Single-Domain Antibodies against SARS-CoV-2. Cell Host Microbe. 2020 S1931-3128(20) 30250-X
128. Shi R, Shan C, Duan X, Chen Z, Liu P, Song J. et al. A human neutralizing antibody targets the receptor binding site of SARS-CoV-2. Nature. 2020 10.1038/s41586-020-2381-y
129. Jiang S, Hillyer C, Du L. Neutralizing Antibodies against SARS-CoV-2 and Other Human Coronaviruses. Trends Immunol. 2020;41(5):355-359
130. Ju B, Zhang Q, Ge X, Wang R, Yu J, Shan S. et al. Potent human neutralizing antibodies elicited by SARS-CoV-2 infection. Nature. 2020 10.1038/s41586-020-2380-z
131. Wu Y, Wang F, Shen C, Peng W, Li D, Zhao C. et al. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science. 2020 eabc2241
132. Chen Z, Mi L, Xu J, Wang X, Jiang J, Xing J. et al. Function of HAb18G/CD147 in invasion of host cells by severe acute respiratory syndrome coronavirus. J Infect Dis. 2005;191(5):755-760
133. Wang K, Chen W, Zhou Y, Lian J, Zhang Z, Du P. et al. SARS-CoV-2 invades host cells via a novel route: CD147-spike protein. BioRxiv preprint. 2020 doi: 10.1101/2020.03.14.988345
134. Wan Y, Shang J, Graham R, Baric RS, Li F. Receptor Recognition by the Novel Coronavirus from Wuhan: an Analysis Based on Decade-Long Structural Studies of SARS Coronavirus. J Virol. 2020;94(7):e00127-20
135. Zhang H, Penninger JM, Li Y, Zhong N, Slutsky AS. Angiotensin-converting enzyme 2 (ACE2) as a SARS-CoV-2 receptor: molecular mechanisms and potential therapeutic target. Intensive Care Med. 2020;46:586-90
136. Gheblawi M, Wang K, Viveiros A, Nguyen Q, Zhong JC, Turner AJ. et al. Angiotensin Converting Enzyme 2: SARS-CoV-2 Receptor and Regulator of the Renin-Angiotensin System. Circ Res. 2020;126(10):1456-1474
137. Yuan M, Wu NC, Zhu X, Lee CD, So RTY, Lv H. et al. A highly conserved cryptic epitope in the receptor-binding domains of SARS-CoV-2 and SARS-CoV. Science. 2020;368(6491):630-633
138. Tian X, Li C, Huang A, Xia S, Lu S, Shi Z. et al. Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerg Microbes Infect. 2020;9:382-5
139. Wang C, Li W, Drabek D, Okba NMA, van Haperen R, Osterhaus ADME. et al. A human monoclonal antibody blocking SARS-CoV-2 infection. Nature Communications. 2020;11:2251
140. Iwasaki A, Yang Y. The potential danger of suboptimal antibody responses in COVID-19. Nat Rev Immunol. 2020;20:339-41
141. Tang B, Xiao Y, Sander B, Kulkarni MA, Radam-Lac Research T, Wu J. Modelling the impact of antibody-dependent enhancement on disease severity of Zika virus and dengue virus sequential and co-infection. R Soc Open Sci. 2020;7:191749
142. Tirado SM, Yoon KJ. Antibody-dependent enhancement of virus infection and disease. Viral Immunol. 2003;16:69-86
143. Ngono AE, Shresta S. Immune Response to Dengue and Zika. Annu Rev Immunol. 2018;36:279-308
144. Dolgin E. Unlocking the potential of vaccines built on messenger RNA. Nature. 2019;574:S10-S2
145. Pardi N, Hogan MJ, Porter FW, Weissman D. mRNA vaccines - a new era in vaccinology. Nat Rev Drug Discov. 2018;17:261-79
146. Richner JM, Himansu S, Dowd KA, Butler SL, Salazar V, Fox JM. et al. Modified mRNA Vaccines Protect against Zika Virus Infection. Cell. 2017;169:176
147. Alberer M, Gnad-Vogt U, Hong HS, Mehr KT, Backert L, Finak G. et al. Safety and immunogenicity of a mRNA rabies vaccine in healthy adults: an open-label, non-randomised, prospective, first-in-human phase 1 clinical trial. Lancet. 2017;390:1511-20
148. Pardi N, Parkhouse K, Kirkpatrick E, McMahon M, Zost SJ, Mui BL. et al. Nucleoside-modified mRNA immunization elicits influenza virus hemagglutinin stalk-specific antibodies. Nat Commun. 2018;9:3361
149. Moderna. Moderna Announces Positive Interim Phase 1 Data for its mRNA Vaccine (mRNA-1273) Against Novel Coronavirus. Revised 18 May 2020. https://www.modernatx.com
150. Lee J, Arun Kumar S, Jhan YY, Bishop CJ. Engineering DNA vaccines against infectious diseases. Acta Biomater. 2018;80:31-47
151. Modjarrad K, Roberts CC, Mills KT, Castellano AR, Paolino K, Muthumani K. et al. Safety and immunogenicity of an anti-Middle East respiratory syndrome coronavirus DNA vaccine: a phase 1, open-label, single-arm, dose-escalation trial. Lancet Infect Dis. 2019;19:1013-22
152. Smith TRF, Patel A, Ramos S, Elwood D, Zhu X, Yan J. et al. Immunogenicity of a DNA vaccine candidate for COVID-19. Nat Commun. 2020;11:2601
153. Zhu FC, Li YH, Guan XH, Hou LH, Wang WJ, Li JX. et al. Safety, tolerability, and immunogenicity of a recombinant adenovirus type-5 vectored COVID-19 vaccine: a dose-escalation, open-label, non-randomised, first-in-human trial. Lancet. 2020 S0140-6736(20)31208-3
154. van Doremalen N, Lambe T, Spencer A, Belij-Rammerstorfer S, Purushotham J, Port J. et al. ChAdOx1 nCoV-19 vaccination prevents SARS-CoV-2 pneumonia in rhesus macaques. bioRxiv. 2020 doi: https://doi.org/10.1101/2020.05.13.093195
155. Vellozzi C, Burwen DR, Dobardzic A, Ball R, Walton K, Haber P. Safety of trivalent inactivated influenza vaccines in adults: background for pandemic influenza vaccine safety monitoring. Vaccine. 2009;27:2114-20
156. Murdin AD, Barreto L, Plotkin S. Inactivated poliovirus vaccine: past and present experience. Vaccine. 1996;14:735-46
157. ClinicalTrials. Safety and Immunogenicity Study of Inactivated Vaccine for Prophylaxis of SARS CoV-2 Infection (COVID-19). Revised 20 April 2020. https://clinicaltrials.gov/
158. NOVAVAX. Novavax Initiates Phase 1/2 Clinical Trial of COVID-19 Vaccine. Revised 25 May 2020. https://novavax.com
159. ClinicalTrials.gov. Immunity and Safety of Covid-19 Synthetic Minigene Vaccine. Revised 19 Febryary 2020. https://clinicaltrials.gov
160. ClinicalTrials.gov. Safety and Immunity of Covid-19 aAPC Vaccine. Revised 9 March 2020. https://clinicaltrials.gov
161. Wang M, Li M, Ren R, Li L, Chen EQ, Li W. et al. International expansion of a novel SARS-CoV-2 mutant. J Virol. 2020;94(12):e00567-20
162. Lau SY, Wang P, Mok BW, Zhang AJ, Chu H, Lee AC. et al. Attenuated SARS-CoV-2 variants with deletions at the S1/S2 junction. Emerg Microbes Infect. 2020;9:837-42
163. Xia S, Liu M, Wang C, Xu W, Lan Q, Feng S. et al. Inhibition of SARS-CoV-2 (previously 2019-nCoV) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion. Cell Res. 2020;30:343-55
164. Xia S, Yan L, Xu W, Agrawal AS, Algaissi A, Tseng CK. et al. A pan-coronavirus fusion inhibitor targeting the HR1 domain of human coronavirus spike. Sci Adv. 2019;5:eaav4580
165. Xia S, Zhu Y, Liu M, Lan Q, Xu W, Wu Y. et al. Fusion mechanism of 2019-nCoV and fusion inhibitors targeting HR1 domain in spike protein. Cell Mol Immunol. 2020:1-3
166. Bao L, Deng W, Huang B, Gao H, Liu J, Ren L. et al. The pathogenicity of SARS-CoV-2 in hACE2 transgenic mice. Nature. 2020 10.1038/s41586-020-2312-y
167. Jiang R, Liu M, Chen Y, Shan C, Zhou Y, Shen X. et al. Pathogenesis of SARS-CoV-2 in transgenic mice expressing human angiotensin-converting enzyme 2. Cell. in press. doi: https://doi.org/10.1016/j.cell. 2020 05.027
168. Hung IF, Lung KC, Tso EY, Liu R, Chung TW, Chu MY. et al. Triple combination of interferon beta-1b, lopinavir-ritonavir, and ribavirin in the treatment of patients admitted to hospital with COVID-19: an open-label, randomised, phase 2 trial. Lancet. 2020;395(10238):1695-1704
169. Cheng VC, Tang BS, Wu AK, Chu CM, Yuen KY. Medical treatment of viral pneumonia including SARS in immunocompetent adult. J Infect. 2004;49:262-73
170. Melville K, Rodriguez T, Dobrovolny HM. Investigating Different Mechanisms of Action in Combination Therapy for Influenza. Front Pharmacol. 2018;9:1207
171. Dunning J, Baillie JK, Cao B, Hayden FG, International Severe Acute R, Emerging Infection C. Antiviral combinations for severe influenza. Lancet Infect Dis. 2014;14:1259-70
172. Hung IFN, To KKW, Chan JFW, Cheng VCC, Liu KSH, Tam A. et al. Efficacy of Clarithromycin-Naproxen-Oseltamivir Combination in the Treatment of Patients Hospitalized for Influenza A(H3N2) Infection: An Open-label Randomized, Controlled, Phase IIb/III Trial. Chest. 2017;151:1069-80
Author contact
Corresponding author: Wenxin Luo (E-mail: wxluoedu.cn).
 Global reach, higher impact
Global reach, higher impact