13.3
Impact Factor
Theranostics 2020; 10(20):9268-9279. doi:10.7150/thno.46341 This issue Cite
Research Paper
Instability of circular RNAs in clinical tissue samples impairs their reliable expression analysis using RT-qPCR: from the myth of their advantage as biomarkers to reality
1. Department of Urology, Charité - University Medicine, 10117 Berlin, Germany.
2. Berlin Institute for Urologic Research, 10115 Berlin, Germany.
3. Department of Pathology, Charité - University Medicine, 10117 Berlin, Germany.
4. Institute of Pathology, Hospital Leverkusen, 51375 Leverkusen, Germany.
5. Max Delbrueck Center for Molecular Medicine in the Helmholtz Association, Cancer Research Program, 13125 Berlin, Germany.
6. Cancer Dynamics Laboratory, The Francis Crick Institute, 1 Midland Road, London NW1 1AT, U.K.
*These authors contributed equally to this work.
#These authors share senior authorship.
Abstract
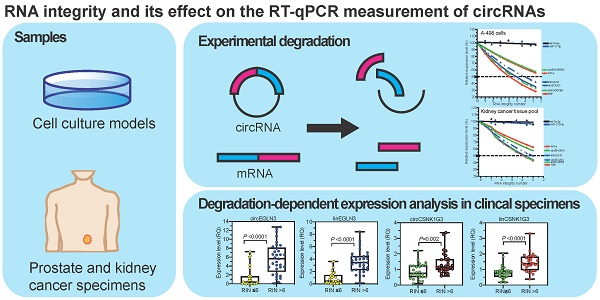
Background: Circular RNAs (circRNAs) are a new class of RNAs with medical significance. Compared to that of linear mRNA transcripts, the stability of circRNAs against degradation owing to their circular structure is considered advantageous for their use as biomarkers. As systematic studies on the stability of circRNAs depending on the RNA integrity, determined as RNA integrity number (RIN), in clinical tissue samples are lacking, we have investigated this aspect in the present study under model and clinical conditions.
Methods: Total RNA isolated from kidney cancer tissue and cell lines (A-498 and HEK-293) with different RIN after thermal degradation was used in model experiments. Further, RNA isolated from kidney cancer and prostate cancer tissue collected under routine surgical conditions, representing clinical samples with RIN ranging from 2 to 9, were examined. Quantitative real-time reverse-transcription polymerase chain reaction (RT-qPCR) analysis of several circRNAs (circEGLN3, circRHOBTB3, circCSNK1G3, circRNA4, and circRNA9), their corresponding linear counterparts, tissue-specific reference genes, and three microRNAs (as controls) was performed. The quantification cycles were converted into relative quantities and normalized to the expression of specific reference genes for the corresponding tissue. The effect of RIN on the expression of different RNA entities was determined using linear regression analysis, and clinical samples were classified into two groups based on RIN greater or lesser than 6.
Results: The results of model experiments and clinical sample analyses showed that all relative circRNA expression gradually decreased with reduction in RIN values. The adverse effect of RIN was partially compensated after normalizing the data and limiting the samples to only those with RIN values > 6.
Conclusions: Our results suggested that circRNAs are not stable in clinical tissue samples, but are subjected to degradative processes similar to mRNAs. This has not been investigated extensively in circRNA expression studies, and hence must be considered in future for obtaining reliable circRNA expression data. This can be achieved by applying the principles commonly used in mRNA expression studies.
Keywords: circular RNAs, RNA integrity, RNA degradation, circRNA stability, normalization, RT-qPCR
 Global reach, higher impact
Global reach, higher impact