13.3
Impact Factor
Theranostics 2020; 10(20):9303-9314. doi:10.7150/thno.46568 This issue Cite
Research Paper
Nanozyme-assisted sensitive profiling of exosomal proteins for rapid cancer diagnosis
1. Research Center for Analytical Sciences in the College of Chemistry, Tianjin First Central Hospital, State Key Laboratory of Medicinal Chemical Biology, and Tianjin Key Laboratory of Molecular Recognition and Biosensing, Nankai University, Tianjin 300071, China.
2. Department of Radiology, The Third Xiangya Hospital, Central South University, Changsha, Hunan 410013, China.
Abstract
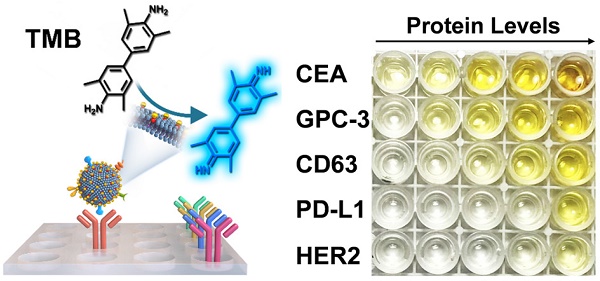
The proteins expressed on exosomes have emerged as promising liquid-biopsy biomarkers for cancer diagnosis. However, molecular profiling of exosomal proteins remains technically challenging. Herein, we report a nanozyme-assisted immunosorbent assay (NAISA) that enables sensitive and rapid multiplex profiling of exosomal proteins. This NAISA system is based on the installation of peroxidase-like nanozymes onto the phospholipid membranes of exosomes, thus avoiding the need for post-labelling detection antibodies. The exosomal proteins are determined by a sensitive nanozyme-catalyzed colorimetric assay less than 3 h, without the need for multi-step incubation and washing operations. Using NAISA to profile exosomal proteins from different cell lines and clinical samples, we reveal that tumor-associated exosomal proteins can serve as promising biomarkers for accurate cancer diagnosis in a cooperative detection pattern.
Methods: Exosomes were engineered with DSPE-PEG-SH through hydrophobic interaction, and then were assembled with gold nanoparticles (2 nm) to produce Exo@Au nanozyme. The proteins on Exo@Au could be selectively captured by their specific antibodies seeded into a 96-well plate. The immobilized Exo@Au shows peroxidase-like activity to perform colorimetric assays by reaction with 3,3′,5,5′-tetramethylbenzidine (TMB) and H2O2. The protein levels of exosomes were recorded on a microplate reader.
Results: The NAISA platform is capable of profiling multiple exosomal proteins from both cancer cell lines and clinical samples. The expression levels of exosomal proteins, such as CD63, CEA, GPC-3, PD-L1 and HER2, were used to classify different cancer cell lines. Moreover, the protein profiles have been applied to differentiate healthy donors, hepatitis B patients, and hepatic cell carcinoma (HCC) patients with high accuracy.
Conclusion: The NAISA nanozyme was allowed to rapidly profile multiple exosomal proteins and could have great promise for early HCC diagnosis and identification of other cancer types.
Keywords: Exosome, nanozyme, protein profiling, cancer diagnosis, nanozyme-assisted immunosorbent assay
 Global reach, higher impact
Global reach, higher impact