13.3
Impact Factor
Theranostics 2021; 11(1):361-378. doi:10.7150/thno.46360 This issue Cite
Research Paper
Histone H3K27 methyltransferase EZH2 and demethylase JMJD3 regulate hepatic stellate cells activation and liver fibrosis
1. Shanghai Fifth People's Hospital of Fudan University, Shanghai Key Laboratory of Medical Epigenetics, Ministry of Science and Technology International Co-Laboratory of Medical Epigenetics and Metabolism, Institutes of Biomedical Sciences, Shanghai Medical College, Fudan University, Shanghai, China.
2. Center for Medical Research and Innovation, Shanghai Pudong Hospital and Pudong Medical Center, Shanghai Medical College, Fudan University, Shanghai, China.
3. State Key Laboratory of Genetic Engineering and Collaborative Innovation Center for Genetics and Development, Ministry of Education Key Laboratory of Contemporary Anthropology and Department of Anthropology and Human Genetics, School of Life Sciences, Fudan University, Shanghai, China.
4. State Key Laboratory of Proteomics, Beijing Proteome Research Center, National Center for Protein Sciences, Beijing Institute of Lifeomics, Beijing, China.
Abstract
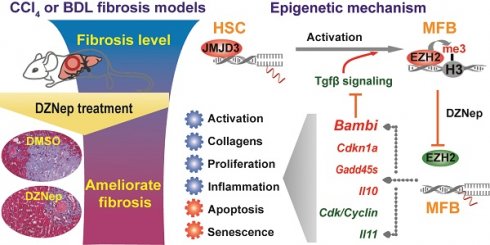
Rationale: As the central hallmark of liver fibrosis, transdifferentiation of hepatic stellate cells (HSCs), the predominant contributor to fibrogenic hepatic myofibroblast responsible for extracellular matrix (ECM) deposition, is characterized with transcriptional and epigenetic remodeling. We aimed to characterize the roles of H3K27 methyltransferase EZH2 and demethylase JMJD3 and identify their effective pathways and novel target genes in HSCs activation and liver fibrosis.
Methods: In primary HSCs, we analyzed effects of pharmacological inhibitions and genetic manipulations of EZH2 and JMJD3 on HSCs activation. In HSCs cell lines, we evaluated effects of EZH2 inhibition by DZNep on proliferation, cell cycling, senescence and apoptosis. In CCl4 and BDL murine models of liver fibrosis, we assessed in vivo effects of DZNep administration and Ezh2 silencing. We profiled rat primary HSCs transcriptomes with RNA-seq, screened the pathways and genes associated with DZNep treatment, analyzed EZH2 and JMJD3 regulation towards target genes by ChIP-qPCR.
Results: EZH2 inhibition by DZNep resulted in retarded growth, lowered cell viability, cell cycle arrest in S and G2 phases, strengthened senescence, and enhanced apoptosis of HSCs, decreased hepatic collagen deposition and rescued the elevated serum ALT and AST activities of diseased mice, and downregulated cellular and hepatic expressions of H3K27me3, EZH2, α-SMA and COL1A. Ezh2 silencing by RNA interference in vitro and in vivo showed similar effects. JMJD3 inhibition by GSK-J4 and overexpression of wild-type but not mutant Jmjd3 enhanced or repressed HSCs activation respectively. EZH2 inhibition by DZNep transcriptionally inactivated TGF-β1 pathway, cell cycle pathways and vast ECM components in primary HSCs. EZH2 inhibition decreased H3K27me3 recruitment at target genes encoding TGF-β1 pseudoreceptor BAMBI, anti-inflammatory cytokine IL10 and cell cycle regulators CDKN1A, GADD45A and GADD45B, and increased their expressions, while Jmjd3 overexpression manifested alike effects.
Conclusions: EZH2 and JMJD3 antagonistically modulate HSCs activation. The therapeutic effects of DZNep as epigenetic drug in liver fibrosis are associated with the regulation of EZH2 towards direct target genes encoding TGF-β1 pseudoreceptor BAMBI, anti-inflammatory cytokine IL10 and cell cycle regulators CDKN1A, GADD45A and GADD45B, which are also regulated by JMJD3. Our present study provides new mechanistic insight into the epigenetic modulation of EZH2 and JMJD3 in HSCs biology and hepatic fibrogenesis.
Keywords: liver fibrosis, HSC, EZH2, JMJD3, epigenetics
 Global reach, higher impact
Global reach, higher impact