13.3
Impact Factor
Theranostics 2021; 11(3):1458-1472. doi:10.7150/thno.48744 This issue Cite
Research Paper
Proteasome activator PA200 maintains stability of histone marks during transcription and aging
1. State Key Laboratory of Cognitive Neuroscience & Learning and Ministry of Education Key Laboratory of Cell Proliferation & Regulation Biology, College of Life Sciences, Beijing Normal University, 19 Xinjiekouwai Avenue, Beijing 100875, China.
2. College of Life Sciences, Beijing Normal University, 19 Xinjiekouwai Avenue, Beijing 100875, China.
3. Graduate School of Life and Environmental Sciences, University of Tsukuba, 1-1-1 Tennodai, Tsukuba, Ibaraki 305-8577, Japan.
4. School of Life Sciences, Tsinghua University, Beijing 100084, China.
*These authors contributed equally to this work.
Abstract
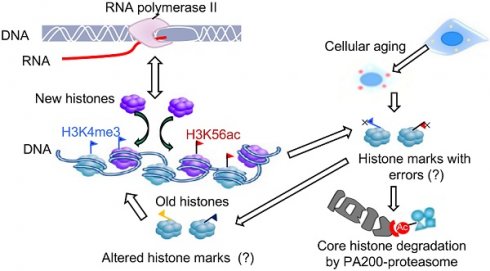
The epigenetic inheritance relies on stability of histone marks, but various diseases, including aging-related disorders, are usually associated with alterations of histone marks. Whether and how the proteasome is responsible for maintaining the histone marks during transcription and aging remain unclear. The core histones can be degraded by the atypical proteasome, which contains the proteasome activator PA200, in an acetylation-dependent manner during somatic DNA damage response and spermiogenesis.
Methods: By utilizing a substitute of methionine to label proteins metabolically, we analyzed histone degradation genome-wide by sequencing the DNA fragments following pulse-chase assays. The genome-wide RNA-sequencing analysis was performed to analyze transcription and chromatin-immunoprecipitation (ChIP)-sequencing was used for analyses of histone marks. The experimental models included gene-manipulated cells (including both mouse and yeast), mouse liver, and mice.
Results: Degradation of H4 or the transcription-coupled histone variant H3.3 could be suppressed by deletion of PA200 or its yeast ortholog Blm10. The histone deacetylase inhibitor accelerated the degradation rates of H3, while the mutations of the putative acetyl-lysine-binding region of PA200 abolished histone degradation in the G1-arrested cells. Deletion of PA200 dramatically altered deposition of the active transcriptional hallmarks (H3K4me3 and H3K56ac) and transcription, especially during cellular aging. Furthermore, deletion of PA200 or Blm10 accelerated cellular aging. Notably, the PA200-deficient mice displayed a range of aging-related deteriorations, including immune malfunction, anxiety-like behavior and shorter lifespan.
Conclusion: PA200 promotes the transcription-coupled degradation of the core histones, and plays an important role in maintaining the stability of histone marks during transcription and aging.
Keywords: Histone marks, aging, histone degradation, proteasome activator PA200, transcription
 Global reach, higher impact
Global reach, higher impact