13.3
Impact Factor
Theranostics 2021; 11(6):2691-2705. doi:10.7150/thno.50571 This issue Cite
Research Paper
KDM6A promotes imatinib resistance through YY1-mediated transcriptional upregulation of TRKA independently of its demethylase activity in chronic myelogenous leukemia
1. Department of Central Laboratory, The Affiliated Huaian No. 1 People's Hospital, Nanjing Medical University, Huai'an, Jiangsu, 223300, China.
2. Department of Hematology, The Affiliated Huaian No.1 People's Hospital,Nanjing Medical University, Huai'an, Jiangsu, 223300, China.
3. Institute of Medicinal Biotechnology, Jiangsu College of Nursing, Huai'an, Jiangsu, 223300, China.
4. The Chinese Academy of Sciences (CAS), Key Laboratory of Innate Immunity & Chronic Disease, CAS Center for Excellence in Cell & Molecular Biology, School of Life Sciences, University of Science & Technology of China, Hefei 230026, China.
5. Department of Oncology, Nanjing First Hospital, Nanjing Medical University, Nanjing, 210006, China.
6. Translational Research Institute of Henan Provincial People's Hospital and People's Hospital of Zhengzhou University, and Molecular Pathology Center, Academy of Medical Sciences, Zhengzhou University, Zhengzhou, Henan 450053, China.
7. School of Environmental and Life Sciences, The University of Newcastle, NSW, 2258, Australia.
8. School of Biomedical Sciences and Pharmacy, The University of Newcastle, NSW, 2308, Australia.
Abstract
Rationale: Despite landmark therapy of chronic myelogenous leukemia (CML) with tyrosine kinase inhibitors (TKIs), drug resistance remains problematic. Cancer pathogenesis involves epigenetic dysregulation and in particular, histone lysine demethylases (KDMs) have been implicated in TKI resistance. We sought to identify KDMs with altered expression in CML and define their contribution to imatinib resistance.
Methods: Bioinformatics screening compared KDM expression in CML versus normal bone marrow with shRNA knockdown and flow cytometry used to measure effects on imatinib-induced apoptosis in K562 cells. Transcriptomic analyses were performed against KDM6A CRISPR knockout/shRNA knockdown K562 cells along with gene rescue experiments using wildtype and mutant demethylase-dead KDM6A constructs. Co-immunoprecipitation, luciferase reporter and ChIP were employed to elucidate mechanisms of KDM6A-dependent resistance.
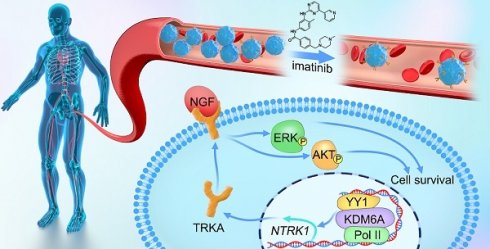
Results: Amongst five KDMs upregulated in CML, only KDM6A depletion sensitized CML cells to imatinib-induced apoptosis. Re-introduction of demethylase-dead KDM6A as well as wild-type KDM6A restored imatinib resistance. RNA-seq identified NTRK1 gene downregulation after depletion of KDM6A. Moreover, NTRK1 expression positively correlated with KDM6A in a subset of clinical CML samples and KDM6A knockdown in fresh CML isolates decreased NTRK1 encoded protein (TRKA) expression. Mechanistically, KDM6A was recruited to the NTRK1 promoter by the transcription factor YY1 with subsequent TRKA upregulation activating down-stream survival pathways to invoke imatinib resistance.
Conclusion: Contrary to its reported role as a tumor suppressor and independent of its demethylase function, KDM6A promotes imatinib-resistance in CML cells. The identification of the KDM6A/YY1/TRKA axis as a novel imatinib-resistance mechanism represents an unexplored avenue to overcome TKI resistance in CML.
Keywords: KDM6A/UTX, YY1, NTRK1/TRKA, CML, imatinib resistance
 Global reach, higher impact
Global reach, higher impact