13.3
Impact Factor
Theranostics 2021; 11(6):3017-3034. doi:10.7150/thno.51887 This issue Cite
Research Paper
Machine learning identifies stroke features between species
1. Werner Siemens Imaging Center, Department of Preclinical Imaging and Radiopharmacy, Eberhard Karls University Tuebingen, Tuebingen, Germany.
2. Department of Nuclear Medicine and Clinical Molecular Imaging, Eberhard Karls University Tuebingen, Tuebingen, Germany.
3. Max Planck Institute for Intelligent Systems, Tuebingen, Germany.
4. Department of Neurology & Stroke, and Hertie Institute for Clinical Brain Research, Eberhard Karls University Tuebingen, Tuebingen, Germany.
5. Department of Diagnostic and Interventional Neuroradiology, Eberhard Karls University Tuebingen, Tuebingen, Germany.
6. Institute of Pathology and Neuropathology and Comprehensive Cancer Center Tübingen, Eberhard Karls University Tuebingen, Tübingen, Germany.
†These authors contributed equally.
Abstract
Identification and localization of ischemic stroke (IS) lesions is routinely performed to confirm diagnosis, assess stroke severity, predict disability and plan rehabilitation strategies using magnetic resonance imaging (MRI). In basic research, stroke lesion segmentation is necessary to study complex peri-infarction tissue changes. Moreover, final stroke volume is a critical outcome evaluated in clinical and preclinical experiments to determine therapy or intervention success. Manual segmentations are performed but they require a specialized skill set, are prone to inter-observer variation, are not entirely objective and are often not supported by histology. The task is even more challenging when dealing with large multi-center datasets, multiple experimenters or large animal cohorts. On the other hand, current automatized segmentation approaches often lack histological validation, are not entirely user independent, are often based on single parameters, or in the case of complex machine learning methods, require vast training datasets and are prone to a lack of model interpretation.
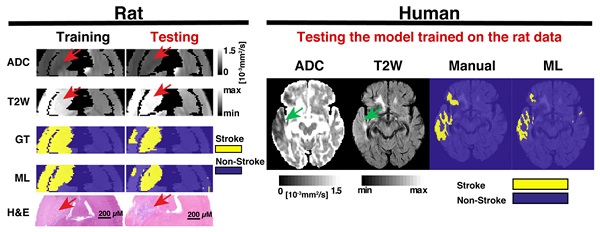
Methods: We induced IS using the middle cerebral artery occlusion model on two rat cohorts. We acquired apparent diffusion coefficient (ADC) and T2-weighted (T2W) images at 24 h and 1-week after IS induction. Subsets of the animals at 24 h and 1-week post IS were evaluated using histology and immunohistochemistry. Using a Gaussian mixture model, we segmented voxel-wise interactions between ADC and T2W parameters at 24 h using one of the rat cohorts. We then used these segmentation results to train a random forest classifier, which we applied to the second rat cohort. The algorithms' stroke segmentations were compared to manual stroke delineations, T2W and ADC thresholding methods and the final stroke segmentation at 1-week. Volume correlations to histology were also performed for every segmentation method. Metrics of success were calculated with respect to the final stroke volume. Finally, the trained random forest classifier was tested on a human dataset with a similar temporal stroke on-set. Manual segmentations, ADC and T2W thresholds were again used to evaluate and perform comparisons with the proposed algorithms' output.
Results: In preclinical rat data our framework significantly outperformed commonly applied automatized thresholding approaches and segmented stroke regions similarly to manual delineation. The framework predicted the localization of final stroke regions in 1-week post-stroke MRI with a median Dice similarity coefficient of 0.86, Matthew's correlation coefficient of 0.80 and false positive rate of 0.04. The predicted stroke volumes also strongly correlated with final histological stroke regions (Pearson correlation = 0.88, P < 0.0001). Lastly, the stroke region characteristics identified by our framework in rats also identified stroke lesions in human brains, largely outperforming thresholding approaches in stroke volume prediction (P<0.01).
Conclusion: Our findings reveal that the segmentation produced by our proposed framework using 24 h MRI rat data strongly correlated with the final stroke volume, denoting a predictive effect. In addition, we show for the first time that the stroke imaging features can be directly translated between species, allowing identification of acute stroke in humans using the model trained on animal data. This discovery reduces the gap between the clinical and preclinical fields, unveiling a novel approach to directly co-analyze clinical and preclinical data. Such methods can provide further biological insights into human stroke and highlight the differences between species in order to help improve the experimental setups and animal models of the disease.
Keywords: Ischemic stroke, Translational medicine, Machine learning, Stroke segmentation Neuroimaging
 Global reach, higher impact
Global reach, higher impact