13.3
Impact Factor
Theranostics 2021; 11(7):3060-3073. doi:10.7150/thno.54150 This issue Cite
Research Paper
Systematic analysis of enhancer regulatory circuit perturbation driven by copy number variations in malignant glioma
1. Key Laboratory of Tropical Translational Medicine of Ministry of Education, College of Biomedical Information and Engineering, Hainan Medical University, Haikou 571199, China
2. College of Bioinformatics Science and Technology, Harbin Medical University, Harbin 150081, China
#These authors contributed equally to this work.
Abstract
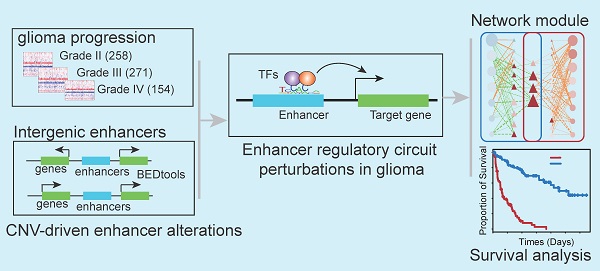
Background: Enhancers are emerging regulatory regions controlling gene expression in diverse cancer types. However, the functions of enhancer regulatory circuit perturbations driven by copy number variations (CNVs) in malignant glioma are unclear. Therefore, we aimed to investigate the comprehensive enhancer regulatory perturbation and identify potential biomarkers in glioma.
Results: We performed a meta-analysis of the enhancer centered regulatory circuit perturbations in 683 gliomas by integrating CNVs, gene expression, and transcription factors (TFs) binding. We found widespread CNVs of enhancers during glioma progression, and CNVs were associated with the perturbations of enhancer activities. In particular, the degree of perturbations for amplified enhancers was much greater accompanied by the glioma malignant progression. In addition, CNVs and enhancers cooperatively regulated the expressions of cancer-related genes. Genome-wide TF binding profiles revealed that enhancers were pervasively regulated by TFs. A network-based analysis of TF-enhancer-gene regulatory circuits revealed a core TF-gene module (58 interactions including seven genes and 14 TFs) that was associated survival of patients with glioma (p < 0.001). Finally, we validated this prognosis-associated TF-gene regulatory module in an independent cohort. In summary, our analyses provided new molecular insights for enhancer-centered transcriptional perturbation in glioma therapy.
Conclusion: Integrative analysis revealed enhancer regulatory perturbations in glioma and also identified a network module that was associated with patient survival, thereby providing novel insights into enhancer-centered cancer therapy.
Keywords: Enhancer, regulatory circuit, copy number variation, prognostic, glioma progression
 Global reach, higher impact
Global reach, higher impact