Impact Factor
Theranostics 2021; 11(8):3624-3641. doi:10.7150/thno.51795 This issue Cite
Research Paper
Genome-wide CRISPR/Cas9 knockout screening uncovers a novel inflammatory pathway critical for resistance to arginine-deprivation therapy
1. TMU Research Center of Cancer Translational Medicine, Taipei Medical University, Taipei 110, Taiwan.
2. CRISPR Gene Targeting Core Lab, Taipei Medical University, Taipei 110, Taiwan.
3. Department of Medical Research & Education, Taipei Medical University - Shuang Ho Hospital, New Taipei City.
4. Department of Hematology & Oncology, Taipei Medical University - Shuang Ho Hospital, New Taipei City, Taiwan.
5. Graduate Institute of Clinical Medicine, College of Medicine, Taipei Medical University, Taipei City, Taiwan.
6. Institute of Biological Chemistry, Academia Sinica, Taipei, Taiwan.
7. Bioinformatics Core Facility, The University of Manchester, Manchester, United Kingdom.
8. Joint Biobank, Office of Human Research, Taipei Medical University, Taipei, Taiwan.
9. Ph.D. Program for Cancer Biology and Drug Discovery, College of Medical Science and Technology, Taipei Medical University, Taipei 110, Taiwan.
10. Graduate Institute of Cancer Biology and Drug Discovery, College of Medical Science and Technology, Taipei Medical University, Taipei 110, Taiwan.
11. Department of Biochemistry and Molecular Cell Biology, School of Medicine, College of Medicine, Taipei Medical University, Taipei, Taiwan.
12. Graduate Institute of Medical Sciences, College of Medicine, Taipei Medical University, Taipei, Taiwan.
13. The Ph.D. Program for Translational Medicine, College of Medical Science and Technology, Taipei Medical University, Taipei 110, Taiwan.
14. Department of Medical Laboratory Science and Biotechnology, Taipei Medical University, Taipei 110, Taiwan.
15. Department of Biotechnology, College of Medical and Health Science, Asia University, Taichung, Taiwan.
16. Department of Medical Research, China Medical University Hospital, China Medical University, Taichung, Taiwan.
17. Department and Graduate Institute of Aquaculture, National Kaohsiung University of Science and Technology, Kaohsiung 81157, Taiwan.
18. Department of Diabetes and Metabolic Diseases Research, Irell & Manella Graduate School of Biological Sciences, Beckman Research Institute, City of Hope, Duarte, CA, United States.
19. Institute of Molecular and Genomic Medicine, National Health Research Institutes, Zhunan, Miaoli County 350, Taiwan.
20. Department of Biochemistry and Molecular Medicine, Comprehensive Cancer Center, University of California at Davis, Sacramento, CA 95817, United States.
Abstract
Arginine synthesis deficiency due to the suppressed expression of ASS1 (argininosuccinate synthetase 1) represents one of the most frequently occurring metabolic defects of tumor cells. Arginine-deprivation therapy has gained increasing attention in recent years. One challenge of ADI-PEG20 (pegylated ADI) therapy is the development of drug resistance caused by restoration of ASS1 expression and other factors. The goal of this work is to identify novel factors conferring therapy resistance.
Methods: Multiple, independently derived ADI-resistant clones including derivatives of breast (MDA-MB-231 and BT-549) and prostate (PC3, CWR22Rv1, and DU145) cancer cells were developed. RNA-seq and RT-PCR were used to identify genes upregulated in the resistant clones. Unbiased genome-wide CRISPR/Cas9 knockout screening was used to identify genes whose absence confers sensitivity to these cells. shRNA and CRISPR/Cas9 knockout as well as overexpression approaches were used to validate the functions of the resistant genes both in vitro and in xenograft models. The signal pathways were verified by western blotting and cytokine release.
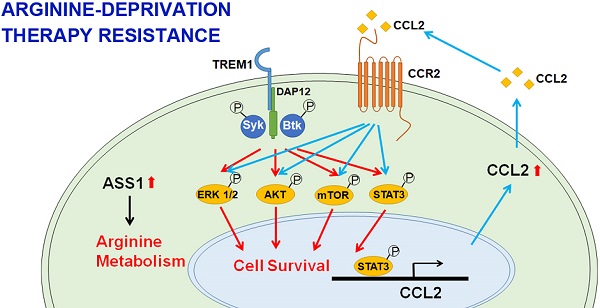
Results: Based on unbiased CRISPR/Cas9 knockout screening and RNA-seq analyses of independently derived ADI-resistant (ADIR) clones, aberrant activation of the TREM1/CCL2 axis in addition to ASS1 expression was consistently identified as the resistant factors. Unlike ADIR, MDA-MB-231 overexpressing ASS1 cells achieved only moderate ADI resistance both in vitro and in vivo, and overexpression of ASS1 alone does not activate the TREM1/CCL2 axis. These data suggested that upregulation of TREM1 is an independent factor in the development of strong resistance, which is accompanied by activation of the AKT/mTOR/STAT3/CCL2 pathway and contributes to cell survival and overcoming the tumor suppressive effects of ASS1 overexpression. Importantly, knockdown of TREM1 or CCL2 significantly sensitized ADIR toward ADI. Similar results were obtained in BT-549 breast cancer cell line as well as castration-resistant prostate cancer cells. The present study sheds light on the detailed mechanisms of resistance to arginine-deprivation therapy and uncovers novel targets to overcome resistance.
Conclusion: We uncovered TREM1/CCL2 activation, in addition to restored ASS1 expression, as a key pathway involved in full ADI-resistance in breast and prostate cancer models.
Keywords: arginine starvation, ADI resistance, CRISPR/Cas9, TREM1, CCL2
 Global reach, higher impact
Global reach, higher impact