Impact Factor
Theranostics 2021; 11(15):7491-7506. doi:10.7150/thno.59196 This issue Cite
Research Paper
Microbial and genetic-based framework identifies drug targets in inflammatory bowel disease
1. West China School of Basic Medical Sciences & Forensic Medicine, Sichuan University, Chengdu, Sichuan 610041, China.
2. Department of Biomedical Sciences, School of Medicine and Health Sciences, University of North Dakota, Grand Forks, ND 58202, USA.
3. Department of Neurology, University of Michigan, Ann Arbor, MI 48109, USA.
4. Medical Research Institute, Wuhan University, Wuhan 430071, China.
5. State Key Laboratory of Trauma, Burns and Combined Injury, Institute of Surgery Research, Daping Hospital, Army Medical University, Chongqing 400038, China.
6. State Key Laboratory of Biotherapy and Cancer Center, West China Hospital, Sichuan University, and Collaborative Innovation Center for Biotherapy, Chengdu, Sichuan 610041, China.
*These authors contributed equally to this work.
Abstract
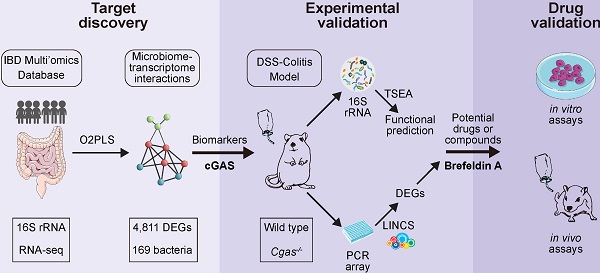
Rationale: With increasing incidence and prevalence of inflammatory bowel disease (IBD), it has become one of the major public health threats, and there is an urgent need to develop new therapeutic agents. Although the pathogenesis of IBD is still unclear, previous research has provided evidence for complex interplays between genetic, immune, microbial, and environmental factors. Here, we constructed a gene-microbiota interaction-based framework to discover IBD biomarkers and therapeutics.
Methods: We identified candidate biomarkers for IBD by analyzing the publicly available transcriptomic and microbiome data from IBD cohorts. Animal models of IBD and diarrhea were established. The inflammation-correlated microbial and genetic variants in gene knockout mice were identified by 16S rRNA sequences and PCR array. We performed bioinformatic analysis of microbiome functional prediction and drug repurposing. Our validation experiments with cells and animals confirmed anti-inflammatory properties of a drug candidate.
Results: We identified the DNA-sensing enzyme cyclic GMP-AMP synthase (cGAS) as a potential biomarker for IBD in both patients and murine models. cGAS knockout mice were less susceptible to DSS-induced colitis. cGAS-associated gut microbiota and host genetic factors relating to IBD pathogenesis were also identified. Using a computational drug repurposing approach, we predicted 43 candidate drugs with high potency to reverse colitis-associated gene expression and validated that brefeldin-a mitigates inflammatory response in colitis mouse model and colon cancer cell lines.
Conclusions: By integrating computational screening, microbiota interference, gene knockout techniques, and in vitro and in vivo validation, we built a framework for predicting biomarkers and host-microbe interaction targets and identifying repurposing drugs for IBD, which may be tested further for clinical application. This approach may also be a tool for repurposing drugs for treating other diseases.
Keywords: Inflammatory bowel disease, cyclic GMP-AMP synthase (cGAS), host transcriptome-microbiome interaction, drug repurposing, brefeldin-a
 Global reach, higher impact
Global reach, higher impact