Impact Factor
Theranostics 2021; 11(20):9752-9771. doi:10.7150/thno.63806 This issue Cite
Research Paper
USP32 confers cancer cell resistance to YM155 via promoting ER-associated degradation of solute carrier protein SLC35F2
1. Graduate School of Biomedical Science and Engineering, Hanyang University, Seoul, South Korea.
2. College of Medicine, Hanyang University, Seoul, South Korea.
Abstract
Background: The most commonly preferred chemotherapeutic agents to treat cancers are small-molecule drugs. However, the differential sensitivity of various cancer cells to small molecules and untargeted delivery narrow the range of potential therapeutic applications. The mechanisms responsible for drug resistance in a variety of cancer cells are also largely unknown. Several deubiquitinating enzymes (DUBs) are the main determinants of drug resistance in cancer cells.
Methods: We used CRISPR-Cas9 to perform genome-scale knockout of the entire set of genes encoding ubiquitin-specific proteases (USPs) and systematically screened for DUBs resistant to the clinically evaluated anticancer compound YM155. A series of in vitro and in vivo experiments were conducted to reveal the relationship between USP32 and SLC35F2 on YM155-mediated DNA damage in cancer cells.
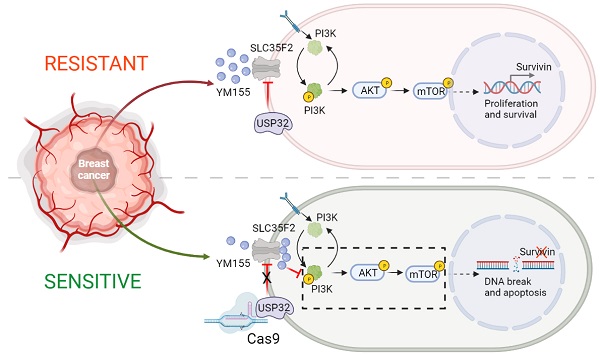
Results: CRISPR-based dual-screening method identified USP32 as a novel DUB that governs resistance for uptake of YM155 by destabilizing protein levels of SLC35F2, a solute-carrier protein essential for the uptake of YM155. The expression of USP32 and SLC35F2 was negatively correlated across a panel of tested cancer cell lines. YM155-resistant cancer cells in particular exhibited elevated expression of USP32 and low expression of SLC35F2.
Conclusion: Collectively, our DUB-screening strategy revealed a resistance mechanism governed by USP32 associated with YM155 resistance in breast cancers, one that presents an attractive molecular target for anti-cancer therapies. Targeted genome knockout verified that USP32 is the main determinant of SLC35F2 protein stability in vitro and in vivo, suggesting a novel way to treat tumors resistant to small-molecule drugs.
Keywords: DNA damage, dose response, human tumor tissues, cell apoptosis, drug transport-cargo
 Global reach, higher impact
Global reach, higher impact