Impact Factor
Theranostics 2021; 11(20):9847-9858. doi:10.7150/thno.65064 This issue Cite
Research Paper
Multiple regression analysis of a comprehensive transcriptomic data assembly elucidates mechanically- and biochemically-driven responses to focused ultrasound blood-brain barrier disruption
1. Department of Biomedical Engineering, University of Virginia, Charlottesville, VA.
2. Department of Radiology & Medical Imaging, University of Virginia, Charlottesville, VA.
Abstract
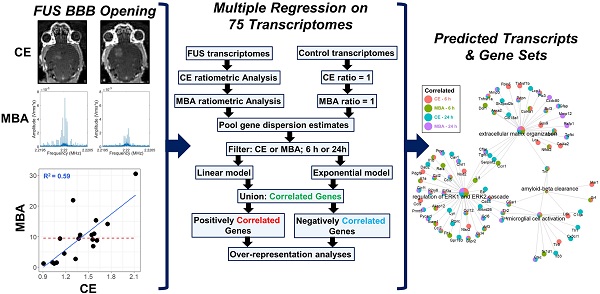
Background: Focused ultrasound (FUS) blood brain barrier disruption (BBBD) permits the noninvasive, targeted, and repeatable delivery of drugs to the brain. FUS BBBD also elicits secondary responses capable of augmenting immunotherapies, clearing amyloid-β and hyperphosphorylated tau, and driving neurogenesis. Leveraging these secondary effects will benefit from an understanding of how they correlate to the magnitude of FUS BBBD and are differentially affected by the mechanical and biochemical stimuli imparted during FUS BBBD.
Methods: We aggregated 75 murine transcriptomes in a multiple regression framework to identify genes expressed in proportion to biochemical (i.e. contrast MR image enhancement (CE)) or mechanical (i.e. harmonic acoustic emissions from MB-activation (MBA)) stimuli associated with FUS BBBD. Models were constructed to control for potential confounders, such as sex, anesthesia, and sequencing batch.
Results: MBA and CE differentially predicted expression of 1,124 genes 6 h or 24 h later. While there existed overlap in the transcripts correlated with MBA vs CE, MBA was principally predictive of expression of genes associated with endothelial reactivity while CE chiefly predicted sterile inflammation gene sets. Over-representation analysis identified transcripts not previously linked to BBBD, including actin filament organization, which is likely important for BBB recovery. Transcripts and pathways associated with neurogenesis, microglial activation, and amyloid-β clearance were significantly correlated to BBBD metrics.
Conclusions: The secondary effects of BBBD may have the potential to be tuned by modulating FUS parameters during BBBD, and MBA and CE may serve as independent predictors of transcriptional reactions in the brain.
Keywords: focused ultrasound, blood-brain barrier, drug delivery, transcriptomics, neurogenesis
 Global reach, higher impact
Global reach, higher impact