Impact Factor
Theranostics 2024; 14(5):2151-2166. doi:10.7150/thno.94161 This issue Cite
Research Paper
FSH induces EMT in ovarian cancer via ALKBH5-regulated Snail m6A demethylation
1. Guangzhou Institute of Cardiovascular Disease, the Second Affiliated Hospital, School of Basic Medical Sciences, Guangzhou Medical University, Guangzhou, China.
2. Department of Gynecology and Obstetrics, The Sixth Affiliated Hospital, Guangzhou Medical University, Guangzhou, China.
3. Department of Gynecology and Obstetrics, Guangzhou First People's Hospital, Guangzhou, China.
#Equal contribution
Received 2024-1-11; Accepted 2024-2-26; Published 2024-3-3
Abstract
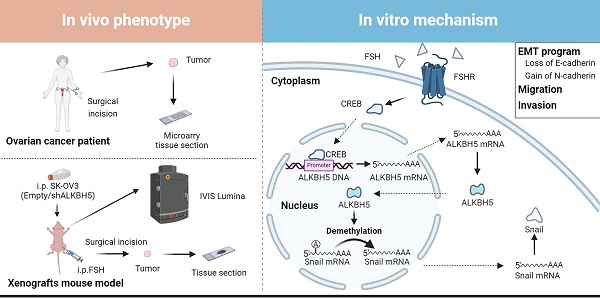
Background: The therapeutic benefits of targeting follicle-stimulating hormone (FSH) receptor in treatment of ovarian cancer are significant, whereas the role of FSH in ovarian cancer progresses and the underlying mechanism remains to be developed.
Methods: Tissue microarray of human ovarian cancer, tumor xenograft mouse model, and in vitro cell culture were used to investigate the role of FSH in ovarian carcinogenesis. siRNA, lentivirus and inhibitors were used to trigger the inactivation of genes, and plasmids were used to increase transcription of genes. Specifically, pathological characteristic was assessed by histology and immunohistochemistry (IHC), while signaling pathway was studied using western blot, quantitative RT-PCR, and immunofluorescence.
Results: Histology and IHC of human normal ovarian and tumor tissue confirmed the association between FSH and Snail in ovarian cancer metastasis. Moreover, in epithelial ovarian cancer cells and xenograft mice, FSH was showed to promote epithelial mesenchymal transition (EMT) progress and metastasis of ovarian cancer via prolonging the half-life of Snail mRNA in a N6-methyladenine methylation (m6A) dependent manner, which was mechanistically through the CREB/ALKBH5 signaling pathway.
Conclusions: These findings indicated that FSH induces EMT progression and ovarian cancer metastasis via CREB/ALKBH5/Snail pathway. Thus, this study provided new insight into the therapeutic strategy of ovarian cancer patients with high level of FSH.
Keywords: Follicle-stimulating hormone, Epithelial ovarian cancer, Epithelial-mesenchymal transition, Snail, ALKBH5
Introduction
Ovarian cancer is most lethal gynecological malignancies [1], however, its pathogenic mechanism is remained to be elucidated [2]. Theories of ovarian cancer pathogenesis, proposed that extended exposure to high levels of gonadotropin [3], e.g., follicle-stimulating hormone (FSH), may excessively stimulate ovarian epithelium growth, ultimately contributing to tumor development [4, 5]. These theories were supported by the observation that FSH was increased in peritoneal fluid of ovarian cancer patients [6]. Studies have demonstrated FSH promoted proliferation [7], migration, and invasion [8] of epithelial ovarian cancer cell via FSH receptor (FSHR) [9]. Antagonist of gonadotropin [10] and antibodies targeting surface-expressed FSHR [11-13] were shown the potential role of FSHR in ovarian cancer therapy, evident by suppression of tumorigenesis in mouse model of ovarian cancer. Nevertheless, in post-menopausal women undergoing hormone replacement therapy, a reduction in FSH was correlated with an elevated risk of ovarian cancer [14]. Unexpectedly, study showed that FSH inhibited proliferation in normal ovarian epithelial cells from both premenopausal and postmenopausal women [15], which did not support the mentioned theories that gonadotrophin is directly involved in ovarian carcinogenesis. These contradictions underscore the critical need of investigation on the role of FSH in ovarian cancer and its underlying mechanisms.
The major cause of mortality in ovarian cancer patient is tumor metastasis [16]. In metastasis, cells go through epithelial-mesenchymal transition (EMT), which involves epithelial cells losing their original surface markers and gaining characteristics of mesenchymal cells, thereby allowing tumors to detach from their original location and spreading to distant areas. EMT is collectively regulated by various transcription factors, e.g., Snail, Slug, Twist, ZEB1, and ZEB2 [17]. Recent studies reported that EMT and its transcription factors were modulated by N6-methyladenosine (m6A) modifications [18-20], and intervention in m6A modifications demonstrated a potential cancer therapy [21].
mRNA m6A modification, a prevalent post-transcriptional modification, is dynamically regulated by methyltransferases (e.g., METTL3, METTL14 and WTAP), demethylases (e.g., ALKBH5 and FTO), and readers (e.g., YTHDF and YTHDC family) [22]. In the context of ovarian cancer, m6A plays a crucial role in mRNA synthesis, transcription, and degradation of oncogenes and tumor suppressor genes [23-25], ultimately influencing various tumorigenic processes, including cancer cell proliferation, migration, metastasis [26], chemoresistance [27] and immune evasion [28]. The modification of mRNA by m6A in EMT transcription factors also impacts EMT and tumor metastasis [29-31]. However, there is currently no clear evidence on whether FSH can regulate m6A modification on EMT transcription factors and influence ovarian cancer metastasis.
In this study, we observed that FSH induced the upregulation of the m6A demethylase ALKBH5 and led to the promotion of EMT in ovarian cancer cells. Further investigation revealed that ALKBH5 regulated the transcription factor Snail, a known suppressor of E-cadherin. Additionally, we uncovered that FSH upregulated ALKBH5 expression through cAMP-response element binding protein (CREB), resulting in the removal of m6A modifications on Snail mRNA. This process enhanced Snail expression and contributed to the initiation of EMT. The crucial role of ALKBH5 in FSH-induced ovarian cancer metastasis was confirmed in both animal model and clinical samples. Our findings suggested that intervening m6A modifications to modulate Snail could be a potential therapeutic approach for ovarian cancer.
Results
FSH induced EMT in epithelial ovarian cancer cells via demethylase ALKBH5
To get a broad view of the influence of FSH on the ovarian cancer, immunohistochemistry (IHC) of FSH receptor (FSHR) was performed on tissue microarray (TMA) of ovarian tumors derived from 160 ovarian cancer patients. As shown in Table 1 and Figure S1A, high expression of FSHR had a higher probability of metastasis. TGF-β has been determined to be a major inducer of EMT during cancer metastasis [32]. Our result showed that FSH induced an EMT phenotype comparable to TGF-β in epithelial ovarian cancer cells (Figure 1A, B and Figure S1B). Moreover, the changes in E-cadherin and N-cadherin expression confirmed FSH-induced EMT in multiple epithelial ovarian cancer cell lines (Figure 1C, D and Figure S1C-E). Among the cell lines, SK-OV3 showed an obvious response and was then chose as a major in vitro tool to investigate the role of FSH in EMT.
Association between FSHR and clinicopathologic characterizations in ovarian cancer
| FSHR expression | ||||
|---|---|---|---|---|
| Characteristics | Patients | Low | High | P-value |
| Ages | ||||
| ≤ | 57 | 24 | 33 | 0.49a |
| >50 | 103 | 45 | 58 | |
| T Stage | ||||
| T1+T2 | 119 | 51 | 68 | 0.525a |
| T3+T4 | 41 | 18 | 23 | |
| Node Metastasis | ||||
| Absent | 123 | 58 | 65 | 0.045a |
| Present | 37 | 11 | 26 | |
| CA-125 | ||||
| ≤ | 47 | 17 | 30 | 0.166a |
| >200 | 113 | 52 | 61 | |
| Tumor Grade | ||||
| Ⅰ | 24 | 14 | 10 | 0.061a |
| Ⅱ | 33 | 18 | 15 | |
| Ⅲ | 103 | 38 | 65 | |
| Histological Type | ||||
| Serous | 51 | 24 | 27 | 0.654b |
| Mucinous | 107 | 45 | 62 | |
| Others | 2 | 1 | 1 | |
Number in the table represented the case number of patients.
a: Pearson Chi-Square test; b: Fisher's exact test;
FSH induced EMT in epithelial ovarian cancer cells via demethylase ALKBH5. (A, B) Immunofluorescence and bright filed images of SK-OV3 cells treated with FSH (75 mIU) or TGF-β (10 ng/μl) for 48 hours, scale bar = 20 µm. (C-H) SK-OV3 cells were treated with different concentrations of FSH for various time points. (C, D) Western blots for E-cadherin and N-cadherin. (E, F) Western blots for METTL3, METTL14, ALKBH5, FTO and YTHDF1/2/3. (G, H) Western blot quantification of ALKBH5 intensity measurements (n = 3), data was analyzed by Mann-Whitney U test. *P < 0.05, **P < 0.01 versus CON. (I) Western blots of E-cadherin, N-cadherin and ALKBH5 in SK-OV3 cells treated with FSH (75 mIU, 48 h) after transfection of ALKBH5 siRNA (siALKBH5). (J-L) SK-OV3 cells transfected with ALKBH5 plasmid followed by FSH treatment (75 mIU, 48 h), n = 3 - 4 per group. (J) Immunofluorescence of E-cadherin- and N-cadherin-stained SK-OV3 cells. Scale bar = 20 µm. (K, L) Wound healing and Transwell assays were performed in SK-OV3 cells, data was analyzed by one-way ANOVA test. *P < 0.05, **P < 0.01, ***P < 0.001 versus CON transfected with Vector.
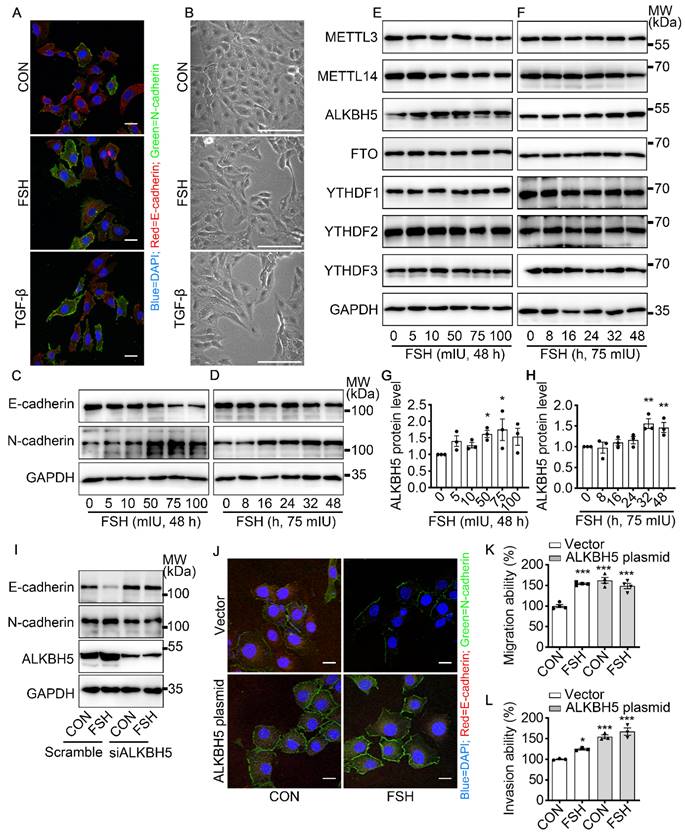
Emerging studies have shown that m6A modification plays an essential role in various cancer progressions [33]. To determine whether m6A modification was involved in FSH-induced EMT progression, we first investigated the expression patterns of seven key regulators for m6A modification, our results show that both demethylase ALKBH5 and methyltransferase METTL14 respond to FSH in epithelial ovarian cancer cells (Figure 1E-H and Figure S1F, H, I). Through a systematic analysis of the expression patterns of EMT and m6A regulators, Yanna Zhang et al. revealed a synergistic effect between ALKBH5 and EMT regulators on ovarian cancer development [34]. Our results confirmed that ALKBH5 had higher expression in epithelial ovarian cancer cell lines (e.g., SK-OV3, ES-2, and A2780) compared with the normal ovary cell line (e.g., ISOE-80) (Figure S1G). The silence of ALKBH5 by siRNA inhibited the regulation of FSH on E-cadherin and N-cadherin expressions (Figure 1I, J and Figure S1J), while overexpression of ALKBH5 enhanced the regulation of FSH on E-cadherin and N-cadherin expressions, thereby reducing EMT-related activity in SK-OV3 and A2780 cells (Figure 1K, L and Figure S1K-M). Taken together, our result showed that demethylase ALKBH5 is essential for FSH-induced EMT progression in epithelial ovarian cancer cells.
Snail is a critical transcription factor in FSH-induced EMT
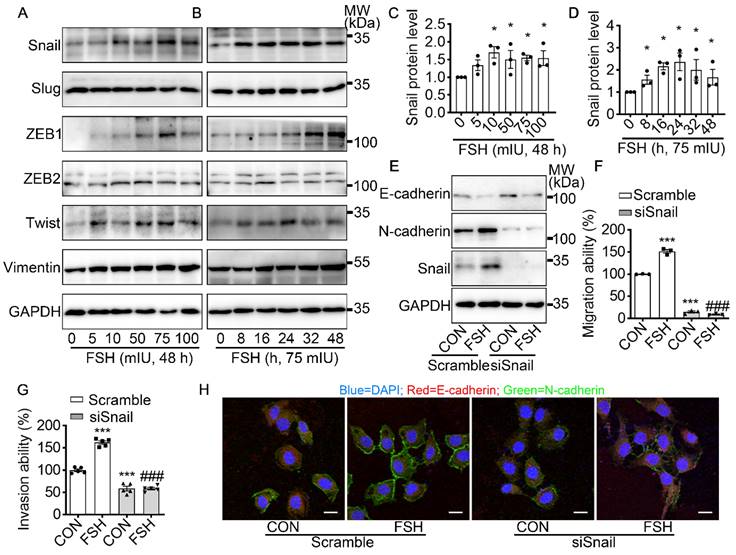
To further explore the underlying mechanism of FSH-induced EMT, we monitored the expression pattern of six main EMT-related protein after FSH treatment. Our result demonstrated that both EMT transcription factors (Snail, ZEB1, Twist) and EMT protein markers (Vimentin) were responds to FSH in epithelial ovarian cancer cells (Figure 2A-D and Figure S2A-D). Snail, a key inducer of EMT, have been reported to be involved in the chemoresistance [35], stemness, and metastasis [36] in ovarian cancer. To confirm the role of Snail in FSH-induced EMT, Snail siRNA was transfected to knock down Snail expression. The regulatory effects of FSH on E- and N-cadherin were abolished by knocking down Snail, indicating that FSH-induced EMT was dependent on Snail signaling (Figure 2E, H and Figure S2E, F). Similarly, FSH-induced migration and invasion of SK-OV3 were attenuated by Snail silencing (Figure 2F, G and Figure S2G, H). These results indicated that FSH promotes EMT of epithelial ovarian cancer cells via Snail signaling.
Snail is a critical transcription factor in FSH-induced EMT. (A-D) SK-OV3 cells were treated with different concentrations of FSH for various time points. (A, B) Western blots for Snail, Slug, ZEB1/2, Twist and Vimentin. (C, D) Western blot quantification of Snail intensity measurements (n = 3), data was analyzed by Mann-Whitney U test. *P < 0.05 versus CON. (E-H) SK-OV3 cells were treated with FSH (75 mIU, 48 h) after transfection of scramble or Snail siRNA (siSnail). (E) Western blots of E-cadherin, N-cadherin and Snail in SK-OV3 cells. (F, G) Wound healing and Transwell assays were performed. Quantitative data of healing area (n = 3) and cell invasion (n = 5) were analyzed by one-way ANOVA test. ***P < 0.001 versus CON group transfected with scramble siRNA, ###P < 0.001 versus FSH group transfected with scramble siRNA. (H) Immunofluorescence of SK-OV3 cells stained with E-cadherin and N-cadherin. Scale bar = 20 µm.
m6A modification of Snail mRNA is essential for FSH-induced EMT via ALKBH5
To revealed the specific role of Snail in FSH-induced EMT via ALKBH5, we first preformed luciferase assay for Snail promoter activity. However, we observed that FSH was not able to increase the transcriptional activity of Snail (Figure 3A). We then used actinomycin D to evaluate the stability of Snail mRNA. Result showed that FSH prolonged the half-life of Snail mRNA (Figure 3B). Studies had shown that m6A modification can extend the half-life of mRNA [37-39], therefore we determined whether m6A modification participate in increased mRNA and protein expression of Snail induced by FSH. Indeed, FSH was able to decrease m6A content of total RNA in SK-OV3 cells as indicated by methylation quantification assay (Figure 3C). According to the substrates preference of ALKBH5 [40], sequence analysis was performed to predict Snail containing consensus m6A motifs located in the CDS2 region (Figure 3G). m6A-Immunoprecipitation (MeRIP) qPCR result confirmed that FSH reduces m6A modification of Snail mRNA in the predicted regions (Figure 3D). Sequentially, we confirmed ALKBH5 reduced both the mRNA expression and m6A modification level of Snail (Figure 3E and Figure S3A). Furthermore, we found ALKBH5 prolonged the half-life of Snail mRNA (Figure 3F). Our result also showed that ALKBH5 prolong Snail protein half-life using protein biosynthesis inhibitor cycloheximide (CHX) pulse-chase assay (Figure S3C, D). The role of ubiquitination in ALKBH5 regulated Snail expression was also assessed via using proteasome inhibitor (MG132), result showed progression of ubiquitination had limited influence on ALKBH5 regulated Snail expression (Figure S3B).
m6A modification of Snail mRNA is essential for FSH-induced EMT via ALKBH5. (A) Snail promoter activity was measured by luciferase assay in 293T cells transfected with FSHR plasmid before PBS or FSH (75 mIU, 48 h) treatment (n = 5), data was analyzed for statistical difference by two-tailed unpaired t test, ns, no significant. (B) qRT-PCR of Snail mRNA in SK-OV3 cells with pretreatment of FSH (75 mIU, 48 h) and treatment of Actinomycin D (ActD) for various time points (n = 3). (C) N6-methyladenosine (m6A) content of total RNA in SK-OV3 cells with FSH treatment (75 mIU, 48 h), n = 3 per group, data was analyzed by two-tailed unpaired t test. **P < 0.01. (D) m6A RIP-qPCR analysis of Snail mRNA in SK-OV3 cells treated with FSH (75 mIU, 48 h), n = 5 per group, data was analyzed by two-tailed unpaired t test. *P < 0.05. (E) m6A RIP-qPCR analysis of Snail mRNA in SK-OV3 cells transfected with ALKBH5 siRNA (siALKBH5) (n = 3), data was analyzed by two-tailed unpaired t test. *P < 0.05. (F) qRT-PCR of Snail mRNA in SK-OV3 cells pretreated with Actinomycin D (ActD) for various time points (n = 7), data was analyzed by two-tailed unpaired t test, *P < 0.05, **P < 0.01, ***P < 0.001. (G) Scheme showed locations of the m6A motifs within the Snail mRNA, and A to T mutation within m6A consensus site in Snail mRNA. (H-K) SK-OV3 cells were transfected with empty vector (Vector), Snail-CDs (CDs) and Snail-mut (Mut) plasmid. Data was analyzed by two-tailed unpaired t test, **P < 0.01, ***P < 0.001. versus Vector. #P < 0.05, ###P < 0.001 versus CDs. (H) qRT-PCR of Snail mRNA. (I) qRT-PCR of E-cadherin mRNA. (J) Western blots for E-cadherin and Snail. (K) RT-PCR of Snail mRNA in SK-OV3 cells transfected with Snail-CDs (CDs) and Snail-mut (Mut) plasmid and treatment of Actinomycin D (ActD) for various time points (n = 3). (L) Western blots of E-cadherin, N-cadherin, and Snail in SK-OV3 cells treated with FSH (75 mIU, 48 h), after transfection of Snail-CDs or Snail-mut plasmid. (M) Western blots for E-cadherin, N-cadherin and Snail in SK-OV3 cells co-transfected with ALKBH5 plasmid and wild type Snail (Snail-CDs) or CDS region mutated (Snail-mut) plasmid.
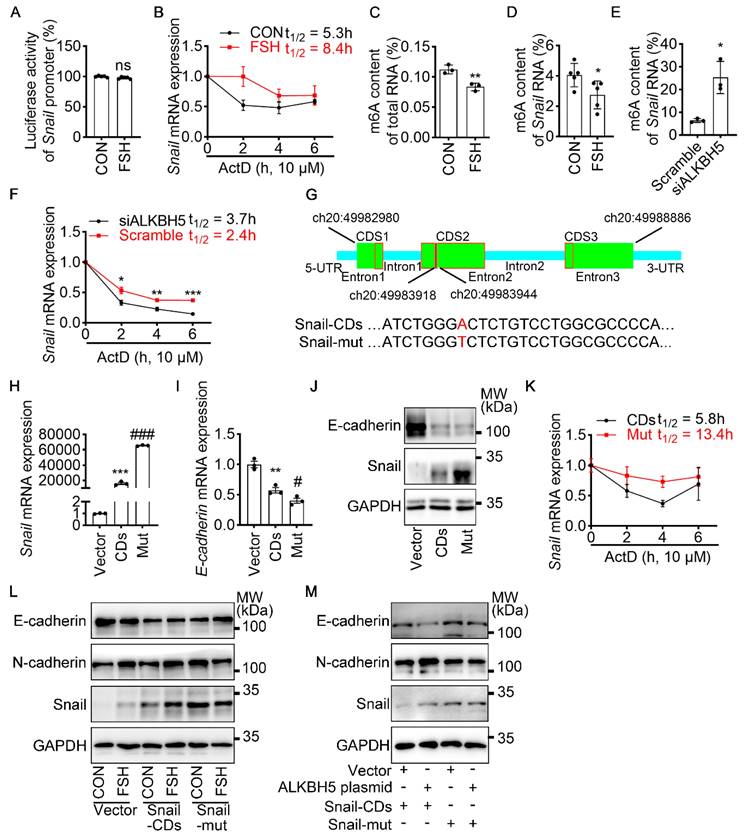
To further confirm the role of m6A modification on Snail mRNA in FSH-regulated EMT, a wild-type Snail (Snail-CDs) or mutant (Snail-mut) plasmid was constructed and transfected into SK-OV3. Both Snail-mut and Snail-CDs plasmid transfection decreased E-cadherin mRNA and protein level compared with vector. Compared with Snail-CDs plasmid transfection, an evaluated Snail expression was observed in Snail-mut plasmid transfection, interestingly, this increased Snail expression induced by the Snail-mut was not able to further decrease the expression of E-cadherin (Figure 3H-J, Figure S3E). Additionally, actinomycin D assay was performed and showed that m6A modification played an essential role in the stability of Snail mRNA (Figure 3K). Further result showed that FSH-induced EMT was blocked with by mutation of Snail on the m6A modification site (Figure 3L). We next assessed whether FSH reduces m6A modification of Snail mRNA via demethylase ALKBH5. Snail-CDS or Snail-mut was co-transfected with ALKBH5 plasmid in SK-OV3 cells. We found a significant decrease in E-cadherin and increase in N-cadherin, while co-transfection of Snail-mut and ALKBH5 plasmid was not able to recapitulate this phenomenon (Figure 3M).
To determine the essential role of ALKBH5 in the FSH-induced EMT process, ALKBH5 siRNA was solely- or co-transfected with Snail plasmid into SK-OV3 cells. We observed that knockdown of ALKBH5 inhibited FSH-promoted EMT, migration and invasion, while these inhibitory effects were rescued by the overexpression of Snail (Figure S3F-H). Thus, we confirmed that ALKBH5 was implicated in FSH-induced EMT by regulating Snail in a m6A-dependent manner. ALKBH5 H204A (catalytic inactive mutation) plasmid transfection [41] further demonstrated that the demethylase activity of ALKBH5 was essential in EMT (Figure S3I).
FSH increases ALKBH5/Snail expression and triggers EMT via CREB
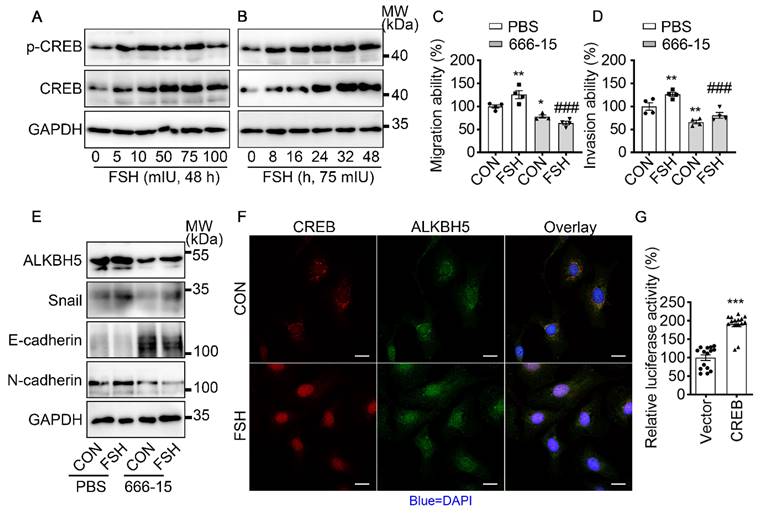
It is well established that cAMP response element-binding protein (CREB) is a classical transcription factor that can be phosphorylated and activated by FSH. To determine if CREB is involved in FSH-induced EMT, we first confirmed FSH activated CREB phosphorylation in a time- and concentration-dependent manner (Figure 4A, B and Figure S4A, B, I). Phosphorylated CREB acted as the transcription factor to regulate various gene transcription [42]. To explore whether ALKBH5 gene transcription was regulated by CERB, a selective inhibitor of CERB, 666-15, was used to inhibit CREB-mediated gene transcription [43]. Our result showed FSH activates CREB to increase ALKBH5 and Snail expression, thereby enhancing migration and invasion of ovarian cancer (Figure 4C-E and Figure S4C-H, J). Moreover, bioinformatics analysis (http://jaspar.genereg.net) predicted that CREB binding site locates in ALKBH5 gene promoter region (Figure S4K). Luciferase assay further confirmed that transcriptional activity of ALKBH5 was regulated by CREB (Figure 4G). Moreover, we monitored that FSH induced accumulation of CREB and ALKBH5 in nucleus (Figure 4F). These findings indicated that CREB-triggered ALKBH5 gene transcription plays an important role in FSH-induced EMT and tumor metastasis.
FSH activates CREB to increase ALKBH5 expression. (A, B) Western blots of p-CREB and CREB in SK-OV3 cells treated with different concentrations of FSH for various time points. (C, D) Wound healing and invasion assays were performed on SK-OV3 cells with pretreatment of CREB inhibitor 666-15 (10 μM, 1 h), then with FSH treatment (75 mIU, 48 h), n = 4 per group, data was analyzed for statistical difference by one-way ANOVA test. *P < 0.05, **P < 0.01 versus CON+PBS, ###P < 0.001 versus FSH+PBS. (E) Western blots of ALKBH5, Snail, E-cadherin and N-cadherin expression in SK-OV3 cells with pretreatment of 666-15 (10 μM, 1 h) and treatment of FSH. (F) Immunofluorescence for CREB and ALKBH5 in SK-OV3 cells treated with FSH (75 mIU, 48 h). Scale bar = 20 µm. (G) ALKBH5 promoter activity was measured by luciferase assay in SK-OV3 cells transfected with Vector or CREB plasmid (n = 15), data was analyzed by two-tailed unpaired t test. ***P < 0.001.
FSH enhances EMT and metastasis of ovarian cancer via ALKBH5 in model of xenografts
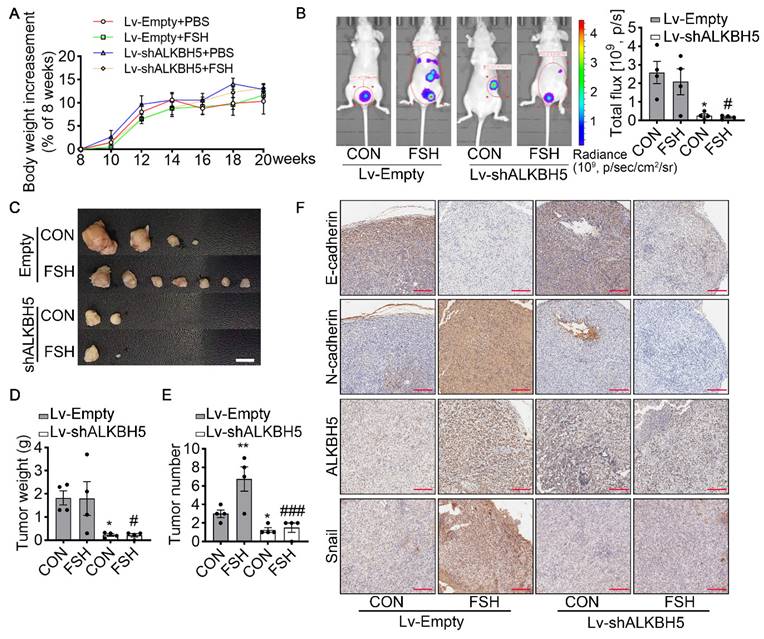
To verify the effect of ALKBH5 in FSH-promoted progression of epithelial ovarian cancer in vivo, the ovariectomized nude mice were intraperitoneally injected with SK-OV3 cells transfected with empty lentivirus (Lv-Empty) or ALKBH5 shRNA lentivirus (Lv-shALKBH5) to establish peritoneal metastatic models of ovarian cancer (Figure S5A). One week after the development of xenograft, mice were treated with FSH for 14 weeks (Figure S5B). During the 14 weeks treatment, there was no significant different in body weight among the 4 groups (Figure 5A). At the endpoint of experiment, in vivo imaging system (IVIS) was performed to assess the presence of peritoneal metastases. As expected, FSH increased the risk of distant metastasis and number of tumors (Figure 5B). However, ALKBH5 knockdown significantly blocked the effect of FSH on the risk of distant metastasis and tumor number in xenograft mice (Figure 5C-E). IHC result showed that FSH increased expression of N-cadherin, ALKBH5, Snail and decreased expression of E-cadherin in tumor tissues, while these effects were reversed by the knockdown of ALKBH5 (Figure 5F). Ki-67 protein has been widely used as a proliferation marker for human tumor cells. IHC of Ki-67 was performed and result showed both FSH treatment and Lv-ALKBH5 transfection had comparable influence on the proliferation level in SK-OV3 xenografts (Figure S5C). Taken together, our finding indicated that FSH increases ALKBH5 to enhance EMT and metastasis of ovarian cancer.
ALKBH5 is essential for FSH-induced EMT and metastasis of ovarian cancer in mouse xenografts model. (A-F) SK-OV3 cells were transfected with a control (Lv-Empty) or shALKBH5 (Lv-shALKBH5) luciferase-tagged lentivirus. SK-OV3 cells were then implanted to bilateral ovariectomized (OVX) nude mice to develop a xenografts model for FSH treatment (3 IU/day per mouse for 14 weeks). (A) Body increasement was calculated from the body weight of 8 weeks (n = 4). (B) In vivo bioluminescence imaging was performed to monitor tumor burden, and the bioluminescent signal was quantified and analyzed by Mann-Whitney U test (n = 4). *P < 0.05 versus CON group with Lv-Empty transfection. #P < 0.05 versus FSH group with Lv-Empty transfection. (C-E) Tumors of the SK-OV3 tumor-bearing mice were excised, imaged and weighed (n = 4). Scale bar = 1 cm. Data was analyzed by Mann-Whitney U test. *P < 0.05, **P < 0.01 versus CON group with Lv-Empty transfection. #P < 0.05, ###P < 0.001 versus FSH group with Lv-Empty transfection. (F) SK-OV3 xenografts were sectioned and stained for E-cadherin, N-cadherin, ALKBH5 and Snail expression by immunohistochemistry. Scale bar = 200 µm.
Both FSHR and Snail expression were positively associated with ALKBH5 expression in ovarian cancer
To determine the role of ALKBH5 and Snail expression in clinicopathologic characteristics of ovarian cancer, IHC of ALKBH5 and Snail were performed on two adjacent TMA containing 88 ovarian tumors and 8 normal ovarian tissues of 96 individual patients (Table 2 and Figure 6A). Based on low and high expression of ALKBH5, patients were divided into two groups. As shown in Table 2, high expression of ALKBH5 was positively associated with ovarian cancer stages (P = 0.019), node metastasis (P = 0.04), distant metastasis (P < 0.001), peritoneal seeding (P = 0.001), ascites formation (P < 0.001), primary tumor origin (P = 0.04), histological type (P = 0.037). By performing IHC of Snail and ALKBH5 on the TMA, we further determined the correlation between Snail and ALKBH5 expression using Cramér's V association analysis (V = 0.711, P < 0.001, Figure 6B). As shown in Table 2, high expression of Snail was correlated to patients age (P = 0.019), ovarian cancer stages (P < 0.001), distant metastasis (P < 0.001), peritoneal seeding (P < 0.001), ascites formation (P < 0.001), primary tumor origin (P = 0.038).
Association between ALKBH5, Snail, and clinicopathologic characterizations in ovarian cancer
| ALKBH5 expression | Snail expression | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Characteristics | Patients | Low | High | P-value | Patients | Low | High | P-value | |
| Ages | |||||||||
| ≤ | 24 | 18 | 6 | 0.218a | 24 | 16 | 8 | 0.019a | |
| >50 | 72 | 44 | 28 | 72 | 43 | 29 | |||
| T Stage | |||||||||
| T1+T2 | 50 | 36 | 14 | 0.019a | 50 | 30 | 20 | < 0.001a | |
| T3+T4 | 38 | 18 | 20 | 38 | 21 | 17 | |||
| Node Metastasis | |||||||||
| Absent | 7 | 7 | 0 | 0.04b | 7 | 6 | 1 | 0.058c | |
| Present | 81 | 47 | 34 | 81 | 45 | 36 | |||
| Distant Metastasis | |||||||||
| Absent | 55 | 36 | 19 | < 0.001a | 55 | 34 | 21 | < 0.001a | |
| Present | 33 | 18 | 15 | 33 | 17 | 16 | |||
| Peritoneal Seeding | |||||||||
| Absent | 51 | 39 | 12 | 0.001a | 51 | 33 | 18 | < 0.001a | |
| Present | 37 | 15 | 22 | 37 | 18 | 19 | |||
| Ascites | |||||||||
| Absent | 81 | 52 | 29 | < 0.001c | 81 | 48 | 33 | < 0.001c | |
| Present | 7 | 2 | 5 | 7 | 3 | 4 | |||
| Ovarian Tissue | |||||||||
| Normal | 8 | 8 | 0 | 0.047d | 8 | 8 | 0 | 0.022d | |
| Cancer | 88 | 54 | 34 | 88 | 51 | 37 | |||
| Primary Tumor Origin | |||||||||
| Ovarian | 81 | 47 | 34 | 0.04b | 81 | 45 | 36 | 0.038b | |
| Others | 7 | 7 | 0 | 7 | 6 | 1 | |||
| Histological Type | |||||||||
| Serous | 49 | 24 | 25 | 0.037d | 49 | 24 | 25 | 0.198d | |
| Mucinous | 9 | 8 | 1 | 9 | 8 | 1 | |||
| Endometrioid | 14 | 6 | 8 | 14 | 6 | 8 | |||
| Clear cell carcinoma | 9 | 9 | 0 | 9 | 7 | 2 | |||
Number in the table represented the case number of patients.
a: Pearson Chi-Square test; b: Fisher's exact test; c: Continuity correction test; d: Mann-Whitney Test;
FSHR and Snail expression are positively associated with ALKBH5 expression. (A) IHC of ALKBH5 and Snail on tissue microarray of ovarian tumor. Scale bar = 250 μm. (B) Correlation of ALKBH5 to Snail expression in ovarian tumor. Cramér's V was used for the association analysis. V = 0.711, P < 0.001. (C) IHC of Snail, ALKBH5 and FSHR in tissue sections of normal and tumor ovarian tissues. Scale bar = 200 μm. (D) Association between Snail and FSHR mRNA expression in The Cancer Genome Atlas Program (TCGA, https://tcga-data.nci.nih.gov/tcga/tcgaHome2.jsp). Cramér's V was used for the association analysis. (E) Association between ALKBH5 and FSHR mRNA expression in TCGA database. Cramér's V was used for the association analysis. (F) Western blots of FSHR, ALKBH5 and Snail protein in normal ovarian tissues (N) and ovarian tumors (T). (G) m6A content of total RNA in normal ovarian tissue and ovarian tumor (n = 5), data was analyzed by two-tailed unpaired t test. *P < 0.05.
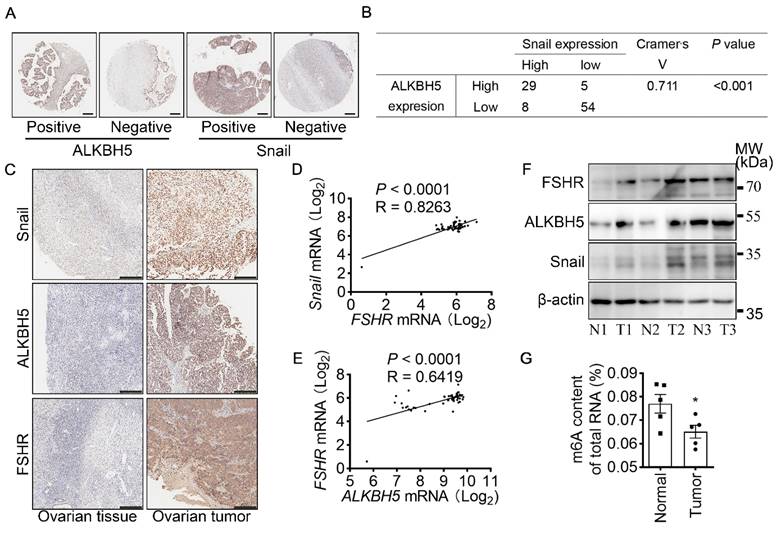
To confirm the role of FSHR, ALKBH5 and Snail in the progression of ovarian cancer, expressions of FSHR, ALKBH5 and Snail in paired freshly normal ovarian (from hysteromyoma patients) and tumor tissues were assessed. Upregulation of FSHR, ALKBH5, Snail was observed in ovarian tumor. IHC analysis also showed that decreased expression of E-cadherin, but increased expression of N-cadherin in ovarian tumor (Figure 6C and Figure S5D, E). TCGA data reveals the positive correlation between FSH, ALKBH5 and Snail expression in ovarian cancer (Figure 6D, E). Survival analysis showed both ALKBH5 and Snail higher expression was related to shorter overall survival times in ovarian cancer patients (Figure S5F, G). In human specimen, we also confirmed that FSHR, ALKBH5 and Snail protein expression showed significant upregulation in ovarian tumors, compared to normal ovarian tissue (Figure 6F), m6A content in total mRNA was higher than those in ovarian tumors (Figure 6G). Interestingly, we also found that METTL14 and YTHDF2 expression were decreased in ovarian tumor (Figure S5H). These results identified the role and strong correlation between FSHR, ALKBH5 and Snail in ovarian cancer progression.
Discussion
Although the "gonadotropin theory" of ovarian cancer pathogenesis remains a subject of controversy [14, 44], recent immunotherapies targeting FSHR have demonstrated high level of safety and efficacy in ovarian cancer treatment [11, 12]. Therefore, confirming the role of FSH in ovarian tumors and its underlying mechanisms would be essential for further optimizing the clinical application of FSHR immunotherapies. Our study revealed that FSH promoted metastasis in ovarian tumors through the EMT process, with ALKBH5-induced demethylation of m6A modifications in Snail mRNA playing a critical role in this procedure. This suggests that regulation on the reversible m6A modifications could be an effective approach to explore FSH-induced ovarian tumor metastasis.
Apart from its role in maintaining reproductive system development and regulating the function of extragonadal organs, FSH is associated with various tumor processes, including breast cancer [45], non-functional pituitary adenomas [46], and ovarian cancer [47]. With the discovery of FSHR in normal ovarian epithelial cells and ovarian cancer cells [4], the role of FSH in the ovarian tumor process induced by abnormal epithelial cell development has progressively attracted research interest. Our study found that, in tumor tissues from 160 patients with different stage of ovarian cancer, the expression of FSHR was found to be correlated with metastasis. Using the epithelial ovarian cancer cell line SK-OV3 and a nude mouse ovarian intraperitoneal metastasis model, we confirmed that FSH induced EMT and promoted ovarian tumor metastasis. Other than cell migration and invasion, FSH participated in ovarian tumor processes by regulating cell proliferation, apoptosis, and promoting tumor angiogenesis [48]. Our study, along with others, suggested that monitoring and intervening in hormone levels could constitute a safe and effective approach for prevention and treatment of hormone-related cancers.
Tumor metastasis is a major cause of death in ovarian cancer patients [16]. EMT is the initial step where cancer epithelial cells acquire the ability to move, leading to tumor metastasis [49]. Recent studies suggested that small molecule targeted therapy of EMT in tumor epithelial cells indicate a potential treatment for tumor metastasis [50]. In our study, FSH was shown to inhibit E-cadherin and increase N-cadherin, contributing to EMT and the metastasis of epithelial ovarian cancer cells in vivo and in vitro. Consistently, in tissues of nonfunctioning pituitary tumors, the concentrations of FSH were inversely associated with E-cadherin expression [51]. Moreover, research has reported that FSHR is present in metastases to lymph nodes, brain, bones, and liver, which were originated from breast, lung, prostate, and renal cancers [52]. Together, these results suggested that FSH influences tumor metastasis, and small molecule targeted therapy of FSH/FSHR holds promise as a safe and effective approach for treating cancer metastasis.
In tumor microenvironment, cancer and immune cells release cytokines, growth factors, and chemokines towards surrounding epithelial cells. This action activates EMT transcription factors (Snail, Slug, Twist, ZEB1, and ZEB2) and initiates EMT in the epithelial cells [53]. Snail is a classical transcription factor of EMT, and its activation is correlated with tumor pathological grading, lymph node metastasis, and poor prognoses of tumor metastasis [54]. Research has demonstrated that transcriptional activity of the Snail promoter is directly modulated by various growth factors and signaling molecules [55]. However, our results indicated that FSH has no impact on the transcriptional activation of Snail promoter. Furthermore, post-translational modifications, including ubiquitination [56], phosphorylation [57], and m6A modification [30], regulate the mRNA stability and protein expression of Snail, thereby impacting EMT. Our study confirmed that FSH increased Snail protein expression by stabilizing Snail mRNA via m6A modification. This result may explain the findings of Yang et al who observed FSH upregulating Snail protein in ovarian cancer [58]. Additionally, our discovery is corroborated by the observation that m6A modification in Snail mRNA was regulated by the methyltransferase METTL3 [30] in nasopharyngeal carcinoma and the demethylase FTO [59] in ovarian cancer. Importantly, our research identified the CDS region of Snail mRNA was the site of FSH-induced m6A modification. These results suggested that by leveraging the reversible nature of m6A regulation, regulation on m6A modification at specific sites on Snail could be a potential approach to treat tumor metastasis.
In tumor metastasis, m6A modification is linked to the tumor immune microenvironment [60], glucose metabolism of cancer cells [61], and migration and invasion capabilities [62]. Yu et al discovered m6A modifications in the EMT transcription factor Slug in head and neck squamous carcinoma [29]. Similarly, Lin et al identified m6A modifications in the EMT transcription factor ZEB1 in triple-negative breast cancer [31]. These observations aligned with our conclusion that the m6A modification in the EMT transcription factor Snail plays a pivotal role in FSH-induced EMT, contributing to ovarian tumor metastasis. Furthermore, we confirmed that the upregulation of demethylase ALKBH5 is a key factor leading to FSH-induced EMT, primarily by increasing Snail protein expression. However, studies on renal fibrosis suggested that the drug Genistein inhibited the occurrence of EMT by downregulating Snail through ALKBH5 [63]. These varied regulatory outcomes suggested that the influence of ALKBH5 on Snail may be contingent on distinct tissues or cell types, exhibiting diverse roles in various diseases [64].
ALKBH5 is implicated in the proliferation, migration, invasion of cancer cell, and tumor metastasis [65]. Our research for the first time demonstrated that ALKBH5 promotes EMT and facilitates intraperitoneal metastasis in ovarian cancer cells. This finding aligned with the discovery by Hu et al in clear cell renal cell carcinoma, showcasing the role of ALKBH5 in tumor metastasis through EMT [66], and consistent with the observation by Sun et al reporting the association of ALKBH5 with ovarian tumor-related lymphatics and lymphatic metastasis [23]. Moreover, we clarified the underlying mechanism by which ALKBH5 promotes EMT by increasing Snail mRNA stability through m6A modification. The stability of Snail mRNA is crucial in the context of tumor metastasis, as previously confirmed in lung cancer [67]. Therefore, intervening in Snail mRNA stability through pathways like ALKBH5 may represent a potential approach to decelerate tumors metastasis. Simultaneously, several studies indicate that ALKBH5, through regulating the stability of downstream gene mRNA, is involved in the progress of non-tumor diseases, particularly infertility resulting from abnormal reproductive system development [40]. These data indicated that ALKBH5 stabilizes the target mRNA, directly regulating tumor cells or influencing the tumor microenvironment, thereby playing an essential role in tumor metastasis.
Studies have indicated that the transcriptional process of ALKBH5 is jointly regulated by epigenetic factors and transcription factors [64]. Our experiments confirmed that the transcription factor CREB, as a classic downstream molecule of FSH, upregulates the level of ALKBH5 in ovarian tumors, ultimately leading to EMT occurrence. This finding was analogous to the results obtained by Guo et al in colorectal cancer cells, where activating the CREB signaling pathway significantly increased EMT in colorectal cancer cells [68]. Research has shown that FSH increased promoter activity of CREB in ovarian epithelial cells [69]. Moreover, CREB highly expressed in various tumors, including ovarian cancer [70], suggesting that CREB, as a responsive hormone activity intermediate regulator, plays a crucial role in tumors. The studies mentioned above suggested that targeting the CREB and downstream m6A-related signaling pathways in hormone-related tumor processes may represent a potential strategy for retarding tumor progression.
While this study provided a new perspective on FSH regulation of ovarian tumor metastasis via m6A modification, limitation in this study should be acknowledged. Although we have observed that FSH decreased METTL14 expression, the 'reader' of m6A, the role of METTL14 in EMT of ovarian cancer remains to be investigated. The role of Snail-regulated m6A modification in FSH-induced tumor metastasis has not been validated in animal experiments. Although m6A inhibitors for in vivo study, e.g., 3-deazaadenosine, are commercially available, the specificity of this inhibitor for m6A modification in Snail mRNA remained limited. Theoretically, development of mice with mutated methylation sites in Snail could address this limitation, however, that will be future work for our next project.
In conclusion, this study demonstrated that FSH activates CREB, leading to increased expression of ALKBH5. Subsequently, ALKBH5 induces m6A demethylation in the CDS region of Snail mRNA, enhancing mRNA stability and upregulating protein expression of Snail. This cascade of events induced EMT and ovarian cancer metastasis. Our findings suggested that disrupting m6A modification through intervention in FSH and its downstream signaling pathways could provide a novel therapeutic approach for ovarian tumor metastasis.
Conclusions
Overall, this study indicated that FSH increased ALKBH5 mRNA and protein expression via CREB activation. ALKBH5 triggers Snail mRNA m6A demethylation on its CDS region to increase Snail protein expression, thus induces EMT and ovarian cancer metastasis. It is anticipated that blocking FSH/FSHR and downstream signaling pathway via m6A alteration would provide a novel therapeutic avenue for ovarian tumor metastasis.
Materials and methods
Cell culture studies
Human epithelial ovarian cancer cell lines (SK-OV3, ES-2, A2780) and HEK293T cell line were purchased from the American Type Culture Collection (VA, USA). SK-OV3 and ES-2 cells were cultured in McCoy's 5A medium (#X054B1, BasalMedia) supplemented with 10% (v/v) heat-inactivated fetal bovine serum (FBS, #10270-106, Gibco), 100 unit/mL penicillin and 100 µg/mL streptomycin (P/S, #SV30010, Hyclone), while A2780 and HEK293T cells were grown in DMEM (#21041025, Gibco) supplemented with 10% (v/v) FBS and P/S. Opti-MEM (#31985) were acquired from Gibco (Waltham MA USA). Cells were cultured in a humidified environment of 95% air, 5% CO2 at 37°C. Experiments were performed when the cells reached 90% confluence.
Clinical specimens
5 pairs of epithelial ovarian cancer and normal ovarian tissue were obtained from the Guangzhou First People's Hospital, normal ovarian tissues samples were obtained from donors during surgery for other benign gynecological diseases. Diagnosis of normal and tumor tissues was performed by experienced pathologists. All patients were informed prior to the samples donation and had provided their written informed consent. The research in this work followed the principles of Helsinki Declaration and was approved by the Institutional Review Board of the Guangzhou First People's Hospital.
Tissue microarray
The tissue microarray (TMA) of ovarian tissues was obtained from Shanghai WellBio Technology. The TMA in Table 1 contained 160 ovarian tumor samples, TMA in Table 2 contained 88 ovarian tumor samples and 8 normal ovarian tissues samples, the patients in two TMA were recruited in Tongxu First Hospital, Kaifeng, Henan, China from 2015 to 2021. The TMA was sectioned at 5 µm thickness. Adjacent sections were used to assess the expression of ALKBH5 and Snail in the TMA using immunohistochemistry as described above. FSHR, ALKBH5 and Snail expression were independently evaluated by two experienced pathologists using double-blind approach. Tissue sections were divided into four groups: negative, weak, medium and strong grades. Negative and weak staining were classified as low expression, according to the protein expression, while medium and strong staining were classified as high expression.
Animals
Female Balb/c nude mice were obtained from Beijing Vital River Laboratory Animal Technology (Beijing, China). All mice were housed in a temperature-controlled room (Physical Containment Level 2, PC2) on a 12-hour light and dark cycle at Guangzhou Medical University Animal Center and were allowed access to water and food ad libitum. All animal experiments were carried out according to the Guide for the Care and Use of Laboratory Animals outlined by the National Institutes of Health (NIH) and conformed to the guidelines of Animal Ethics Committees of Guangzhou Medical University (Protocol: 2020-077).
Female Balb/c nude mice at 8 weeks of age were anesthetized using pentobarbital sodium (1%, 80 mg/kg), followed by bilateral ovariectomy (OVX), where two sides of ovaries were surgically excised with a single surgical incision performed by one surgeon as previously described [71]. After surgery, mice were allowed two weeks of recovery period before subjected to further experiments.
To established tumor xenotransplants, recovered OVX Balb/c nude mice were randomly divided into two groups. SK-OV3 cells stably expressing control shRNA (n = 8) or ALKBH5 shRNA (n = 8) were harvested and counted, respectively. SK-OV3 (2×106 cells per mouse) were resuspended in 100 μL PBS was then intraperitoneal injected into each mouse according to its respective group. One week after, PBS (vehicle control, n = 4 in each group) or recombinant FSH (3 IU per mouse, #F4021, Merck) dissolved in PBS (FSH, n = 4 in each group) was subcutaneously injected into mice every day for 14 weeks, according to previous study [71]. Throughout the study, animal body weight was monitored in a weekly base.
IVIS Lumina XRMS was used to track the progression of in vivo tumor peritoneum metastasis weekly one week after the intraperitoneal injection of SK-OV3 cells. OVX tumor bearing mice were anesthetized using isoflurane (4% for induction, 2% for maintenance). D-luciferin (15 mg/mL, 150 mg/kg, #122799, PerkinElmer) was intraperitoneally administrated into mice 10 minutes before IVIS imaging of luminescence (5 minutes exposure). The data were analyzed using Living Image software.
Cell transfection and lentiviral infection
The Snail siRNA was purchased from Santa Cruz Biotechnology (Santa Cruz, CA), and ALKBH5 siRNA (CCATGACGTCCCGGGACAACTATAA) was constructed and obtained from Invitrogen. siRNAs (40 nM diluted in 1 mL Opti-MEM) were transfected to SK-OV3 cells using Lipofectamine RNAiMAX (#13778150, Invitrogen). The plasmid of wild-type (pENTER-ALKBH5) and mutated ALKBH5-H204A (pENTER-ALKBH5-H204A) was obtained from WZ Biosciences Inc, while plasmid of wild-type (pSLenti-Snail) and mutated Snail (pSLenti-Snail-mut, adenosine bases in m6A consensus sites replaced by cytosine nucleotides) was constructed and obtained from OBiO Technology. Plasmid (2 µg diluted in 1 mL Opti-MEM) was transfected to SK-OV3 cells using Lipofectamine 3000 Transfection Reagent (#L3000015, Invitrogen). The shRNA of human ALKBH5 was cloned into a lentiviral vector (pLent-3in1-shRNA-CMV-copGFP-P2A-Puro) for knockdown. Sequences of the shRNAs were as followed, stably transfected SK-OV3 cells were selected using puromycin (#ST551, Beyotime, China): siRNA1: GTCCTTCTTTAGCGACTCT; siRNA2: CCATGACGTCCCGGGACAACTATAA; siRNA3: GCTCAGTGGATATGCTGCTGATGAA.
Immunofluorescence assay
SK-OV3 cells were cultured on glass coverslips (#801009, Nest) in six-well plates, fixed with 4% paraformaldehyde (#430012, Asegene), and permeabilized with 0.3% Triton X-100 (#DH351-1, Ding Guo Prosperous). The cells were then blocked in 5% BSA and incubated with primary antibodies against E-cadherin (1:1000, #60335-1-Ig, Proteintech), N-cadherin (1:1000, #22018-1-AP, Proteintech), ALKBH5 (1:1000, #16837-1-AP, Proteintech) and CREB (1:1000, #9104S, Cell Signaling Technology) at 4°C overnight, followed by FITC-conjugated secondary antibodies incubations (1:1000; Alexa Fluor® 555 Conjugate, #4409S, Cell Signaling Technology; 1:1000, Alexa Fluor™ Plus 488, #A32731, Invitrogen) at room temperature for 1 hour. Nuclei were counterstained with DAPI (1:5000, #4083S, Cell Signaling Technology). Finally, cells on glass coverslips were mounted on glass slides (#188105W, Citoglas) with antifade mounting reagent (#S2100, Asegene). The fluorescent signal was visualized using Zeiss microscope (LM980, Zeiss).
Western blotting and antibodies
After treatment, tissue or cells were washed with ice-cold PBS, lysed with lysis buffer (100 mM Tris-HCl (pH=6.8), 4% SDS, 20% glycerol, 1× protease inhibitors cocktail (Sigma, #P8340), 1× phosphatase inhibitors cocktail (Sigma, #P0044), 1 mM sodium orthovanadate, 1 mM NaF, and 1 mM phenylmethylsulfonylfluoride, PMSF). Lysate were scraped, harvested, denatured with boiling water, then centrifuged. The resulting supernatant were collected, and protein contents were determined using BCA assay (Thermo scientific, #23235). 25 µg protein was then added onto a 10% SDS-PAGE gel for separation by electrophoresis. Separated protein was then transferred to PVDF membrane, blocked with 5% BSA at room temperature for 60 min, and incubated with corresponding primary antibodies at 4°C overnight. Membrane was then washed with TBST and incubated with corresponding HRP-conjugated secondary antibodies at room temperature for 1 hour. Membrane was washed, while protein was detected and visualized using Enhanced Chemiluminescence reagent (#WBKLS0500, Millipore) and Imaging Systems (Amersham Imager 600, GE).
All antibodies used in this study were as follows: anti-FSHR (#22665-1-AP), anti-E-cadherin (#20874-1-AP), anti-N-cadherin (#22018-1-AP), anti-Snail (#13099-1-AP), anti-ZEB1 (#66279-1-Ig), anti-ZEB2 (#14026-1-AP), anti-ALKBH5 (#16837-1-AP), anti-Vimentin (#10366-1-AP), anti-GAPDH (#60004-1-Ig) were obtained from Proteintech. Anti-Slug (#9585S), anti-FTO (#45980S), anti-p-CREB (#9198S), anti-CREB (#9104S), anti-Twist (#90445), anti-Mouse IgG HRP-linked (#7076S), anti-Rabbit IgG HRP-linked (#7074S) were obtained from Cell Signaling Technology. Anti-β-actin (#sc-81178) was obtained from Santa Cruz. Anti-FTO (#ab124892), anti-METTL3 (#ab195352), anti-YTHDF1 (#ab99080), anti-YTHDF3 (#ab103328) were obtained from Abcam. Anti-YTHDF2 (#ABE542) was obtained from Millipore. Anti-METTL14 (#HPA038002) was obtained from Sigma. Before using, all primary antibodies were diluted in 1:2000 with 5% BSA, and the corresponding HRP-conjugated secondary antibodies were diluted in 1:10000 with 5% BSA.
RNA isolation and RT-qPCR
RNA extraction was performed using SteadyPure Universal RNA Extraction Kit (#AG21017, Accurate Biology) according to the manufacturer's instruction. Complementary DNA was prepared by reverse transcription using Evo M-MLV reverse transcriptase (#AG11706, Accurate Biology) following the user's manual. Real-time quantitative PCR was performed on thermal cycler (CFX96, Bio-Rad) using the SYBR Green probe (#AG11701, Accurate Biology). RNA transcripts were determined by relative to GAPDH [72] using the 2-△△Ct method as described in previous publication [73]. The detailed information of PCR primers was listed in the Supplementary Table 1.
Wound-healing assay
SK-OV3 cells were seeded onto a six-well plate with culture-insert (ibidi, #80496), which provide a defined cell-free gap for wound healing assays. 24 hours after seeding, culture-insert was removed. Cells were gently washed twice with PBS, cultured with FBS-free medium and ready for the subsequent treatments. 0 and 48 hours after treatment, three random fields of the wound healing area were imaged and used to determine cell migration.
Transwell invasion assay
SK-OV3 cell suspension (2×104 cells in FBS-free culture media) were added into the upper chamber of Matrigel-coated inserts (#356234, BD Bioscience). The Matrigel-coated inserts was then carefully transferred to the 24-well plate containing 10% FBS culture media for cell invasion. After pre-determined invasion period of 48 hours, inserts were carefully washed with PBS without disturbing the Matrigel layer, then fixed with 4% paraformaldehyde and stained with 0.1% crystal violet (#G1064, Solarbio). Non-invaded cells on the upper chamber were carefully removed. Insert were then placed on a slide for imaging. Five random fields were captured and used for cell counting.
Promoter activity and luciferase reporter assay
DNA fragment of ALKBH5 promoter (2000 bp) was cloned into pGL3 promoterless vector upstream of the luciferase gene to construct a recombinant plasmid of pGL3-Basic-ALKBH5-luc. The plasmids (800 ng diluted in 0.5 mL Opti-MEM) of pGL3-Basic-ALKBH5-luc and pRL-TK (80 ng diluted in 0.5 mL Opti-MEM) were then co-transfected into HEK293T cells using Lipofectamine 3000 Transfection Reagent (Invitrogen). After drug treatment for the transfected cells, promoter activity was assessed by measuring the luciferase activity using the Promega Reporter Assays (#E1910, Promega) according to the manufacturer's instruction (SPARK 10M, TECAN). Promoter activity of pGL3-Basic-ALKBH5-luc was then determined by relative to pRL-TK promoter activity using the comparative method.
Immunohistochemistry
Human ovarian tumor, health ovarian tissues, and SK-OV3 xenografts were collected, formalin-fixed and paraffin-embedded. Tissue paraffin blocks were then sectioned at 4 µm thickness (RM2016, Leica biosystems) and placed on a glass slide. Tissue sections were deparaffined using in xylene and rehydrated in serial dilution of ethanol from 100% to 70%. Rehydrated tissue sections were incubated with 0.3% H2O2 to quench the endogenous peroxidase, and then blocked with 3% BSA (#G5001, Servicebio). Primary antibodies of FSHR (1:200, #22665-1-AP, Proteintech), ALKBH5 (1:200, #ab195377, Abcam), Snail (1:200, #SC-271977, Santa Cruz), anti-E-cadherin (1:200, #20874-1-AP, Proteintech), anti-N-cadherin (1:200, #22018-1-AP, Proteintech), Ki-67 (1:200, #12202S, Cell Signaling Technology) and isotype antibodies (1:200, #31235, Cell Signaling Technology) were used to treat the tissue section at 4 °C overnight, followed by the correspond HRP-conjugated secondary antibodies. After washing, protein signal was visualized using DAB (#GK500710, Gene Tech). Stained sections were imaged using a bright field microscope (Leica CS2, Leica biosystems). Image J software was used to quantify the average optical density (AOD).
Analysis of m6A content
Total RNA was extract using SteadyPure Universal RNA Extraction Kit (#AG21017, Accurate Biology). Total m6A content was measured using the m6A RNA Methylation Assay Kit (#ab185912, Abcam). Briefly, 80 ul of binding solution and 200 ng of sample RNA were add into pre-coated plate and incubate at 37°C for 90 min for RNA binding. The dissociated RNA was discarded using washing buffer. After incubation with capture antibody, detection antibody and enhancer solution in sequence, m6A modification RNA was captured. Finally, the wells were incubated with developer solution in the dark, then stop solution was used to stop the reaction, m6A content was quantified using a microplate reader at 450 nm wavelength.
Analysis of Snail mRNA methylation level
Total RNA was extracted from cells using Trizol (#15596081, Ambion) according to the manufacturer's instruction. mRNA was separated from total RNA according to the protocol of the GenElute™ mRNA Low Dose Preparation Kit (MRN70, Sigma). Then the above mRNA was concentrated with glycogen (AM9510, Invitrogen), and the mRNA fragment was processed. RNA was incubated with m6A antibody for immunoprecipitation using Magna methylated RNA immunoprecipitation m6A Kit (MAGNAPIP, Millipore), qRT-PCR was performed using 2 µL m6A IP RNA as described above. The detailed information of the PCR primers is listed in the Supplementary Table 1.
Statistical analysis
The individual data was expressed as mean ± standard error of the mean (SEM) from at least three independent experiments unless otherwise specified. Unpaired Student's t-test and Mann-Whitney tests were used in the comparison of two groups, while one-way ANOVA was used for multiple comparison. The Pearson's χ2, Pearson's corrected χ2, and Fisher's exact tests were used to analyze the association between ALKBH5/Snail expression, FSHR expression and clinicopathological characteristics. Statistical analyses were performed with SPSS 17.0. A P-value of < 0.05 was considered to be statistically significant. *P < 0.05, **P < 0.01, ***P < 0.001, ns. not significant.
Abbreviations
FSH, follicle-stimulating hormone; EMT, epithelial-mesenchymal transition; m6A, N6-methyladenosine; ALKBH5, AlkB Homolog 5, RNA Demethylase; qRT-PCR, quantitative real time PCR; m6A-RT-PCR, methylated RIP-qPCR; FSHR, follicle stimulating hormone receptor; ActD, actinomycin D; CHX, cycloheximide; OVX, ovariectomized; CCK8, cell counting kit-8; TMA, tissue microarray; IHC, immunohistochemistry; CREB: cAMP response element-binding protein.
Supplementary Material
Supplementary figures and table.
Acknowledgements
Funding
This work was supported by National Natural Science Foundation (No. 81871137, 82171631 to X.D. Fu), Guangdong Natural Science Foundation (No. 2021A1515010074 to Y Chen). Guangzhou Science and Technology Project (No. 202102020060 to J. Teoh).
Author contributions
XD. F. conceived and supervised the study. WY. C. and XY. X. wrote the original draft of the manuscript. XD. F., WY. C., XY. X., HW. Y., Y. C. and XF. Z. revised the manuscript. XY. X., XF. Z., P. L., XS. L., HP. L., J. T., YW. C. and YL. Y. performed the experiments and analyzed the data.
Ethics approval and consent to participate
The clinical specimens' collection in this study was approved by the Institutional Review Board of the Guangzhou First People's Hospital (Ref. no. K-2021-165-01). Animal experiments was approved by experiments Animal Ethics Committees of Guangzhou Medical University (Protocol number: 2020-077). Two tissue microarray (TMA) sample collection in this study was approved by the Institutional Review Board of Tongxu First Hospital.
Consent for publication
All authors read and approved the final manuscript for publication.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2022. CA Cancer J Clin. 2022;72:7-33
2. Veneziani AC, Gonzalez-Ochoa E, Alqaisi H, Madariaga A, Bhat G, Rouzbahman M. et al. Heterogeneity and treatment landscape of ovarian carcinoma. Nat Rev Clin Oncol. 2023;20:820-42
3. Konishi I, Kuroda H, Mandai M. Review: Gonadotropins and development of ovarian cancer. Oncology. 1999;57(Suppl 2):45-8
4. Choi KC, Kang SK, Tai CJ, Auersperg N, Leung PC. Follicle-stimulating hormone activates mitogen-activated protein kinase in preneoplastic and neoplastic ovarian surface epithelial cells. J Clin Endocrinol Metab. 2002;87:2245-53
5. Choi JH, Choi KC, Auersperg N, Leung PC. Gonadotropins upregulate the epidermal growth factor receptor through activation of mitogen-activated protein kinases and phosphatidyl-inositol-3-kinase in human ovarian surface epithelial cells. Endocr Relat Cancer. 2005;12:407-21
6. Halperin R, Hadas E, Langer R, Bukovsky I, Schneider D. Peritoneal fluid gonadotropins and ovarian hormones in patients with ovarian cancer. Int J Gynecol Cancer. 1999;9:502-7
7. Li Y, Ganta S, Cheng C, Craig R, Ganta RR, Freeman LC. Fsh stimulates ovarian cancer cell growth by action on growth factor variant receptor. Mol Cell Endocrinol. 2007;267:26-37
8. Liu L, Zhang J, Fang C, Zhang Z, Feng Y, Xi X. Oct4 mediates fsh-induced epithelial-mesenchymal transition and invasion through the erk1/2 signaling pathway in epithelial ovarian cancer. Biochem Biophys Res Commun. 2015;461:525-32
9. Zheng W, Magid MS, Kramer EE, Chen YT. Follicle-stimulating hormone receptor is expressed in human ovarian surface epithelium and fallopian tube. Am J Pathol. 1996;148:47-53
10. Blaakaer J, Baeksted M, Micic S, Albrectsen P, Rygaard J, Bock J. Gonadotropin-releasing hormone agonist suppression of ovarian tumorigenesis in mice of the wx/wv genotype. Biol Reprod. 1995;53:775-9
11. Bordoloi D, Bhojnagarwala PS, Perales-Puchalt A, Kulkarni AJ, Zhu X, Liaw K. et al. A mab against surface-expressed fshr engineered to engage adaptive immunity for ovarian cancer immunotherapy. JCI Insight. 2022 7
12. Perales-Puchalt A, Svoronos N, Rutkowski MR, Allegrezza MJ, Tesone AJ, Payne KK. et al. Follicle-stimulating hormone receptor is expressed by most ovarian cancer subtypes and is a safe and effective immunotherapeutic target. Clin Cancer Res. 2017;23:441-53
13. Perales-Puchalt A, Wojtak K, Duperret EK, Yang X, Slager AM, Yan J. et al. Engineered dna vaccination against follicle-stimulating hormone receptor delays ovarian cancer progression in animal models. Mol Ther. 2019;27:314-25
14. Beral V, Bull D, Green J, Reeves G. Ovarian cancer and hormone replacement therapy in the million women study. Lancet. 2007;369:1703-10
15. Ivarsson K, Sundfeldt K, Brännström M, Hellberg P, Janson PO. Diverse effects of fsh and lh on proliferation of human ovarian surface epithelial cells. Hum Reprod. 2001;16:18-23
16. Matulonis UA, Sood AK, Fallowfield L, Howitt BE, Sehouli J, Karlan BY. Ovarian cancer. Nat Rev Dis Primers. 2016;2:16061
17. Stemmler MP, Eccles RL, Brabletz S, Brabletz T. Non-redundant functions of emt transcription factors. Nat Cell Biol. 2019;21:102-12
18. Yue B, Song C, Yang L, Cui R, Cheng X, Zhang Z. et al. Mettl3-mediated n6-methyladenosine modification is critical for epithelial-mesenchymal transition and metastasis of gastric cancer. Mol Cancer. 2019;18:142
19. Jeschke J, Collignon E, Al Wardi C, Krayem M, Bizet M, Jia Y. et al. Downregulation of the fto m(6)a rna demethylase promotes emt-mediated progression of epithelial tumors and sensitivity to wnt inhibitors. Nat Cancer. 2021;2:611-28
20. Zhao X, Li X, Li X. Multiple roles of m(6)a methylation in epithelial-mesenchymal transition. Mol Biol Rep. 2022;49:8895-906
21. Deng LJ, Deng WQ, Fan SR, Chen MF, Qi M, Lyu WY. et al. M6a modification: recent advances, anticancer targeted drug discovery and beyond. Mol Cancer. 2022;21:52
22. Wang Y, Wang Y, Patel H, Chen J, Wang J, Chen ZS. et al. Epigenetic modification of m(6)a regulator proteins in cancer. Mol Cancer. 2023;22:102
23. Sun R, Yuan L, Jiang Y, Wan Y, Ma X, Yang J. et al. Alkbh5 activates fak signaling through m6a demethylation in itgb1 mrna and enhances tumor-associated lymphangiogenesis and lymph node metastasis in ovarian cancer. Theranostics. 2023;13:833-48
24. Liu T, Wei Q, Jin J, Luo Q, Liu Y, Yang Y. et al. The m6a reader ythdf1 promotes ovarian cancer progression via augmenting eif3c translation. Nucleic Acids Res. 2020;48:3816-31
25. Zhang Y, Qiu JG, Jia XY, Ke Y, Zhang MK, Stieg D. et al. Mettl3-mediated n6-methyladenosine modification and hdac5/yy1 promote iffo1 downregulation in tumor development and chemo-resistance. Cancer Lett. 2023;553:215971
26. Shi Y, Xiong X, Sun Y, Geng Z, Chen X, Cui X. et al. Igf2bp2 promotes ovarian cancer growth and metastasis by upregulating ckap2l protein expression in an m(6) a-dependent manner. FASEB J. 2023;37:e23183
27. Nie S, Zhang L, Liu J, Wan Y, Jiang Y, Yang J. et al. Alkbh5-hoxa10 loop-mediated jak2 m6a demethylation and cisplatin resistance in epithelial ovarian cancer. J Exp Clin Cancer Res. 2021;40:284
28. Wang R, Ye H, Yang B, Ao M, Yu X, Wu Y. et al. M6a-modified circnfix promotes ovarian cancer progression and immune escape via activating il-6r/jak1/stat3 signaling by sponging mir-647. Int Immunopharmacol. 2023;124:110879
29. Yu D, Pan M, Li Y, Lu T, Wang Z, Liu C. et al. Rna n6-methyladenosine reader igf2bp2 promotes lymphatic metastasis and epithelial-mesenchymal transition of head and neck squamous carcinoma cells via stabilizing slug mrna in an m6a-dependent manner. J Exp Clin Cancer Res. 2022;41:6
30. Yu X, Zhao H, Cao Z. The m6a methyltransferase mettl3 aggravates the progression of nasopharyngeal carcinoma through inducing emt by m6a-modified snail mrna. Minerva Med. 2022;113:309-14
31. Lin Y, Jin X, Nie Q, Chen M, Guo W, Chen L. et al. Ythdf3 facilitates triple-negative breast cancer progression and metastasis by stabilizing zeb1 mrna in an m(6)a-dependent manner. Ann Transl Med. 2022;10:83
32. Xu J, Lamouille S, Derynck R. Tgf-beta-induced epithelial to mesenchymal transition. Cell Res. 2009;19:156-72
33. Deng X, Qing Y, Horne D, Huang H, Chen J. The roles and implications of rna m(6)a modification in cancer. Nat Rev Clin Oncol. 2023;20:507-26
34. Zhang Y, Wang X, Duan X, Du T, Chen X. The synergistic effect of emt regulators and m6a modification on prognosis-related immunological signatures for ovarian cancer. Sci Rep. 2023;13:14872
35. Cao L, Wan Q, Li F, Tang CE. Mir-363 inhibits cisplatin chemoresistance of epithelial ovarian cancer by regulating snail-induced epithelial-mesenchymal transition. BMB Rep. 2018;51:456-61
36. Hojo N, Huisken AL, Wang H, Chirshev E, Kim NS, Nguyen SM. et al. Snail knockdown reverses stemness and inhibits tumour growth in ovarian cancer. Sci Rep. 2018;8:8704
37. Uzonyi A, Dierks D, Nir R, Kwon OS, Toth U, Barbosa I. et al. Exclusion of m6a from splice-site proximal regions by the exon junction complex dictates m6a topologies and mrna stability. Mol Cell. 2023;83:237-51.e7
38. Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D. et al. N6-methyladenosine-dependent regulation of messenger rna stability. Nature. 2014;505:117-20
39. Gong C, Wu J, Li H, Luo C, Ji G, Guan X. et al. Mettl3 achieves lipopolysaccharide-induced myocardial injury via m(6)a-dependent stabilization of myh3 mrna. Biochim Biophys Acta Mol Cell Res. 2023;1870:119503
40. Zheng G, Dahl JA, Niu Y, Fedorcsak P, Huang CM, Li CJ. et al. Alkbh5 is a mammalian rna demethylase that impacts rna metabolism and mouse fertility. Mol Cell. 2013;49:18-29
41. Feng C, Liu Y, Wang G, Deng Z, Zhang Q, Wu W. et al. Crystal structures of the human rna demethylase alkbh5 reveal basis for substrate recognition. J Biol Chem. 2014;289:11571-83
42. Lee HJ, Mignacca RC, Sakamoto KM. Transcriptional activation of egr-1 by granulocyte-macrophage colony-stimulating factor but not interleukin 3 requires phosphorylation of camp response element-binding protein (creb) on serine 133. J Biol Chem. 1995;270:15979-83
43. Xie F, Li BX, Kassenbrock A, Xue C, Wang X, Qian DZ. et al. Identification of a potent inhibitor of creb-mediated gene transcription with efficacious in vivo anticancer activity. J Med Chem. 2015;58:5075-87
44. Stadel BV. Letter: the etiology and prevention of ovarian cancer. Am J Obstet Gynecol. 1975;123:772-4
45. Eskelinen M, Nordén T, Lindgren A, Wide L, Adami HO, Holmberg L. Preoperative serum levels of follicle stimulating hormone (fsh) and prognosis in invasive breast cancer. Eur J Surg Oncol. 2004;30:495-500
46. Wang Y, Cheng T, Lu M, Mu Y, Li B, Li X. et al. Tmt-based quantitative proteomics revealed follicle-stimulating hormone (fsh)-related molecular characterizations for potentially prognostic assessment and personalized treatment of fsh-positive non-functional pituitary adenomas. EPMA J. 2019;10:395-414
47. Wei S, Lai L, Yang J, Zhuandi G. Expression levels of follicle-stimulating hormone receptor and implication in diagnostic and therapeutic strategy of ovarian cancer. Oncol Res Treat. 2018;41:651-4
48. Choi JH, Wong AS, Huang HF, Leung PC. Gonadotropins and ovarian cancer. Endocr Rev. 2007;28:440-61
49. Bakir B, Chiarella AM, Pitarresi JR, Rustgi AK. Emt, met, plasticity, and tumor metastasis. Trends Cell Biol. 2020;30:764-76
50. Lengrand J, Pastushenko I, Vanuytven S, Song Y, Venet D, Sarate RM. et al. Pharmacological targeting of netrin-1 inhibits emt in cancer. Nature. 2023;620:402-8
51. Kolnes AJ, Øystese KAB, Olarescu NC, Ringstad G, Berg-Johnsen J, Casar-Borota O. et al. FSH levels are related to e-cadherin expression and subcellular location in nonfunctioning pituitary tumors. J Clin Endocrinol Metab. 2020;105:2587-94
52. Siraj A, Desestret V, Antoine M, Fromont G, Huerre M, Sanson M. et al. Expression of follicle-stimulating hormone receptor by the vascular endothelium in tumor metastases. BMC Cancer. 2013;13:246
53. Dongre A, Weinberg RA. New insights into the mechanisms of epithelial-mesenchymal transition and implications for cancer. Nat Rev Mol Cell Biol. 2019;20:69-84
54. Wang Y, Shi J, Chai K, Ying X, Zhou BP. The role of snail in emt and tumorigenesis. Curr Cancer Drug Targets. 2013;13:963-72
55. Barberà MJ, Puig I, Domínguez D, Julien-Grille S, Guaita-Esteruelas S, Peiró S. et al. Regulation of snail transcription during epithelial to mesenchymal transition of tumor cells. Oncogene. 2004;23:7345-54
56. Viñas-Castells R, Beltran M, Valls G, Gómez I, García JM, Montserrat-Sentís B. et al. The hypoxia-controlled fbxl14 ubiquitin ligase targets snail1 for proteasome degradation. J Biol Chem. 2010;285:3794-805
57. Xie W, Jiang Q, Wu X, Wang L, Gao B, Sun Z. et al. Ikbke phosphorylates and stabilizes snail to promote breast cancer invasion and metastasis. Cell Death Differ. 2022;29:1528-40
58. Yang Y, Zhang J, Zhu Y, Zhang Z, Sun H, Feng Y. Follicle-stimulating hormone induced epithelial-mesenchymal transition of epithelial ovarian cancer cells through follicle-stimulating hormone receptor pi3k/akt-snail signaling pathway. Int J Gynecol Cancer. 2014;24:1564-74
59. Sun M, Zhang X, Bi F, Wang D, Zhou X, Li X. et al. Fto inhibits epithelial ovarian cancer progression by destabilising snai1 mrna through igf2bp2. Cancers (Basel). 2022 14
60. Yin H, Zhang X, Yang P, Zhang X, Peng Y, Li D. et al. Rna m6a methylation orchestrates cancer growth and metastasis via macrophage reprogramming. Nat Commun. 2021;12:1394
61. Hou Y, Zhang Q, Pang W, Hou L, Liang Y, Han X. et al. Ythdc1-mediated augmentation of mir-30d in repressing pancreatic tumorigenesis via attenuation of runx1-induced transcriptional activation of warburg effect. Cell Death Differ. 2021;28:3105-24
62. Yang X, Zhang S, He C, Xue P, Zhang L, He Z. et al. Mettl14 suppresses proliferation and metastasis of colorectal cancer by down-regulating oncogenic long non-coding rna xist. Mol Cancer. 2020;19:46
63. Ning Y, Chen J, Shi Y, Song N, Yu X, Fang Y. et al. Genistein ameliorates renal fibrosis through regulation snail via m6a rna demethylase alkbh5. Front Pharmacol. 2020;11:579265
64. Qu J, Yan H, Hou Y, Cao W, Liu Y, Zhang E. et al. Rna demethylase alkbh5 in cancer: from mechanisms to therapeutic potential. J Hematol Oncol. 2022;15:8
65. Wang J, Wang J, Gu Q, Ma Y, Yang Y, Zhu J. et al. The biological function of m6a demethylase alkbh5 and its role in human disease. Cancer Cell Int. 2020;20:347
66. Hu W, Klümper N, Schmidt D, Ritter M, Ellinger J, Hauser S. Depletion of the m(6)a demethylases fto and alkbh5 impairs growth and metastatic capacity through emt phenotype change in clear cell renal cell carcinoma. Am J Transl Res. 2023;15:1744-55
67. Wang X, Liu R, Zhu W, Chu H, Yu H, Wei P. et al. Udp-glucose accelerates snai1 mrna decay and impairs lung cancer metastasis. Nature. 2019;571:127-31
68. Guo YH, Wang LQ, Li B, Xu H, Yang JH, Zheng LS. et al. Wnt/β-catenin pathway transactivates microrna-150 that promotes emt of colorectal cancer cells by suppressing creb signaling. Oncotarget. 2016;7:42513-26
69. Gubbay O, Rae MT, McNeilly AS, Donadeu FX, Zeleznik AJ, Hillier SG. Camp response element-binding (creb) signalling and ovarian surface epithelial cell survival. J Endocrinol. 2006;191:275-85
70. Steven A, Friedrich M, Jank P, Heimer N, Budczies J, Denkert C. et al. What turns creb on? and off? and why does it matter? Cell Mol Life Sci. 2020;77:4049-67
71. Li X, Chen W, Li P, Wei J, Cheng Y, Liu P. et al. Follicular stimulating hormone accelerates atherogenesis by increasing endothelial vcam-1 expression. Theranostics. 2017;7:4671-88
72. Vermani L, Kumar R, Senthil Kumar N. Gapdh and pum1: optimal housekeeping genes for quantitative polymerase chain reaction-based analysis of cancer stem cells and epithelial-mesenchymal transition gene expression in rectal tumors. Cureus. 2020;12:e12020
73. Xu X, Yan Q, Liu X, Li P, Li X, Chen Y. et al. 17β-estradiol nongenomically induces vascular endothelial h(2)s release by promoting phosphorylation of cystathionine γ-lyase. J Biol Chem. 2019;294:15577-92
Author contact
Corresponding authors: MD Yang Cheng, Guangzhou First People's Hospital, 1st Panfu Rd, Yuxiu District, Guangzhou, China, 510180. Email: 391031160com; Dr Weiyu Chen, Guangzhou Medical University, 1st Xinzao Rd, Xinzao, Panyu District Guangzhou, China, 510260. Email: wychenedu.cn; Dr Xiaodong Fu, Guangzhou Medical University, 1st Xinzao Rd, Xinzao, Panyu District Guangzhou, China, 510260. Email: fuxiaodedu.cn (send all correspondence to this email).
 Global reach, higher impact
Global reach, higher impact