13.3
Impact Factor
Theranostics 2015; 5(11):1214-1224. doi:10.7150/thno.12403 This issue Cite
Research Paper
Identification of 7 Proteins in Sera of RA Patients with Potential to Predict ETA/MTX Treatment Response
1. INSERM, U905, Rouen, France
2. CNRS, UMR6270, Mont Saint Aignan, France
3. PISSARO Proteomic Platform, Mont Saint Aignan, France
4. Normandy University, IRIB, Rouen, France
5. Department of Rheumatology, Rouen University Hospital, Rouen, France
6. CIC/CRB1404, Rouen University, Rouen, France
7. Department of Immunology, Rouen University Hospital, Rouen, France
8. Department of Rheumatology, Amiens University Hospital, Amiens, France
9. Department of Rheumatology, Lille University Hospital, Lille, France
10. Department of Rheumatology, Caen University Hospital, Caen, France
Abstract
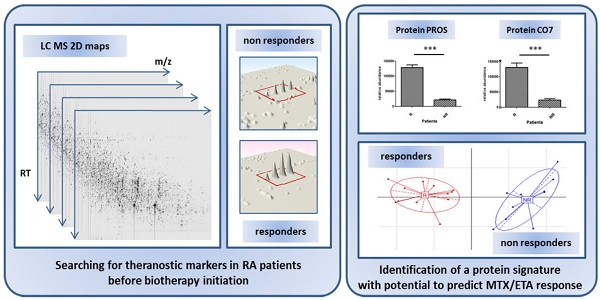
Objective: The recent growth of innovating biologics has opened fascinating avenues for the management of patients. In rheumatoid arthritis, many biologics are currently available, the choice of which being mostly determined empirically. Importantly, a given biologic may not be active in a fraction of patients and may even provoke side effects. Here, we conducted a comparative proteomics study in attempt to identify a predictive theranostic signature of non-response in patients with rheumatoid arthritis treated by etanercept/methotrexate combination.
Methods: A serum sample was collected prior to treatment exposure from a cohort of 22 patients with active RA. A proteomic “label free” approach was then designed to quantitate protein biomarkers using mass spectrometry. To verify these results, a relative quantification followed by an absolute quantification of interesting protein candidates were performed on a second cohort. The criterion of judgment was the response to etanercept/methotrexate combination according to the EULAR criteria assessed at 6 months of treatment.
Results: These investigations led to the identification of a set of 12 biomarkers with capacity to predict treatment response. A targeted quantitative analysis allowed to confirm the potential of 7 proteins from the latter combination on a new cohort of 16 patients. Two highly discriminating proteins, PROS and CO7, were further evaluated by ELISA on this second cohort. By combining the concentration threshold of each protein associated to a right classification (responders vs non-responders), the sensitivity and specificity reached 88.9 % and 100 %, respectively.
Conclusion: Prior to methotrexate/etanercept treatment, abundance of several sera proteins, notably PROS and CO7, were associated to response status of RA patients 6 month after treatment initiation.
Keywords: biomarkers, rheumatoid arthritis
 Global reach, higher impact
Global reach, higher impact