13.3
Impact Factor
Theranostics 2017; 7(1):213-227. doi:10.7150/thno.16044 This issue Cite
Research Paper
Genome-Wide lncRNA Microarray Profiling Identifies Novel Circulating lncRNAs for Detection of Gastric Cancer
1. Department of General Surgery & Institute of General Surgery, Chinese People's Liberation Army General Hospital, Fuxing Road 28, Beijing 100853, P.R. China;
2. Department of General Surgery, General Hospital of Armed Police Force, Yongding Road 69, Beijing 100039, P.R. China.
*Contributed equally to this work.
Abstract
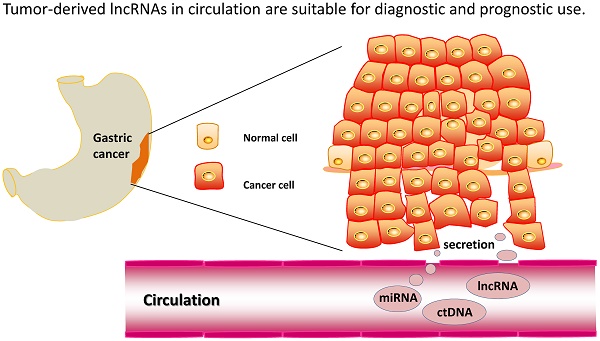
Long non-coding RNAs (lncRNAs) can serve as blood-based biomarkers for cancer detection. To identify novel lncRNA biomarkers for gastric cancer (GC), we conducted, for the first time, genome-wide lncRNA screening analysis in two sets of samples: five paired preoperative and postoperative day 14 plasma samples from GC patients, and tissue samples from tumor and adjacent normal tissues. Candidate tumor-related lncRNAs were then quantitated and evaluated in three independent phases comprising 321 participants. The expression levels of lncRNAs were also measured in GC cell lines and the corresponding culture medium. Biomarker panels, lncRNA-based Index I and carcinoembryonic antigen (CEA)-based Index II, were constructed using logistic regression, and their diagnostic performance compared. Fagan's nomogram was plotted to facilitate clinical application. As a result, we identified five novel plasma lncRNAs (TINCR, CCAT2, AOC4P, BANCR and LINC00857), which, when combined in the lncRNA-based Index I, outperformed the CEA-based Index II (P < 0.001) and could distinguish GC patients from healthy controls with an area under the receiver-operating curve (AUC) of 0.91 (95% confidence interval (CI): 0.88-0.95). The lncRNA-based index decreased significantly by postoperative day 14 (P = 0.016), indicating its ability to monitor tumor dynamics. High values of the lncRNA-based index were correlated with tumor size (P = 0.036), depth of invasion (P = 0.025), lymphatic metastasis (P = 0.012) and more advanced tumor stages (P = 0.003). The lncRNA-based index was also able to discriminate GC patients from precancerous individuals and patients with gastrointestinal stromal tumor with AUC values of 0.82 (95% CI: 0.71-0.92) and 0.80 (95% CI: 0.68-0.91), respectively. Taken together, our findings demonstrate that this panel of five plasma lncRNAs could serve as a set of novel diagnostic biomarkers for GC detection.
Keywords: circulating lncRNAs, gastric cancer, diagnosis, microarray, nomogram.
 Global reach, higher impact
Global reach, higher impact