Impact Factor
Theranostics 2019; 9(9):2637-2645. doi:10.7150/thno.32224 This issue Cite
Research Paper
Synergistically enhanced colorimetric molecular detection using smart cup: a case for instrument-free HPV-associated cancer screening
1. Department of Mechanical Engineering and Applied Mechanics, University of Pennsylvania, 220 South 33rd St. Philadelphia, Pennsylvania 19104-6315, USA
2. Department of Pathology and Laboratory Medicine, University of Pennsylvania, 3400 Spruce St. Philadelphia, PA 19104-4283, USA
3. Department of Microbiology, University of Pennsylvania, 3610 Hamilton Walk, Philadelphia, PA 19104-6076, USA
4. Present address: Department of Biomedical Engineering, University of Connecticut Health Center, 263 Farmington Avenue, Farmington, CT 06030, USA
Received 2018-12-12; Accepted 2019-2-7; Published 2019-4-13
Abstract
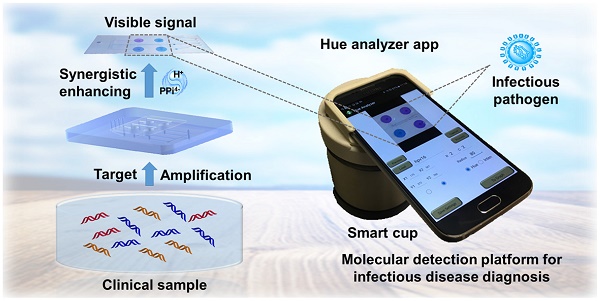
Rationale: Early and accurate detection of disease is crucial for its prevention, identification, and treatment. However, most of disease diagnostics is still limited in clinical laboratories due to the need of complicated instruments and professional personnel. Herein, we reported a smartphone-based synergistically enhanced colorimetric method for molecular diagnostics in our point of care (POC) smart cup platform.
Methods: A disposable microfluidic chip was developed for colorimetric loop-mediated isothermal amplification (LAMP) detection of multiple HPV DNA in our POC smart cup platform. The colorimetric detection takes advantage of synergistic effect of PPi4- and H+ ions, two byproducts of LAMP reaction. Color signal of LAMP assay was recorded and analyzed by our custom Android app (dubbed “Hue Analyzer”).
Results: Our method not only significantly improves colorimetric readout, but also provides a 10-fold increase in detection sensitivity. It has been successfully applied for HPV-associated cancer screening with spiked saliva and clinical swab samples.
Conclusion: The proposed POC diagnostic platform is completely compatible with other nucleic acid biomarkers and has great potential for personalized health monitoring and disease prevention.
Keywords: synergistic enhancing effect, hue based colorimetric detection, point-of-care testing, HPV-associated cancer screening
Introduction
Early and rapid disease diagnosis, including cancer screening, infectious disease detection, remains a significant global challenge. Nowadays, cancer and infectious disease have become the leading causes of death and disability, posing a considerable threat to human health globally [1]. For example, human papillomavirus (HPV) causes nearly all cervical cancers (99%) and some other cancers (i.e., oropharynx, vagina, vulva) [2]. Recent research demonstrated that primary HPV DNA testing detects cervical neoplasia earlier and more accurately than cytology test widely used for cervical cancer screening [3]. However, current HPV DNA tests, such as QIAGEN's careHPV™, Cepheid Xpert HPV Assay (Xpert), Atila HPV diagnostics kit and Roche's cobas® HPV test, typically require well-trained technicians, expensive instruments, and long detection time (several hours to days), which are not suitable for widespread application at home and low-resource settings [4]. Therefore, there is an unmet need for simple, fast, accurate, point of care (POC) molecular detection technologies.
Recently, simple isothermal amplification techniques such as loop-mediated isothermal amplification (LAMP) has emerged as a powerful method for POC molecular diagnostics due to its simplicity, sensitivity and low resource requirement [5, 6]. Especially, simple colorimetric detection of LAMP reaction can be achieved with various indicators including Mg2+ indicator, pH-sensitive dyes, and functional gold nanoparticles [7-13]. However, most of them only monitor single ion changes (e.g., H+ or Mg2+) during LAMP amplification, leading to low reliability and non-sensitive colorimetric LAMP detection.
Today smartphones have become ubiquitous with internet connectivity, including developing countries with resource-limited settings. The embedded camera coupled with powerful computing capabilities enables smartphone to become an alternative to conventional dedicated optical detector, allowing low cost and reliable signal acquisition, image analysis, and result reporting. Especially, image analysis using smartphone can be easily adapted for objective color readout, eliminating variations caused by potential differences in color perception and environmental factors during analysis [14-17]. For example, smartphone-based approach has been adopted for the fluorescence-based and colorimetric-based detection of LAMP reaction [12, 18-23]. An inexpensive fiber-optic bundle-based fluorescence microplate reader integrated with a mobile phone has been developed for highly stable and sensitive detection of nucleic acid amplification [18]. CIE xyY colorspace (chromaticity-luminance) method has been developed to analyze LAMP fluorescence signals [19]. However, fluorescence-based LAMP assay typically requires fluorescence filters and excitation light source. Colorimetric-based LAMP assay is simpler, lower cost and more suitable for instrument-free POC diagnostic applications.
Herein, we developed a smartphone-based synergistically enhanced colorimetric molecular detection strategy for LAMP assay, enabling instrument-free molecular diagnostics at the point of care. The target nucleic acids of causative pathogens from raw samples can be directly amplified through LAMP reaction in a disposable microfluidic chip in combination with our portable smart cup platform powered by exothermic chemical reaction [23, 24]. The synergistically enhanced visible color signal can be recorded by a smartphone camera and readout by our custom Android app (dubbed “Hue Analyzer”). As a demonstration of clinical applications, we successfully achieved HPV-associated cancer screening by detecting multiple HPV DNAs (i.e., HPV 16, 18 and 31) in saliva and clinical cervical swab samples in the smart cup platform.
Experimental
Chemicals and instruments
(NH4)2SO4, Tween 20, KCl, KOH, acetonitrile and hydroxy naphthol blue (HNB) were purchased from Sigma-Aldrich. The stock solution of HNB was 2 mM in ddH2O. Sterile molecular biology-grade water was purchased from Thermo Fisher Scientific Inc. EvaGreen® dye was purchased from Biotium. 10X isothermal amplification reaction buffer, Bst 2.0 DNA Polymerase (8,000 U/mL), thermostable inorganic pyrophosphatase (2,000 U/mL), MgSO4 and dNTP were purchased from New England BioLabs (NEB) Inc. GspSSD2.0 Isothermal Mastermix was purchased from OptiGene (UK). Hydrophobic PTFE membrane filter was purchased from Sterlitech Inc. HPV 16, 18 and 31 genomes were obtained from Dr. Jianxin You Lab at the University of Pennsylvania. Escherichia coli O157:H7 was purchased from ATCC (ATCC® 43895TM). LAMP primers and PCR primers were purchased from Integrated DNA Technologies, Inc. All other chemicals used in this study were analytical reagent grade or better. Homemade non-buffered LAMP reaction solution was prepared by mixing 10 μL 1M (NH4)2SO4, 1 μL Tween 20, 25 μL 2 M KCl, 4 μL 1M KOH, 140 μL dNTP, 40 μL Bst 2.0 DNA Polymerase and 270 μL ddH2O. LAMP primers mix was prepared by mixing 40 μL 100 µM FIP/BIP, 20 μL 100 µM LF/LB (or LF), and 5 μL 100 µM F3/B3 [25, 26]. The non-buffered LAMP reaction solution and LAMP primers mix were stored at -20 oC. UV-vis spectra were measured by a μ-Quant microplate reader Nanodrop 2000C (Thermo Scientific, USA). Real-time LAMP and PCR reactions were carried out on CFX96 Touch™ Real-Time PCR Detection System (Bio-Rad, USA).
LAMP experimental protocol
Three different LAMP reaction solutions (15 μL LAMP reaction) were used in our LAMP assay: i) homemade, non-buffered LAMP reaction solution: 7.5 μL pre-prepared non-buffered LAMP reaction solution, 3.6 μL ddH2O, 0.8 μL LAMP primers mix, 0.9 μL HNB and 1.2 μL 100 mM MgSO4; ii) NEB LAMP reaction buffer: 2.1 μL dNTP, 1.5 μL 10X isothermal amplification reaction buffer (New England BioLabs Inc), 0.6 μL Bst 2.0 DNA polymerase, 0.9 μL 100 mM MgSO4, 0.8 μL LAMP primers mix, 0.9 μL HNB and 7.2 μL ddH2O; and iii) Optigene GspSSD2.0 Isothermal Mastermix (Optigene, UK) for real-time fluorescence LAMP assay: 9 μL GspSSD2.0 Isothermal Mastermix, 3.7 μL dd H2O, 0.8 μL LAMP primers mix and 0.5 μL Eva green. All LAMP reactions in the presence/absence of 1 μL target (ddH2O for negative control) were incubated at 65 °C. Endpoint imaging and hue value analysis of colorimetric LAMP assay were achieved by Galaxy S6 smartphone (Samsung, South Korea) with our custom Hue Analyzer app (Figure S1).
Smartphone app for hue value analysis
The Hue Analyzer app (Figure S1) was developed with Eclipse Integrated Development Environment (IDE) in Android Developer Tools and Java. The app includes: i) Main menu, ii) Instructions, iii) Parameter settings and image capturing, and iv) Data analysis and result reporting. The app (Figure S1 (B)) provides operating instructions for user and allows the user to select the detection regions of the reactor array displayed in the preview window (Figure S1 (C)). The app takes photos of the microfluidic chip or tube at the end of LAMP reaction. All images are saved in Joint Photographic Experts Group (JPEG) format. The app analyzes the hue values from the user-defined regions and displays their results.
Microfluidic chip fabrication and operation
The 36 mm x 21 mm x 3.50 mm microfluidic chip was assembled by three layers: i) a 250 µm polymethyl methacrylate (PMMA) film top; ii) a 3 mm PMMA chip; iii) and a 250 µm PMMA film bottom (Figure S2). The top/bottom cover films were cut by a CO2 laser (Universal Laser Systems). The chip body was fabricated by a computer controlled milling machine (HAAS Automation Inc.) to create four LAMP reaction chambers, microchannels, inlet and outlet (Figure S2A). 1 μL of specific LAMP primers mix was pre-loaded into individual LAMP reaction chamber before bonding the chip body with PMMA film top by using acetonitrile [27]. In the experiment, the outlet of the chip (Figure S2B) was firstly covered by a hydrophobic porous membrane to provide an obstruction to liquid sample but not air, enabling all four reaction chambers to be fully filled. After thermally-treated lysate sample along with LAMP reaction solution (without LAMP primers) was introduced into the chip through its inlet (Figure S2B), the hydrophobic porous membrane was removed. Then, the inlet and outlet of the chip were sealed by PCR Sealers™ tape (Microseal® 'B' Film) (Bio-Rad) and the chip was inserted into our homemade smart cup for LAMP amplification.
Portable smart cup platform
Smart cup platform developed previously by our Lab [23, 24] was used for LAMP reaction without need of electricity, enabling instrument-free POC molecular diagnostics. Briefly, the smart cup (Figure S3) utilized water-triggered, exothermic chemical reaction to supply heat for isothermal amplification. In the test, one Mg-Fe alloy pouch and 7.5 mL of tap water were added into the smart cup to produce chemical heat for isothermal amplification. The temperature was regulated by a phase change material (PCM) (Innotech Products Ltd., USA) that melts at 68 °C. After preheating the smart cup for approximately 10 min, the microfluidic chip loaded with samples was inserted into the smart cup for isothermal amplification. After 60 min incubation for LAMP amplification, the chip image was taken by the smartphone and the hue value of each reaction chamber was quantitatively read by the Hue Analyzer app. The smart cup is reusable except for Mg-Fe alloy pouch (~0.15 $ per pouch) and the estimated cost for chip material and assay reagents is ~2 $ per assay, enabling simple, cost-effective POC molecular diagnostics.
Saliva samples preparation
Saliva samples were collected from participants without swallowing or expectorating for 5 min and then spit into the collection vial (IRB protocol #: 823266). 2 mL saliva was collected through this protocol. Known concentration HPV DNA (i.e., HPV 16, 18 and 31) were spiked into saliva samples and then incubated at 95 oC for 4 min to inactivate potential inhibitors of LAMP reaction existing in saliva matrix. The saliva samples could be used immediately or stored at -20 °C until use. To detect spiked HPV DNA in saliva samples, 37.5 μL non-buffered LAMP reaction solution, 23 μL saliva samples, 4.5 μL HNB and 6 μL 100 mM MgSO4 were initially mixed in a tube and then added into the microfluidic chip by a digital pipette (Figure S2B).
Clinical cervical swab samples preparation
Clinical cervical swab samples obtained from the Hospital of the University of Pennsylvania (IRB protocol #: 829760) were utilized for liquid-based Pap smear test, real-time quantitative PCR and HPV genotype analysis in our smart cup-based POC diagnostic platform, respectively. The Pap smear test was completed at the Hospital of the University of Pennsylvania. For HPV DNA detection by real-time PCR and our smart cup platform, 200 μL cervical swab samples were firstly centrifuged at 1000X g for 5 min. Next, the liquid supernatant including methanol was removed and left cell pellet. Then, the concentrated cervical cells were resuspended by 50 μL ddH2O and repeated the above steps for 3 times. Lastly, the concentrated cervical cells were incubated at 95 °C for 10 min for cell lysis and released the nucleic acids [28]. After cells lysis at 95 °C for 10 min, the solution was concentrated at 1000X g for 10 seconds and the liquid supernatant was used for further LAMP amplification. For LAMP reaction in our smart cup platform, 37.5 μL non-buffered LAMP reaction solution, 23 μL cervical cell lysis solution, 4.5 μL HNB, 4 μL ddH2O and 6 μL 100 mM MgSO4 were mixed and added into the four-chamber chip which have already been pre-loaded with HPV 16, 18 and 31 LAMP primers (ddH2O for blank control group) in their individual chambers, respectively. Then, the microfluidic chip was incubated in the smart cup platform for 60 min and the images were recorded by smartphone camera and analyzed by the Hue Analyzer app. The hue value of 238° was defined as the threshold value at 5 times standard deviations below the mean baseline level (blank control). Real-time PCR detection of HPV DNAs was similar to the previously reported method [29].
Results and Discussion
Synergistically enhanced colorimetric molecular detection
A synergistically enhanced colorimetric detection strategy for LAMP assay has, for the first time, been proposed by taking advantage of the synergistic effect of pyrophosphate (PPi4-) and H+ ions, two byproducts of LAMP reaction in non-buffered LAMP reaction solution (without Tris salt) (Figure 1A). The polymerization during LAMP reaction produces large amounts of byproducts including PPi4- and H+ ions. When LAMP reaction occurs in non-buffered solution, its pH value changes from 8.8 to ~ 6.8. When HNB is added to LAMP reaction solution as a colorimetric indicator, its color changes from initial blue to fuchsia due to the electron transfer from HNB to Mg2+ after formation of HNB-Mg2+ complex. During LAMP reaction, the produced pyrophosphate ions react with Mg2+ ions and form insoluble product magnesium pyrophosphate (Mg2P2O7), which promotes the dissociation of HNB-Mg2+ complex (Figure 1A). Simultaneously, the produced H+ ion in the non-buffered LAMP reaction solution can also decrease the stability of HNB-Mg2+ complex and further release the blue HNB (Figure 1A) [30-32]. Thus, the synergistic reaction of PPi4- and H+ ions with HNB-Mg2+ complex significantly improves the release of blue HNB, producing an enhanced colorimetric signal in LAMP assay. To validate it, different concentrations of Mg2+ ions were added in LAMP reaction solution with 120 µM of HNB at pH values of 8.8 and 6.8, respectively. As shown in Figure 1B, the hue value change of HNB indicator between 4 mM and 8 mM Mg2+ was, respectively, ~25o and ~36o at a constant pH value (8.8) and various pH values (8.8 to 6.8), which showed that synergistic effect can significantly improve the colorimetric LAMP detection. Due to absence of buffer solution, the pH value in the non-buffered LAMP solution dropped more than that of the buffered LAMP solution after LAMP amplification (Figure S4). But we did not observe any obvious effect of the non-buffered LAMP solution on the LAMP reaction, which is consistent with the previous literature [10]. In addition, there is no significant difference in pH change for different nucleic acid targets (e.g., HPV 16, E. coli O157, Salmonella and λDNA) in our non-buffered LAMP assay (Figure S5). All of these results show that non-buffered LAMP solution is suitable for high sensitive colorimetric detection of nucleic acids.
Synergistically enhanced colorimetric detection strategy of LAMP assay in non-buffered reaction solution. (A) Detection mechanism. (B) Colorimetric detection and hue value analysis of HNB indicator with 4, 6 and 8 mM Mg2+ at a constant pH value (8.8) and various pH values (8.8 to 6.8), respectively. (C) Colorimetric LAMP detection of HPV 16 DNA and hue value analysis in non-buffered LAMP solution and NEB LAMP reaction buffer, respectively. Error bars denote s.d. (n=3).
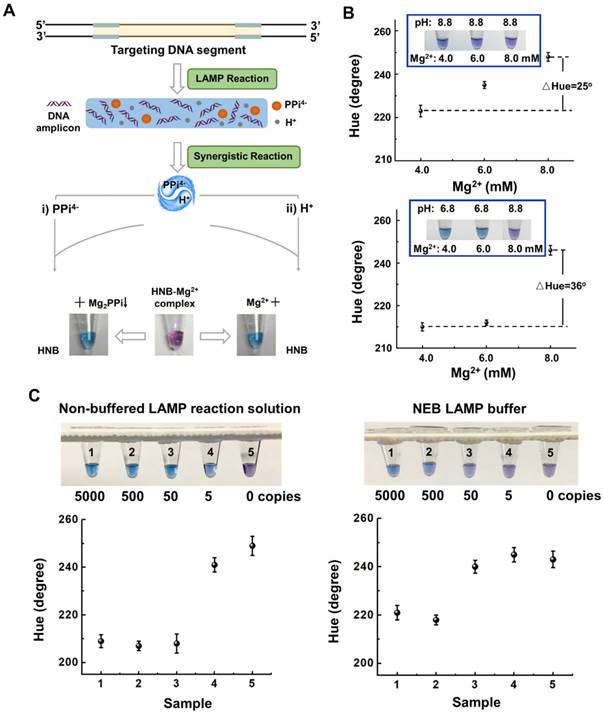
To evaluate the sensitivity of our synergistically enhanced colorimetric detection method, we chose HPV 16 as a model analyte. We carried out colorimetric LAMP assay with 10-fold serially diluted DNA template in the homemade non-buffered LAMP reaction solution and NEB LAMP reaction buffer (New England Biolabs, MA), respectively. A limit of detection of 50 copies per test for HPV 16 can be achieved with the non-buffered LAMP solution, which is 10-fold higher than that of NEB LAMP reaction buffer (Figure 1C). To further validate that the synergistic effect improves colorimetric LAMP detection compared to single pH change, 10 mU of thermostable inorganic pyrophosphatase was added into the non-buffered LAMP solution to digest pyrophosphate and maintain a constant Mg2+ ion concentration. As shown in Figure S6A and S6B, a more significant color change was observed in our synergistically enhanced colorimetric detection without pyrophosphatase. In addition, HNB indicator showed better sensitivity than phenol red (pH sensitive dye) in our non-buffered colorimetric LAMP assay (Figure 1C and Figure S6C). Therefore, the synergistic effect in the non-buffered LAMP solution not only significantly enhances the color change between positive and negative samples, but also improves the detection sensitivity of LAMP reaction, allowing us to develop an ultrasensitive colorimetric molecular detection.
Smartphone-based hue image analysis
Colorimetric readout with naked eye is a simple, straightforward and instrument-free biochemical analysis method. However, it is susceptible to variations in human color perception. To this end, a custom Hue Analyzer app was developed for objective and reliable colorimetric readout by quantifying hue value of images, eliminating need for sophisticated optical instrumentation (Figure 2A and Figure S1). The app can take photos, process images, quantify hue values of images, as well as report results. In color models, the hue, saturation and intensity (HSI) have widely applied for artificial intelligence image analysis such as real-time flame detection, lane-marking detection as well as analytical chemistry [15, 33]. The hue component describes the attribute of pure color, which is close to human perception of color and its sensitivity to image noise is lower compared with saturation and intensity. Hue value can be represented as an angle in the color ranging from 0 to 360°. To investigate the reliability of our smartphone-based hue image analysis method, we determined the hue values of a series of 120 µM HNB indicator solution with different concentration of Mg2+ (0, 2, 4, 6, 8, and 10 mM) by the Hue Analyzer app (Figure 2A and B). The hue values changed from 205° to 260° and showed a good linear relationship (r2 = 0.993) at Mg2+ ion concentrations ranging from 0 to 10 mM (Figure 2C). For comparison, absorption spectra of HNB indicator solution were measured with different Mg2+ ion concentrations by determining their absorbance peak at 650 nm (Figure 2D) on conventional UV spectrophotometer. The results show that our smartphone-based hue analysis method has wider linear range and lower variation compared to UV spectrophotometer method (Figure 2C and E).
Comparison of smartphone-based hue value analysis method and traditional absorption spectrometry in colorimetric detection. (A) Custom app interface. (B) Optical image of tubes containing 120 µM HNB indicator and various Mg2+ ion concentrations. (C) Hue values of HNB indicator as a function of various concentrations of Mg2+ ions. (D) Absorption spectra of HNB with various Mg2+ ion concentrations. (E) Absorbance peak of HNB indicator at 650 nm as a function of various Mg2+ ion concentrations. Error bars denote s.d. (n=3).
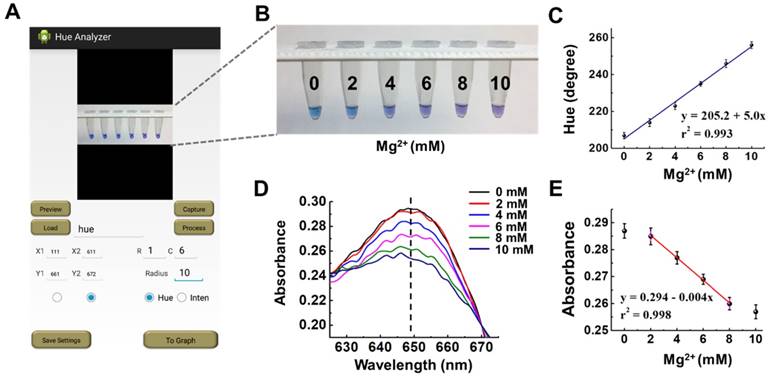
In addition to human color perception difference, colorimetric detection can be affected by variations of environment light. We evaluated the effect of different environmental light on hue image analysis and compared its performance with that of RGB value detection which has been widely used for color analysis, including colorimetric LAMP assay [12, 34, 35]. Unlike traditional RGB model, hue value in HSI model is independent of the light intensity because it is multiplicative/scale invariant [36]. As shown in Figure S7, the hue value analysis exhibits more reliable and reproducible results compared to RGB model. Additionally, we tested the long-term stability of hue-based colorimetric detection in LAMP assay. As shown in Figure S8, the hue image analysis shows that there is no significant hue value change in LAMP reaction products after more than 21-days storage under daylight exposure at room temperature, enabling POC molecular diagnostics without need for dark storage condition. In addition, the sensitivity and specificity of smartphone-based hue value analysis method in the non-buffered colorimetric LAMP assay is comparable with that of conventional fluorescence detection in Optigene GspSSD2.0 Isothermal Mastermix (Figure S9 to Figure S13). These results show that the smartphone-based hue image analysis is a reliable, stable and sensitive approach for objective, digital colorimetric detection without need of sophisticated optical instrumentation, allowing us to develop simple, inexpensive, instrument-free POC diagnostics.
Multiple HPV DNA detection in saliva samples
To validate the feasibility of our proposed method for simultaneous detection of multiple nucleic acid targets at the point of care, we designed and fabricated a four-chamber microfluidic chip (Figure 3A, top, and Figure S2) to simultaneously detect three high-risk HPV genotypes, HPV 16, 18 and 31 in saliva and clinical swab samples. The LAMP primers targeted to HPV 16, 18, and 31 (Table S1) were pre-stored and dried in chambers 2, 3 and 4, respectively. The specific LAMP primers pre-stored in the chambers determine which HPV DNA will be amplified. Chamber 1 serves as a blank control without any primers (Figure 3A, top). If needed, more chambers can be added in the chip to simultaneously detect more nucleic acid targets.
HPV DNA detection in saliva samples in our POC diagnostic platform. (A) Photographs of four-chamber microfluidic chip (top) and smart cup equipped with smartphone (bottom). (B) Screenshots of the app interfaces: imaging/ parameter setting (left) and hue value quantitative readout (right). (C) Detection of saliva samples spiked 103 copies of HPV 16, 18 and 31, respectively. *** indicates a significant difference in the hue value (p < 0.001, t-test). Error bars denote s.d. (n=3).
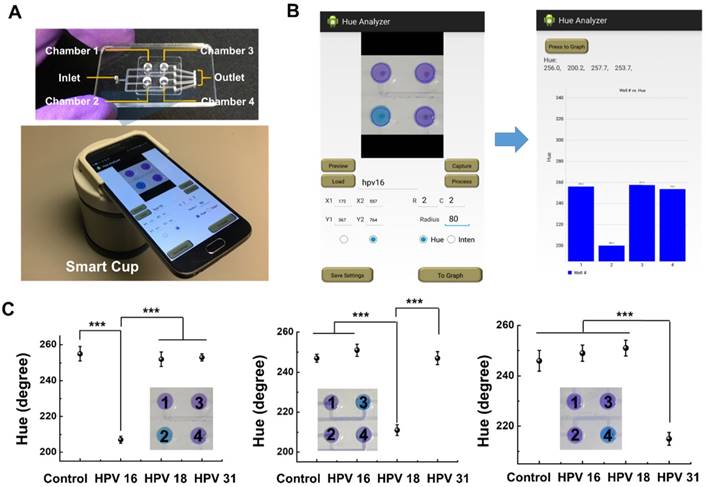
Saliva-based screening of high-risk HPV DNA is a simple, non-invasive and reliable method for early diagnosis of HPV-related cancers, such as oral/oropharyngeal cancer and throat cancer [37]. In the testing, saliva samples spiked with HPV DNA were first incubated at 95 °C for 4 min to inactivate potential inhibitors before introducing into the microfluidic chip. Next, saliva samples were mixed with non-buffered LAMP reaction solution (no LAMP primers) and pipetted into the microfluidic chip with pre-stored HPV LAMP primers. Then, the chip was inserted into the smart cup (Figure 3A, bottom, Figure S3) for isothermal amplification. After 60-min incubation, the chip image was taken by the smartphone (Figure 3B, left) and hue value of each chamber was quantitatively determined by the Hue Analyzer app (Figure 3B, right). The combination of simple microfluidic chip with our portable smart cup platform allows us to develop an inexpensive, instrument-free molecular point of care diagnostics.
As shown in Figure 3C, the groups of saliva samples spiked with 103 copies of HPV 16, 18, and 31 DNAs, respectively shows significantly different hue values with control groups (p < 0.001, t-test). In our experiment, we did not observe any significant effect of potential buffering capacity of saliva (Figure 3C and Figure S14). As shown in Figure 1C, the sensitivity of the developed smartphone-based hue value analysis method for HPV DNA detection could reach 102 copies, which is comparable to that of previous methods [25,38]. Additionally, we further demonstrated the feasibility of our method for simultaneous detection of multiple HPV genotypes in the same saliva sample in the smart cup platform (Figure S15). All these results verified that saliva samples do not influence our colorimetric LAMP detection, which has great potential in saliva-based diagnostics.
HPV-associated cervical cancer screening in clinical swab samples
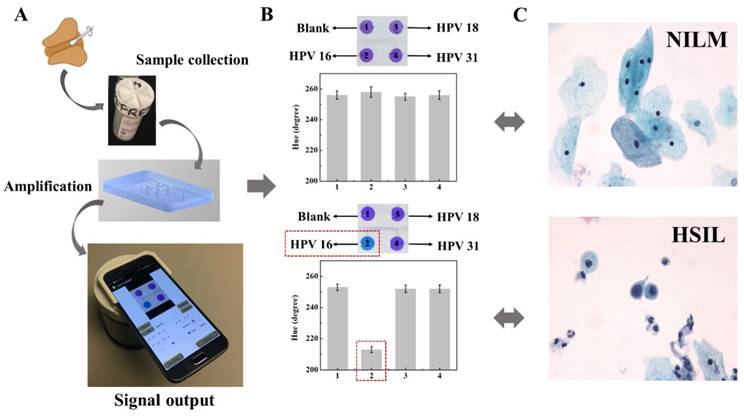
To further validate the clinical application of our smart cup diagnostic platform, we tested fifteen clinical swab samples including three positive samples and twelve negative samples. The collected clinical samples can be directly added into the microfluidic chip after thermal lysis [28]. Results can be easily recorded and reported by smartphone after 60-min amplification (Figure 4A, B and Figure S16). As shown in Table 1, the results obtained from the smart cup platform agree well with that of real-time PCR method [29]. The detection results of the smart cup are also compared and correlated with that of conventional Pap smear test which screens for abnormal cell changes (Figure 4C and Table 1). Therefore, our smartphone-based synergistically enhanced colorimetric molecular detection platform is suitable for clinical applications in cervical cancer screening, and other infectious disease detection at the point of care.
Cervical cancer screening with clinical swab samples in our POC diagnostic platform. (A) Screening workflow. (B) Detection results of our smart cup platform. Red dash box indicates positive. (C) Pap smear results (HSIL: High-Grade Squamous Intraepithelial Lesion; NILM: Negative for Intraepithelial Lesion or Malignancy).
Cervical cancer screening with clinical swab samples by our smart cup platform, real-time PCR method and Pap smear test.
| Clinical swab samples | Smart cup platform | Real time PCR | Pap smear test | ||||
|---|---|---|---|---|---|---|---|
| HPV16 | HPV18 | HPV31 | HPV16 | HPV18 | HPV31 | ||
| 1 | - | - | - | - | - | - | NILM |
| 2 | - | - | - | - | - | - | NILM |
| 3 | - | - | - | - | - | - | NILM |
| 4 | - | - | - | - | - | - | NILM |
| 5 | - | - | - | - | - | - | NILM |
| 6 | - | - | - | - | - | - | NILM |
| 7 | - | - | - | - | - | - | NILM |
| 8 | - | - | - | - | - | - | NILM |
| 9 | - | - | - | - | - | - | NILM |
| 10 | - | - | - | - | - | - | NILM |
| 11 | - | - | - | - | - | - | NILM |
| 12 | - | - | - | - | - | - | NILM |
| 13 | - | - | + | - | - | + | HSIL |
| 14 | - | + | - | - | + | - | ASCUS |
| 15 | + | - | - | + | - | - | HSIL |
+: detected; -: not detected; NILM: Negative for Intraepithelial Lesion or Malignancy; HSIL: High-Grade Squamous Intraepithelial Lesion; ASCUS: Atypical squamous cells of undetermined significance
Conclusion
In summary, we proposed a synergistically enhanced colorimetric method for LAMP assay and developed smart cup-based POC diagnostic platform for HPV-associated cervical cancer screening at the point of care. Our method allows the rapid, portable, and inexpensive molecular detection of a wide range of nucleic acid targets without need for complex equipment, offering several advantages: i) the synergistic enhancing effect in colorimetric LAMP assay not only produces improved color difference, but increases the detection sensitivity, ii) smartphone-based hue image analysis method provides reliable, objective, digital readout in colorimetric detection without need of any complex optical detector, and iii) the chemically heated-smart cup combined with disposable microfluidic chips enables instrument-free POC molecular diagnostics. The POC diagnostic platform described here has great potential for personalized health monitoring and prevention of cancer and infectious disease at the point of care and in resource-limited clinical settings.
Abbreviations
HNB: hydroxy naphthol blue; HPV: human papillomavirus; HSI: hue, saturation and intensity; LAMP: loop-mediated isothermal amplification; PCR: polymerase chain reaction; PMMA: polymethyl methacrylate; POC: point of care; RGB: red green blue.
Supplementary Material
Supplementary figures and table.
Acknowledgements
The work was supported, in part, by R01EB023607, R01CA214072, R01CA187718 and R21TW010625.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Shaman J. Pandemic preparedness and forecast. Nat Microbiol. 2018;3:265
2. Schiffman M, Castle PE, Jeronimo J, Rodriguez AC, Wacholder S. Human papillomavirus and cervical cancer. Lancet. 2007;370:890-907
3. Ogilvie GS, van Niekerk D, Krajden M, Smith LW, Cook D, Gondara L. et al. Effect of screening with primary cervical HPV testing vs cytology testing on high-grade cervical intraepithelial neoplasia at 48 months: the HPV FOCAL randomized clinical trial. JAMA. 2018;320:43-52
4. Castle PE, Smith KM, Davis TE, Schmeler KM, Ferris DG, Savage AH. et al. Reliability of the Xpert HPV assay to detect high-risk human papillomavirus DNA in a colposcopy referral population. Am J Clin Pathol. 2015;143:126-33
5. Loo J, Kwok H, Leung C, Wu S, Law I, Cheung Y. et al. Sample-to-answer on molecular diagnosis of bacterial infection using integrated lab-on-a-disc. Biosens Bioelectron. 2017;93:212-9
6. Du Y, Pothukuchy A, Gollihar JD, Nourani A, Li B, Ellington AD. Coupling sensitive nucleic acid amplification with commercial pregnancy test strips. Angew Chem Int Ed Engl. 2017;56:992-6
7. Soli KW, Kas M, Maure T, Umezaki M, Morita A, Siba PM. et al. Evaluation of colorimetric detection methods for Shigella, Salmonella, and Vibrio cholerae by loop-mediated isothermal amplification. Diagn Microbiol Infect Dis. 2013;77:321-3
8. Kadimisetty K, Song J, Doto AM, Hwang Y, Peng J, Mauk MG. et al. Fully 3D printed integrated reactor array for point-of-care molecular diagnostics. Biosens Bioelectron. 2018;109:156-63
9. Kumvongpin R, Jearanaikool P, Wilailuckana C, Sae-ung N, Prasongdee P, Daduang S. et al. High sensitivity, loop-mediated isothermal amplification combined with colorimetric gold-nanoparticle probes for visual detection of high risk human papillomavirus genotypes 16 and 18. J Virol Methods. 2016;234:90-5
10. Tanner NA, Zhang Y, Evans Jr TC. Visual detection of isothermal nucleic acid amplification using pH-sensitive dyes. Biotechniques. 2015;58:59-68
11. Goto M, Honda E, Ogura A, Nomoto A, Hanaki K-I. Colorimetric detection of loop-mediated isothermal amplification reaction by using hydroxy naphthol blue. Biotechniques. 2009;46:167-72
12. Rodriguez-Manzano J, Karymov MA, Begolo S, Selck DA, Zhukov DV, Jue E. et al. Reading out single-molecule digital RNA and DNA isothermal amplification in nanoliter volumes with unmodified camera phones. ACS nano. 2016;10:3102-13
13. Karthik K, Rathore R, Thomas P, Arun T, Viswas K, Dhama K. et al. New closed tube loop mediated isothermal amplification assay for prevention of product cross-contamination. MethodsX. 2014;1:137-43
14. Brigham R, Grau-Bové J, Rudnicka A, Cassar M, Strlic M. Crowdsourcing as an Analytical Method: Metrology of Smartphone Measurements in Heritage Science. Angew Chem Int Ed Engl. 2018
15. Wang X, Mahoney M, Meyerhoff ME. Inkjet-Printed Paper-Based Colorimetric Polyion Sensor Using a Smartphone as a Detector. Anal Chem. 2017;89:12334-41
16. Barnes L, Heithoff DM, Mahan SP, Fox GN, Zambrano A, Choe J. et al. Smartphone-based pathogen diagnosis in urinary sepsis patients. EBioMedicine. 2018;36:73-82
17. Mauk MG. Calling in the test: Smartphone-based urinary sepsis diagnostics. EBioMedicine. 2018;37:11-2
18. Kong JE, Wei Q, Tseng D, Zhang J, Pan E, Lewinski M. et al. Highly stable and sensitive nucleic acid amplification and cell-phone-based readout. ACS nano. 2017;11:2934-43
19. Priye A, Ball CS, Meagher RJ. Colorimetric-Luminance Readout for Quantitative Analysis of Fluorescence Signals with a Smartphone CMOS Sensor. Anal Chem. 2018;90:12385-9
20. Kaarj K, Akarapipad P, Yoon J-Y. Simpler, Faster, and Sensitive Zika Virus Assay Using Smartphone Detection of Loop-mediated Isothermal Amplification on Paper Microfluidic Chips. Sci Rep. 2018;8:12438
21. Priye A, Bird SW, Light YK, Ball CS, Negrete OA, Meagher RJ. A smartphone-based diagnostic platform for rapid detection of Zika, chikungunya, and dengue viruses. Sci Rep. 2017;7:44778
22. Shen L, Hagen JA, Papautsky I. Point-of-care colorimetric detection with a smartphone. Lab Chip. 2012;12:4240-3
23. Song J, Pandian V, Mauk MG, Bau HH, Cherry S, Tisi LC. et al. Smartphone-Based Mobile Detection Platform for Molecular Diagnostics and Spatiotemporal Disease Mapping. Anal Chem. 2018;90:4823-31
24. Liao S-C, Peng J, Mauk MG, Awasthi S, Song J, Friedman H. et al. Smart cup: a minimally-instrumented, smartphone-based point-of-care molecular diagnostic device. Sens Actuators B Chem. 2016;229:232-8
25. Satoh T, Matsumoto K, Fujii T, Sato O, Gemma N, Onuki M. et al. Rapid genotyping of carcinogenic human papillomavirus by loop-mediated isothermal amplification using a new automated DNA test (Clinichip HPV™). J Virol Methods. 2013;188:83-93
26. Zhao X, Li Y, Wang L, You L, Xu Z, Li L. et al. Development and application of a loop-mediated isothermal amplification method on rapid detection Escherichia coli O157 strains from food samples. Mol Biol Rep. 2010;37:2183-8
27. Liu C, Geva E, Mauk M, Qiu X, Abrams WR, Malamud D. et al. An isothermal amplification reactor with an integrated isolation membrane for point-of-care detection of infectious diseases. Analyst. 2011;136:2069-76
28. Pipper J, Zhang Y, Neuzil P, Hsieh TM. Clockwork PCR including sample preparation. Angew Chem Int Ed Engl. 2008;47:3900-4
29. Moberg M, Gustavsson I, Gyllensten U. Real-time PCR-based system for simultaneous quantification of human papillomavirus types associated with high risk of cervical cancer. J Clin Microbiol. 2003;41:3221-8
30. Yin K, Lv M, Wang Q, Wu Y, Liao C, Zhang W. et al. Simultaneous bioremediation and biodetection of mercury ion through surface display of carboxylesterase E2 from Pseudomonas aeruginosa PA1. Water Res. 2016;103:383-90
31. Schiewer S, Volesky B. Modeling of the proton-metal ion exchange in biosorption. Environ Sci Technol. 1995;29:3049-58
32. Schuster E. The behavior of mercury in the soil with special emphasis on complexation and adsorption processes-a review of the literature. Water Air Soil Pollut. 1991;56:667-80
33. Wang X, Zhang Q, Nam C, Hickner M, Mahoney M, Meyerhoff ME. An Ionophore-Based Anion-Selective Optode Printed on Cellulose Paper. Angew Chem Int Ed Engl. 2017;56:11826-30
34. Su K, Zou Q, Zhou J, Zou L, Li H, Wang T. et al. High-sensitive and high-efficient biochemical analysis method using a bionic electronic eye in combination with a smartphone-based colorimetric reader system. Sens Actuators B Chem. 2015;216:134-40
35. de Oliveira HJS, de Almeida Jr PL, Sampaio BA, Fernandes JPA, Pessoa-Neto OD, de Lima EA. et al. A handheld smartphone-controlled spectrophotometer based on hue to wavelength conversion for molecular absorption and emission measurements. Sens Actuators B Chem. 2017;238:1084-91
36. Cappi G, Spiga FM, Moncada Y, Ferretti A, Beyeler M, Bianchessi M. et al. Label-free detection of tobramycin in serum by transmission-localized surface plasmon resonance. Anal Chem. 2015;87:5278-85
37. Wasserman JK, Rourke R, Purgina B, Caulley L, Dimitroulakis J, Corsten M. et al. HPV DNA in saliva from patients with SCC of the head and neck is specific for p16-positive oropharyngeal tumours. J Otolaryngol Head Neck Surg. 2017;46:3
38. Sargent A, Bailey A, Turner A, Almonte M, Gilham C, Baysson H. et al. Optimal threshold for a positive hybrid capture 2 test for detection of human papillomavirus: data from the ARTISTIC trial. J Clin Microbiol. 2010;48:554-8
Author contact
Corresponding author: Changchun Liu, Ph.D. E-mail: chaliuedu
 Global reach, higher impact
Global reach, higher impact