13.3
Impact Factor
Theranostics 2018; 8(7):1956-1965. doi:10.7150/thno.23767 This issue Cite
Research Paper
Integrative analysis of imaging and transcriptomic data of the immune landscape associated with tumor metabolism in lung adenocarcinoma: Clinical and prognostic implications
1. Department of Nuclear Medicine, Seoul National University Hospital, Seoul, Republic of Korea.
2. Cheonan Public Health Center, Chungnam, Republic of Korea.
3. Department of Community Health, Korea Health Promotion Institute, Seoul, Republic of Korea.
4. Department of Clinical Medical Sciences, Seoul National University, College of Medicine, Seoul, Republic of Korea.
*These authors contributed equally to this work
Abstract
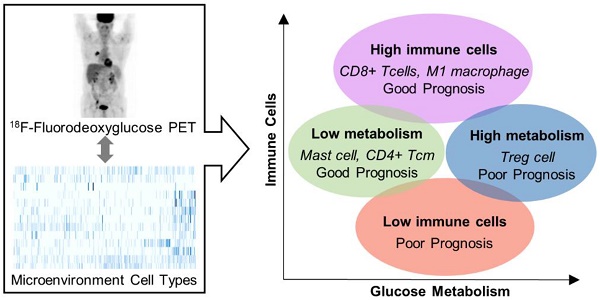
Although metabolic modulation in the tumor microenvironment (TME) is one of the key mechanisms of cancer immune escape, there is a lack of understanding of the comprehensive immune landscape of the TME and its association with tumor metabolism based on clinical evidence. We aimed to investigate the relationship between the immune landscape in the TME and tumor glucose metabolism in lung adenocarcinoma.
Methods: Using RNA sequencing and image data, we developed a transcriptome-based tumor metabolism estimation model. The comprehensive TME cell types enrichment scores and overall immune cell enrichment (ImmuneScore) were estimated. Subjects were clustered by cellular heterogeneity in the TME and the clusters were characterized by tumor glucose metabolism and immune cell composition. Moreover, the prognostic value of ImmuneScore, tumor metabolism and the cell type-based clusters was also evaluated.
Results: Four clusters were identified based on the cellular heterogeneity in the TME. They showed distinct immune cell composition, different tumor metabolism, and close relationship with overall survival. A cluster with high regulatory T cells showed relative hypermetabolism and poor prognosis. Another cluster with high mast cells and CD4+ central memory T cells showed relative hypometabolism and favorable prognosis. A cluster with high ImmuneScore showed favorable prognosis. Multivariate Cox analysis demonstrated that ImmuneScore was a predictive prognostic factor independent of other clinical features.
Conclusions: Our results showed the association between predicted tumor metabolism and immune cell composition in the TME. Our studies also suggest that tumor glucose metabolism and immune cell infiltration in the TME can be clinically applicable for developing a prognostic stratification model.
Keywords: tumor microenvironment, cancer immunoediting, tumor metabolism, next generation sequencing, lung adenocarcinoma
 Global reach, higher impact
Global reach, higher impact