Impact Factor
Theranostics 2026; 16(11):6011-6031. doi:10.7150/thno.128026 This issue Cite
Research Paper
Edge-rich Cu-N3 single atom nanozyme drives lipid switching to potentiate tumor catalytic therapy
1. Department of Stomatology, The First Affiliated Hospital of Wannan Medical College (Yijishan Hospital of Wannan Medical College), Wuhu 241001, China.
2. Department of Nutrition and Food Hygiene, School of Public Health; Department of Radiology, Zhongda Hospital, Nurturing Center of Jiangsu Province for State Laboratory of AI Imaging & Interventional Radiology, School of Medicine; Southeast University, Nanjing 210009, China.
3. Central Laboratory, Department of Biobank, Nanjing Hospital of Chinese Medicine Affiliated to Nanjing University of Chinese Medicine, Nanjing 210022, China.
4. Key Laboratory of Medical Imaging Precision Theranostics and Radiation Protection, College of Hunan Province, Hengyang Medical School, The Affiliated Changsha Central Hospital, University of South China, Changsha, China.
5. Max Planck Institute of Microstructure Physics, Weinberg 2, Halle, 06120, Germany.
#These authors contributed equally to this work.
Received 2025-11-7; Accepted 2026-3-11; Published 2026-4-8
Abstract
Rationale: Tumor catalytic therapy represents a promising antitumor approach by inducing ferroptosis and overcoming apoptosis-related resistance mechanisms. Its efficacy is primarily dictated by the extent of membrane lipid peroxidation (LPO). However, tumor cells may evade ferroptosis through metabolic reprogramming that enriches monounsaturated fatty acids (MUFAs) in membrane lipids, thereby diminishing oxidative vulnerability. Hence, strategies that simultaneously enhance catalytic ROS production and reprogram lipid metabolism are required to address this challenge.
Methods: To overcome this limitation, a novel Cu-N3 single-atom nanozyme with edge enrichment (ER-Cu1SAZyme) was developed, characterized by a hollow porous structure and catalytically active sites concentrated along the edges. This design optimizes atom utilization, increases local electronic density, and lowers the reaction energy barrier, thereby promoting potent intracellular reactive oxygen species (ROS) generation. To further sensitize tumors to ferroptosis, ER-Cu1SAZyme was combined with sirolimus (Srl), an FDA-approved drug, to create the Srl/ER-Cu1SAZyme nanomedicine for coordinated catalytic and metabolic regulation.
Results: The Srl/ER-Cu1SAZyme formulation simultaneously inhibits stearoyl-CoA desaturase 1 (SCD1)-mediated MUFA synthesis and upregulates ACSL4, thereby shifting the membrane lipid composition toward a ferroptosis-sensitive phenotype and enhancing nanozyme-induced LPO. This dual catalytic-metabolic strategy increases ferroptosis susceptibility while reducing metastatic potential linked to excessive membrane fluidity. In tumor-bearing mouse models, Srl/ER-Cu1SAZyme treatment led to a 33-fold reduction in tumor volume compared to the untreated group, without observable systemic toxicity.
Conclusions: These results highlight the effectiveness of integrating edge-enriched single-atom catalysis with lipid metabolic modulation to enhance ferroptosis-based tumor therapy. The Srl/ER-Cu1SAZyme nanomedicine offers a safe and highly potent approach for dual catalytic-metabolic regulation in cancer treatment.
Keywords: nanozyme, single atom catalyst, catalytic therapy, ferroptosis, tumor therapy
Introduction
Nanozymes are synthetic nanomaterials that mimic the catalytic functions of natural enzymes, gaining increasing attention in tumor therapy [1-4]. By catalyzing redox reactions and facilitating substrate conversions in vivo [5-7], nanozymes have emerged as promising agents for tumor catalytic treatment, particularly in inducing ferroptosis [8, 9]. Ferroptosis is a regulated form of cell death triggered by excessive oxidative stress and the accumulation of lipid peroxides (LPO), which leads to irreversible membrane damage [10]. This mechanism offers an effective alternative for eliminating tumor cells resistant to apoptosis-based therapies [11-14]. However, many tumor cells acquire adaptive resistance to oxidative damage by reprogramming lipid metabolism, typically by increasing the proportion of monounsaturated fatty acids (MUFAs) in membrane lipids [15-18]. The single double bond in MUFAs imparts high oxidative stability, protecting membrane integrity and suppressing lipid peroxidation (LPO), thereby reducing sensitivity to ferroptosis. Correcting this metabolic shift, by reducing MUFA levels and restoring the ratio of polyunsaturated fatty acids (PUFAs) to MUFAs, has thus become a promising strategy to enhance ferroptosis and improve therapeutic outcomes.
Recent developments in nanozyme engineering have significantly enhanced catalytic performance by optimizing the structural precision and atomic distribution of active sites [19]. Among these, single-atom nanozymes (SAZymes) represent a new generation of catalytic materials, offering maximized atomic efficiency and well-defined coordination environments. These properties enable superior catalytic activity and selectivity compared to conventional nanoparticle-based systems [20]. However, despite these advances, the catalytic efficiency of many SAzymes remains limited by suboptimal atomic utilization and electronic structure [21]. Most conventional SAzymes exhibit in-plane M-N4 coordination, where only a fraction of catalytic atoms are accessible for substrate interaction, and the local electronic configuration is not ideally tuned for reactive oxygen species (ROS) generation [22]. To overcome these limitations, developing single-atom nanozymes with abundant accessible active sites and optimized atomic coordination is crucial. Such advancements would amplify oxidative stress within the tumor microenvironment (TME) while disrupting metabolic pathways that facilitate membrane remodeling.
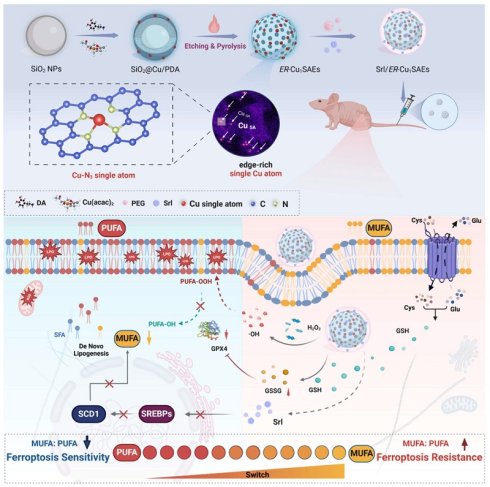
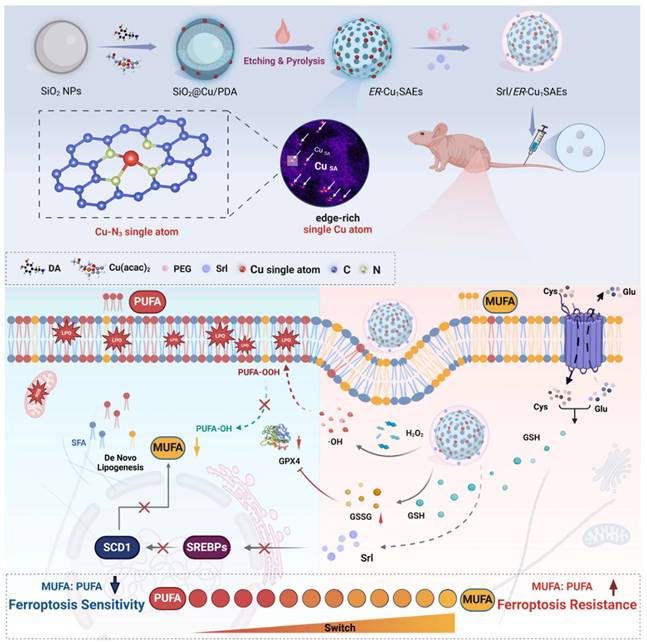
In this study, a novel edge-enriched Cu-N3 single-atom nanozyme (ER-Cu1SAZyme) was designed to meet the dual requirements of enhanced catalytic efficiency and metabolic regulation. Unlike conventional planar SAzymes, ER-Cu1SAZyme features a hollow porous nitrogen-doped carbon framework with Cu atoms preferentially anchored at edge sites, forming abundant Cu-N3 coordination centers. This unique edge-enriched architecture increases atomic utilization, boosts local electronic density, and reduces the reaction energy barrier, thereby facilitating the efficient conversion of H2O2 into highly reactive hydroxyl radicals (•OH) and depleting intracellular glutathione (GSH). The resultant ROS surge disrupts redox balance, triggers extensive LPO, and ultimately induces ferroptosis. Additionally, the hollow, porous structure of ER-Cu1SAZyme serves as an ideal platform for loading and delivering small molecules, including clinical drugs. To sensitize tumors to ferroptosis, the FDA-approved drug sirolimus (Srl) was incorporated into ER-Cu1SAZyme, forming the Srl/ER-Cu1SAZyme nanomedicine. Srl inhibits stearoyl-CoA desaturase 1 (SCD1)-mediated MUFA biosynthesis while promoting ACSL4-mediated PUFA utilization, thereby shifting membrane lipid composition toward a ferroptosis-susceptible phenotype. This synergistic combination amplifies ferroptosis sensitivity while mitigating the excessive membrane fluidity and metastatic potential often associated with PUFA supplementation. As illustrated in Scheme 1, Srl/ER-Cu1SAZyme integrates single-atom catalytic ROS generation with metabolic modulation of membrane lipids, offering a robust and safe ferroptosis-based antitumor strategy. This approach presents a promising solution and a new opportunity to conjugate nanocatalysts with clinical drugs for more effective tumor treatment.
Schematic illustration of Srl/ER-Cu1SAZyme in tumor catalytic therapy. The edge-enriched Cu-N3 single-atom nanozyme (ER-Cu1SAZyme) induces intense oxidative stress in tumor cells by depleting intracellular glutathione (GSH) and catalyzing the conversion of hydrogen peroxide (H2O2) into reactive hydroxyl radicals (•OH), triggering oxidative damage. Moreover, the FDA-approved sirolimus (Srl) effectively inhibits MUFA synthesis by downregulating SREBP1 and SCD1 activity, thereby restoring the MUFA/PUFA ratio in membrane lipids and rendering tumor cells more susceptible to LPO and ferroptosis.
Results and Discussion
Preparation and characterization of the edge-rich single-Cu-atom SAZyme (ER-Cu1SAZyme)
Copper (Cu) atoms are key cofactors in biological enzymatic reactions due to their redox flexibility and biocompatibility [23]. Additionally, Cu offers a cost-effective alternative to noble metals such as platinum (Pt), palladium (Pd), and gold (Au), making it an attractive candidate for single-atom nanozyme design [24]. To optimize Cu catalytic efficiency within the TME, a novel hollow porous structure was engineered to serve as the substrate for the nanozyme, providing abundant edge sites to anchor Cu catalytic centers. The synthesis of ER-Cu1SAZyme follows a three-step process, as illustrated in Figure 1A. First, mesoporous silica (SiO2) was synthesized using the sol-gel method as a hard template. Dopamine and a copper precursor were introduced to initiate in situ polymerization, forming a continuous polydopamine (PDA) coating on the SiO2 surface. Following high-temperature pyrolysis, the PDA coating converted into a nitrogen-doped carbon (CN) framework with atomically dispersed Cu species. Finally, the SiO2 core was selectively removed using NaOH, resulting in a hollow structure with a porous CN shell. This architecture significantly increases the surface area and exposes edge sites, providing an ideal scaffold for stabilizing Cu single atoms.
Synthesis and structural characterizations of the ER-Cu1SAZyme. (A) Illustration of the preparation process of ER-Cu1SAZyme via a thermal anchoring strategy. (B-C) TEM images of ER-Cu1SAZyme. (D-E) Atomic-resolution HAADF-STEM image and corresponding intensity maps with color profile. (F) HAADF-STEM image. (G) Corresponding energy-dispersive X-ray elemental mapping, indicating the homogeneous distribution of copper (Cu), nitrogen (N), and carbon (C) in ER-Cu1SAZyme.
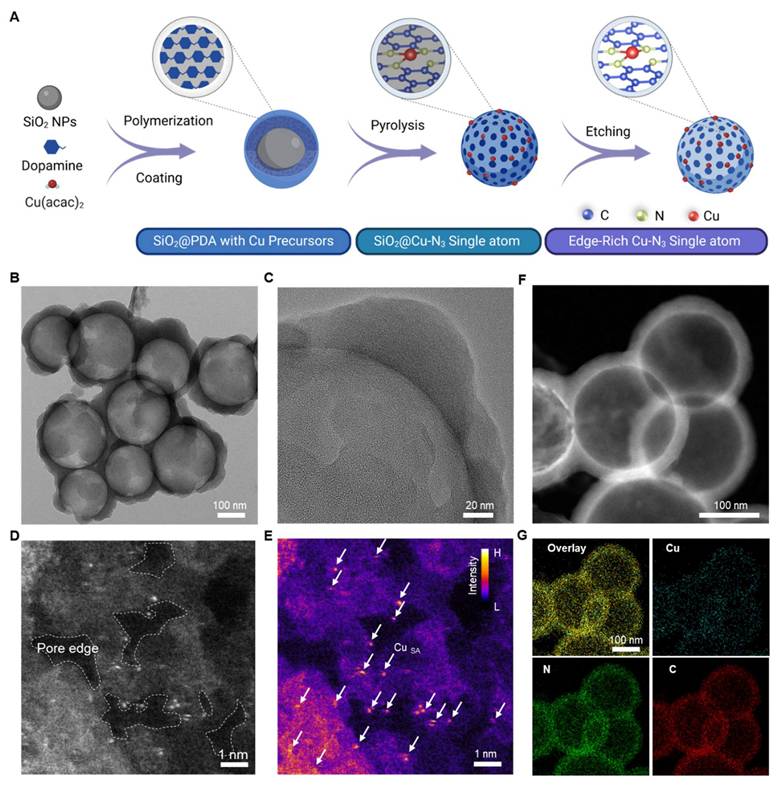
The ER-Cu1SAZyme was extensively characterized using multiple techniques. Transmission electron microscopy (TEM) revealed a well-defined hollow spherical morphology with an average diameter of approximately 250 nm (Figures 1B and S1), and a porous shell thickness ranging from 20 to 40 nm (Figure 1C). To further confirm the hollow and porous structure of ER-Cu1SAZyme, Brunauer-Emmett-Teller (BET) surface area analysis was conducted. The N2 adsorption-desorption isotherms and corresponding pore-size distribution are shown (Figure S2). These results reveal a mesoporous structure with a high surface area of 1376 m2 g-1, complementing the TEM and HAADF-STEM observations. This design promotes the uniform dispersion and stable coordination of single Cu atoms, particularly at the edge sites, ensuring maximal surface exposure. The increased surface area and enhanced accessibility of active sites contribute to the improved catalytic efficiency of the SAZyme. Atomic-resolution high-angle annular dark-field scanning transmission electron microscopy (HAADF-STEM) provided detailed insight into the distribution of Cu atoms on the ER-Cu1SAZyme. Discrete Cu atoms were clearly observed as bright spots, highlighted within white circles (Figure 1D). In contrast to the widely reported in-plane single-atom catalysts with Cu-N4 atomic structures, the synthesized ER-Cu1SAZyme features a hollow CN framework with abundant pores, with the active Cu species predominantly located at the edges of these pores. Accordingly, color-contrast images further highlight the edge-localized dispersion of single Cu atoms on the porous CN framework, clearly verifying their effective dispersion throughout the structure (Figure 1E). Additionally, inductively coupled plasma mass spectrometry (ICP-MS) analysis revealed that the Cu content in ER-Cu1SAZyme is approximately 0.6% (w/w), further confirming the presence of copper species (Table S1). The PEGylated nanomedicine exhibited a hydrated particle size of 300 nm with a zeta potential of +6 mV (Figure S3). The particle size remained stable for over 24 hours, with only a slight increase observed at 48 hours and stabilization by 72 hours, indicating good colloidal stability and excellent dispersibility, making it suitable for in vivo applications (Figure 2F).
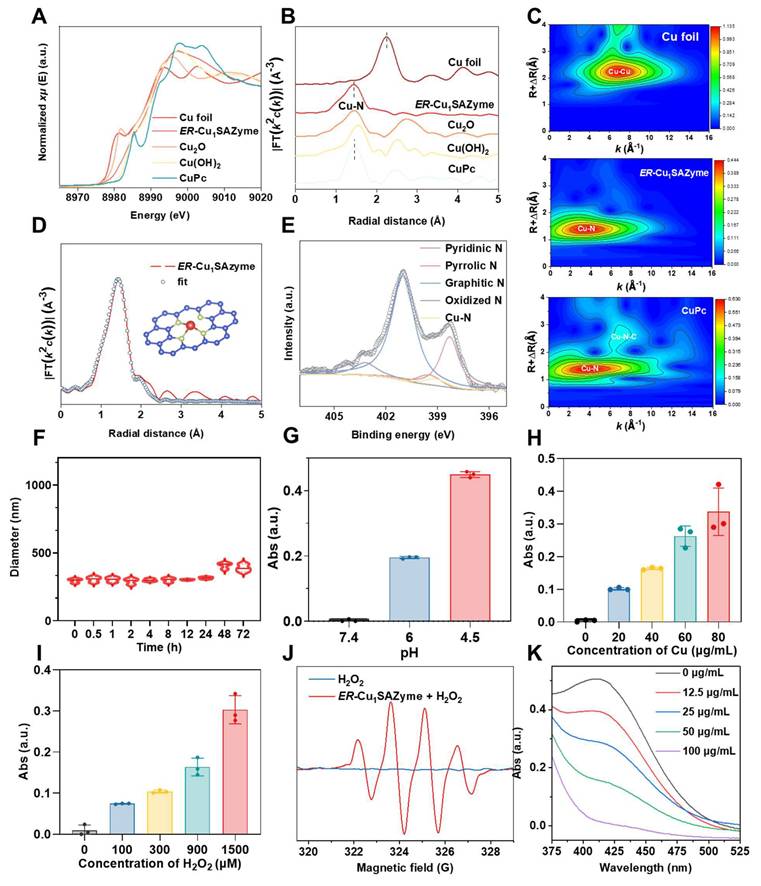
Atomic structural and catalytic characterizations of ER-Cu1SAZyme. (A) Cu K-edge XANES spectra of ER-Cu1SAZyme and reference materials. (B) Fourier-transformed EXAFS signal of ER-Cu1SAZyme and references. (C) EXAFS wavelet-transformed curves of Cu foil, ER-Cu1SAZyme, and CuPc. (D) EXAFS fitting of ER-Cu1SAZyme. (E) N 1s XPS spectra of ER-Cu1SAZyme. (F) Stability analysis of the nanomedicine. (G-I) POD-like activity of ER-Cu1SAZyme in TMB chromogenic reaction at pH 7.4, 6.0, and 4.5 (G); varying ER-Cu1SAZyme concentrations from 0 to 80 μg/mL (H); and varying H2O2 concentrations from 0 to 1500 μM (I). (J) ESR spectra for •OH detection using DMPO as a trapping agent. (K) GSH depletion activity of ER-Cu1SAZyme via the DTNB reduction assay.
Atomic coordination characterizations of the ER-Cu1SAZyme
Synchrotron-based X-ray absorption spectroscopy (XAS) was employed to investigate the chemical state and local atomic environment of Cu in ER-Cu1SAZyme. The Cu K-edge X-ray absorption near-edge structure (XANES) spectra revealed that the absorption edge of ER-Cu1SAZyme shifted to higher energy compared to Cu foil, indicating that the Cu in ER-Cu1SAZyme is in an oxidized state (Figure 2A). By comparing with reference materials such as Cu foil, Cu2O, Cu(OH)2, and CuPc, the average Cu valence in ER-Cu1SAZyme was found to lie between +1 and +2, as the absorption edge of ER-Cu1SAZyme falls between those of Cu2O and Cu(OH)2 (Figure S4) [25]. Fourier transform extended X-ray absorption fine structure (EXAFS) analysis provided further insights into the local coordination of Cu. Unlike Cu foil, which exhibited a characteristic Cu-Cu peak around 2.2 Å in R-space, ER-Cu1SAZyme displayed a prominent Cu-N peak at approximately 1.4 Å, confirming the isolated nature of Cu atoms and their predominant Cu-N coordination (Figures 2B and S5). Additionally, wavelet transform (WT) EXAFS analysis revealed strong Cu-N coordination at ~1.4 Å in both ER-Cu1SAZyme and CuPc, whereas Cu foil showed strong Cu-Cu coordination at ~2.2 Å (Figure 2C). These results further supported the isolated atomic dispersion of Cu in ER-Cu1SAZyme.
EXAFS fitting suggested a Cu-N coordination number of 3.12 for ER-Cu1SAZyme (Figure 2D and Table S2), confirming the presence of a Cu-N3 structure in the first coordination shell. Furthermore, X-ray photoelectron spectroscopy (XPS) validated the interaction between Cu, N, and C. The N 1s spectrum showed characteristic peaks corresponding to pyridinic N at ~398.5 eV, Cu-N coordination at ~399.1 eV, pyrrolic N at ~399.6 eV, graphitic N at ~401.1 eV, and oxidized N at ~403.1 eV, consistent with previously reported Cu-N-C single-atom systems (Figure 2E) [26, 27]. The presence of pyridinic and pyrrolic nitrogen confirmed that isolated Cu is coordinated with approximately three pyridinic N atoms, predominantly forming the Cu-N3 structure in ER-Cu1SAZyme.
Despite the low Cu content, characteristic peaks for Cu 2p3/2 and Cu 2p1/2 were detected at 934.7 eV and 954.6 eV, respectively, corresponding to Cu2+ species, while peaks at 932.7 eV and 952.4 eV were attributed to Cu+ species. A satellite peak at 944.4 eV further confirmed the oxidation state of copper in ER-Cu1SAZyme (Figure S6) [28]. The Cu content determined by XPS was 1.15% by mass, which is higher than the value obtained from energy-dispersive X-ray spectroscopy (EDS) analysis (0.73%), indicating that Cu species are more enriched on the surface of the hollow NC structure (Table S1). This observation aligns with the surface-sensitive nature of XPS, which captures surface-loaded species with greater precision than bulk analysis techniques like EDS [6]. Additionally, graphitic and oxidized nitrogen enhance the conductivity and catalytic stability of the nanomaterial. The C 1s spectrum showed three distinct peaks at 290.0 eV, 287.9 eV, 286.1 eV, and 284.8 eV, corresponding to C=O, C-O, C-N, and C=C bonds, respectively (Figure S7) [29]. These results validate the successful fabrication of ER-Cu1SAZyme, with atomically dispersed Cu sites supported by the CN framework, a configuration expected to yield excellent catalytic performance based on its structural and compositional attributes.
To improve the dispersibility and biocompatibility of ER-Cu1SAZyme under physiological conditions, nanoparticles were coated with 1,2-distearoyl-sn-glycero-3-phosphoethanolamine-polyethylene glycol (DSPE-PEG). In this process, the hydrophobic DSPE segment preferentially adsorbs onto the nanozyme surface, while the hydrophilic PEG chains extend outward into the aqueous environment, forming a monolayer on the nanoparticle surface [30-32]. Simultaneously, Srl was loaded via hydrophobic interactions, forming a stable therapeutic nanomedicine (Srl/ER-Cu1SAZyme). As the feeding ratio of Srl increased, the amount of drug loaded onto ER-Cu1SAZyme increased accordingly, though encapsulation efficiency began to decline once the ratio exceeded 10:1. Therefore, a 10:1 feeding ratio was selected for subsequent experiments (Figure S8). The PEGylated nanomedicine remained stable in physiological mimicking conditions for over 24 hours, with only a slight increase in size observed after 48 hours, stabilizing at 72 hours, indicating excellent dispersibility for potential in vivo applications (Figure 2F).
The catalytic performance of ER-Cu1SAZyme
The catalytic activity of ER-Cu1SAZyme was evaluated through its peroxidase (POD)-like activity using the 3,3',5,5'-tetramethylbenzidine (TMB) colorimetric assay. ER-Cu1SAZyme effectively catalyzed the conversion of H2O2 into highly reactive hydroxyl radicals (•OH), leading to the rapid oxidation of the TMB reagent to its blue oxidized form, which exhibited a characteristic absorption peak at 652 nm, correlating with concentration. As shown in Figure 2G, no significant absorbance signal was observed at pH 7.4, but a notable increase in absorbance occurred at lower pH levels, indicating the ability of the SAZyme to catalyze ROS production specifically within the acidic TME. To further assess catalytic performance under TME-simulating conditions, the catalytic activity was closely monitored at pH 6. Figure 2H illustrates a gradual increase in absorbance with higher concentrations of ER-Cu1SAZyme, suggesting that elevated SAZyme concentrations enhance the catalytic reaction. Similar concentration-dependent results were observed with the TMB (Figure S9). Moreover, as the concentration of H2O2 increased, absorbance also increased, demonstrating that the ROS generation reaction is more effective in the TME, where H2O2 levels are typically higher compared to normal tissues, thus enhancing tumor-targeting capabilities (Figure 2I).
Kinetic analyses were conducted using both the Michaelis-Menten and Lineweaver-Burk models. The calculated Km and Vmax values were 2.06 mM and 0.13 μM·s-1 for the Michaelis-Menten model, and 1.92 mM and 0.13 μM·s-1 for the Lineweaver-Burk model, respectively (Figures S10-11). These results indicate that ER-Cu1SAZyme exhibits significantly higher catalytic activity than a planar non-edge Cu-N4 single-atom counterpart, which was prepared by lowering the pyrolysis temperature to suppress edge-site formation. Despite having a similar elemental composition, this non-edge control sample lacks abundant low-coordinated edge sites, resulting in a planar coordination environment (Figures S12-13). Furthermore, ER-Cu1SAZyme demonstrates superior performance compared to several previously reported Cu-based single-atom nanozymes [33-36], confirming the effectiveness of the edge-enriched structural design. Additionally, the catalase (CAT), superoxide dismutase (SOD), and glutathione peroxidase (GPX)-like activities of ER-Cu1SAZyme were examined to confirm its selective catalytic properties. Results showed only weak CAT-like activity, while SOD- and GPX-like activities were barely measurable (Figures S14-S16). This selective catalytic behavior ensures stable and predictable catalytic performance, which is advantageous for controlled ROS generation and maintaining therapeutic precision in tumor treatment.
To identify the ROS generated by ER-Cu1SAZyme, electron spin resonance (ESR) analysis was conducted using 5,5-dimethyl-1-pyrroline N-oxide (DMPO) as a spin-trapping agent. The resulting 1:2:2:1 peak pattern confirmed the presence of •OH, indicating that ER-Cu1SAZyme acts as a POD mimic by catalyzing the conversion of H2O2 into •OH in tumor tissue (Figure 2J). Furthermore, the GSH-depleting activity of ER-Cu1SAZyme was assessed using the 5,5'-dithiobis (2-nitrobenzoic acid) (DTNB) reduction assay. The decrease in absorbance of the yellow DTNB indicates GSH oxidation in a reaction catalyzed by ER-Cu1SAZyme (Figure 2K). These findings highlight the exceptional ability of ER-Cu1SAZyme to generate highly reactive •OH radicals while simultaneously depleting key endogenous antioxidants like GSH. This dual action facilitates lipid oxidation in cellular membranes, making ER-Cu1SAZyme a potent agent for inducing oxidative stress in tumor cells.
The catalytic performance of ER-Cu1SAZyme before and after DSPE-PEG modification was also evaluated. The modified ER-Cu1SAZyme retained its efficient POD-like catalytic activity under physiological conditions, with only a slight decrease in activity compared to the unmodified nanozyme. This reduction may be due to partial blockage of catalytic sites or limited substrate diffusion (Figure S17).
DFT studies on the activity of ER-Cu1SAZyme
To gain deeper insight into the remarkable catalytic performance of ER-Cu1SAZyme in generating •OH radicals, density functional theory (DFT) calculations were performed to investigate the conversion process of H2O2 during the POD-like catalytic reaction. Initially, the coordination structure of ER-Cu1SAZyme was confirmed through EXAFS data fitting, revealing that Cu atoms are coordinated with an average of 3.12 nitrogen atoms at a path distance of about 1.92 Å (Table S2). This confirms that the primary coordination structure of ER-Cu1SAZyme is Cu-N3 (Figures 3A and S18). Following this, two catalyst models were constructed for analysis: ER-Cu1SAZyme and a control model lacking the Cu active center, referred to as NC. The density of states (DOS) analysis revealed distinct differences between the CN substrate and the Cu-containing model (Figure 3B). In the CN substrate without Cu, the DOS is primarily influenced by C states, with minimal contributions from N states. In contrast, the inclusion of Cu significantly alters the DOS, particularly in the energy range near the Fermi level (0 eV). The Cu atom introduces states around -3 eV, -2 eV, and just below the Fermi level, indicating strong interactions between the Cu atom and the surrounding CN framework. These interactions enhance the electron transport capacity of the SAZyme. Additionally, the increased complexity of the DOS near the Fermi level suggests that the presence of Cu creates new electronic states and modifies the distribution of existing ones, which is essential for the efficient decomposition of H2O2 during POD-like reactions.
DFT studies on the activity of ER-Cu1SAZyme. (A) The coordination structure of ER-Cu1SAZyme. (B) The density of states of CN and ER-Cu1SAZyme. (C) Adsorption energy contrast for H2O2 molecule adsorption on NC and ER-Cu1SAZyme. (D) Energy profile diagram illustrating the possible reaction paths for H2O2 activation over the catalyst.
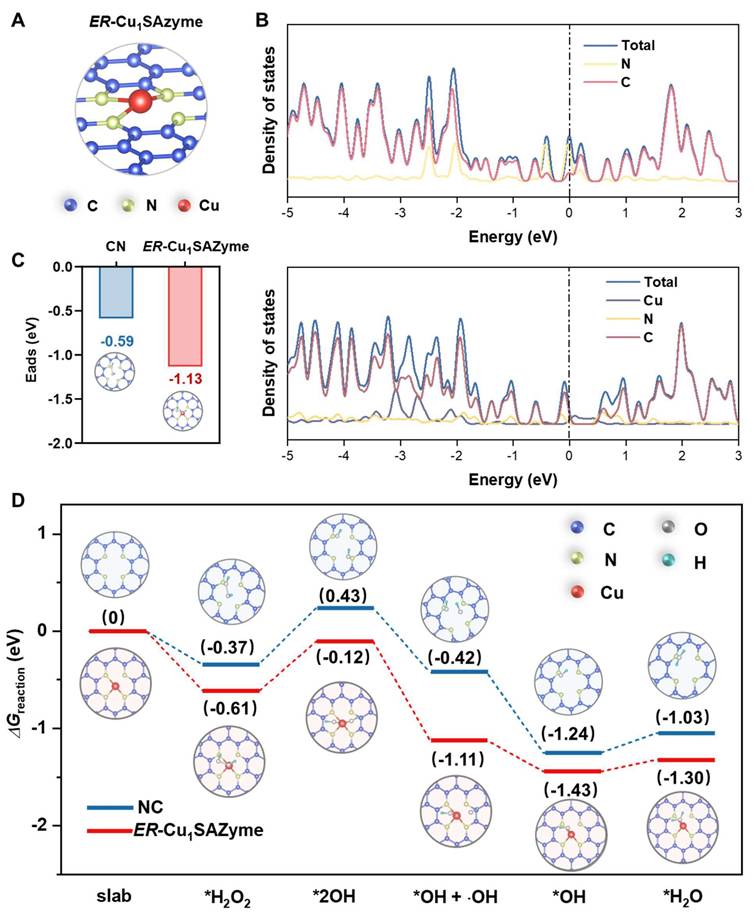
The catalytic performance of ER-Cu1SAZyme is further supported by its stronger interaction with H2O2 compared to the CN substrate, as evidenced by the lower adsorption energy (-1.13 eV vs. -0.59 eV). This indicates that ER-Cu1SAZyme forms a more stable bond with H2O2, which enhances its catalytic activity. The energy profile (Figure 3D) outlines the entire catalytic process, beginning with the adsorption of H2O2 onto the catalyst. ER-Cu1SAZyme demonstrates lower reaction free energy, reflecting a higher adsorption affinity that facilitates subsequent steps. The adsorbed H2O2 dissociates into two *OH intermediates, with one desorbing to release an •OH radical. The remaining *OH reacts with hydrogen to form *H2O, and finally, the catalyst returns to its original state after H2O desorption. Throughout this process, ER-Cu1SAZyme consistently exhibits lower free energy than the CN substrate, confirming that the inclusion of Cu active centers significantly enhances the reaction efficiency, making ER-Cu1SAZyme an effective POD catalyst.
The catalytic performance of Srl/ER-Cu1SAZyme in tumor cells
Oral squamous cell carcinoma (OSCC) is an aggressive malignancy associated with dysregulated lipid metabolism, which drives tumor progression and therapeutic resistance [37, 38]. In this study, OSCC Cal-27 cells were used to evaluate the therapeutic efficacy of Srl/ER-Cu1SAZyme. To assess the uptake efficiency of the SAZymes, red fluorescent-labeled ER-Cu1SAZyme (RF/ER-Cu1SAZyme) was incubated with the cells. The results showed that RF/ER-Cu1SAZyme was effectively internalized by Cal-27 cells in a time-dependent manner, laying the groundwork for subsequent therapeutic applications (Figures 4A and S19).
In vitro catalytic performance of ER-Cu1SAZyme. (A) Confocal microscopy images of Cal-27 cells after incubation with RF/ER-Cu1SAZyme (red: RF/ER-Cu1SAZyme, blue: Hoechst). (B) Cell viability of Cal-27 cells after treatment with Srl/ER-Cu1SAZyme for 48 hours. (C) Live/dead cell assay showing co-staining with Calcein-AM (green for live cells) and PI (red for dead cells). (D-E) Analysis of ROS levels in Cal-27 cells using DCFH-DA staining by flow cytometry (D) and confocal laser scanning microscopy (CLSM) (E). (F) Intracellular GSH levels in Cal-27 cells after incubation with Srl/ER-Cu1SAZyme. (G) Intracellular MDA levels in Cal-27 cells after incubation with Srl/ER-Cu1SAZyme. (H) CLSM images analyzing mitochondrial depolarization using a JC-1 kit. Data are presented as mean ± s.d., with all measurements (n) being biologically independent. Two-group comparisons were performed using a two-tailed unpaired Student's t-test, and multiple comparisons were analyzed by one-way ANOVA with Tukey's post hoc test.
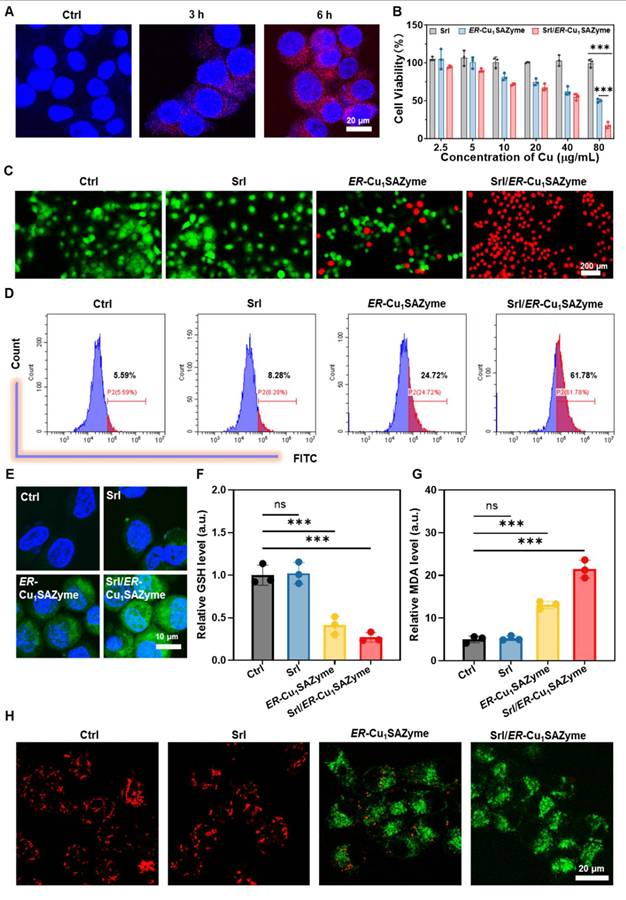
Following confirmation of cellular uptake, the cytotoxic effects of Srl/ER-Cu1SAZyme on Cal-27 cells were assessed using a CCK-8 assay. Treatment with ER-Cu1SAZyme significantly decreased the viability of Cal-27 cells, and the reduction was further pronounced when combined with Srl (Srl/ER-Cu1SAZyme) (Figure 4B). At a concentration of 80 μg/mL, the survival rate of cells treated with ER-Cu1SAZyme was approximately 50%, while the Srl/ER-Cu1SAZyme group exhibited a dramatic decrease in survival, with only 17% cell viability. Notably, Srl alone at equivalent concentrations did not induce significant cytotoxicity, clearly indicating the synergistic effect of ER-Cu1SAZyme in combination with Srl. This synergistic effect was further validated by live/dead cell staining analysis. As shown in Figure 4C, co-staining with Calcein-AM (green fluorescence for live cells) and PI (red fluorescence for dead cells) revealed a substantial increase in dead cells in the ER-Cu1SAZyme-treated group, while the Srl/ER-Cu1SAZyme group displayed almost no green fluorescence, indicating near-complete cell death. Since dying cells lose their spread morphology and become more rounded, they may appear slightly smaller under identical magnification. These results demonstrate that Srl/ER-Cu1SAZyme significantly enhances antitumor efficacy, offering a promising strategy for therapeutic interventions.
To investigate whether the therapeutic effects of Srl/ER-Cu1SAZyme are associated with ROS-mediated cell damage, intracellular ROS levels were measured. Using the green fluorescent probe 2,7-dichlorofluorescein diacetate (DCFH-DA), a significant increase in ROS levels was observed following ER-Cu1SAZyme treatment. Specifically, ROS levels in Cal-27 cells increased from 5.59% to 61.78%, demonstrating that Srl substantially enhanced the catalytic activity of ER-Cu1SAZyme in tumor cells (Figure 4D). This increase in ROS was also confirmed by stronger fluorescence signals observed in confocal microscopy after treatment with ER-Cu1SAZyme and Srl/ER-Cu1SAZyme (Figure 4E). Additionally, the role of Srl/ER-Cu1SAZyme in boosting ROS generation was further explored by measuring intracellular GSH levels. While Srl alone had no significant effect on GSH levels, ER-Cu1SAZyme reduced GSH levels to approximately 40%, and in the Srl/ER-Cu1SAZyme group, GSH dropped further to about 27% (Figure 4F). This confirms that ER-Cu1SAZyme exhibits potent catalytic activity, effectively depleting GSH and significantly amplifying oxidative stress in cancer cells with the assistance of Srl. Furthermore, malondialdehyde (MDA), a key marker of oxidative stress, was measured. MDA levels in the ER-Cu1SAZyme group were 2.6 times higher than in the control group, and this increased further to 4.3 times in the Srl/ER-Cu1SAZyme group (Figure 4G). These results highlight the exceptional catalytic performance of Srl/ER-Cu1SAZyme in amplifying oxidative stress within cancer cells, thereby enhancing its potent therapeutic effect.
After evaluating ROS levels, the effect of Srl/ER-Cu1SAZyme on mitochondria was investigated. Mitochondria are not only the primary source of ROS in cells but also major targets of ROS-induced damage, which can lead to ferroptosis in cancer cells [39]. Mitochondrial damage was assessed by measuring the mitochondrial membrane potential (MMP) using the JC-1 fluorescent probe. In healthy cells, JC-1 aggregates in the mitochondrial matrix, emitting red fluorescence. However, in damaged cells with depolarized mitochondrial membranes, JC-1 remains in its monomeric form, emitting green fluorescence [40]. Following treatment with Srl/ER-Cu1SAZyme, a significant increase in green fluorescence and a reduction in red fluorescence were observed, indicating severe mitochondrial damage caused by ROS generated by the nanomedicine (Figures 4H and S20). These results suggest that Srl/ER-Cu1SAZyme effectively depletes GSH and generates ROS, inducing oxidative stress that results in mitochondrial dysfunction. Consequently, Srl/ER-Cu1SAZyme-induced cancer cell death appears to be linked to ROS-mediated oxidative damage and ferroptosis.
Srl/ER-Cu1SAZyme induces robust ferroptosis in tumor cells
Building on the observed catalytic performance of Srl/ER-Cu1SAZyme, its capacity to induce ROS-mediated ferroptosis was further assessed by examining key markers such as GPX4 expression and LPO levels. GPX4 plays a critical role in regulating cellular redox homeostasis and is directly associated with ferroptosis. Since GSH is essential for GPX4 synthesis, its depletion naturally leads to reduced GPX4 levels [41]. As anticipated, ER-Cu1SAZyme significantly reduced GPX4 expression to 40%, and treatment with Srl/ER-Cu1SAZyme further decreased it to 20%, reflecting a substantial disruption of cellular redox balance induced by this nanoformulation (Figures 5A, B, and S21). LPO, a key hallmark of ferroptosis, was also assessed. Intracellular MDA levels, a terminal product of LPO, were significantly elevated in Srl/ER-Cu1SAZyme-treated cells, indicating heightened oxidative stress. This increase in MDA suggests considerable LPO in the cell membrane, consistent with ferroptotic cell death [42]. To directly measure LPO, the BODIPY-C11 fluorophore was used. This probe undergoes oxidation, resulting in a fluorescence shift from red (reduced state) to green (oxidized state), making it widely applicable in ferroptosis and oxidative stress studies [43, 44]. A distinct green fluorescence was observed in Srl/ER-Cu1SAZyme-treated cells, confirming high levels of LPO (Figures 5C and S22). These results validate that Srl/ER-Cu1SAZyme-induced cancer cell death is closely linked to ferroptosis, driven by oxidative stress and membrane LPO.
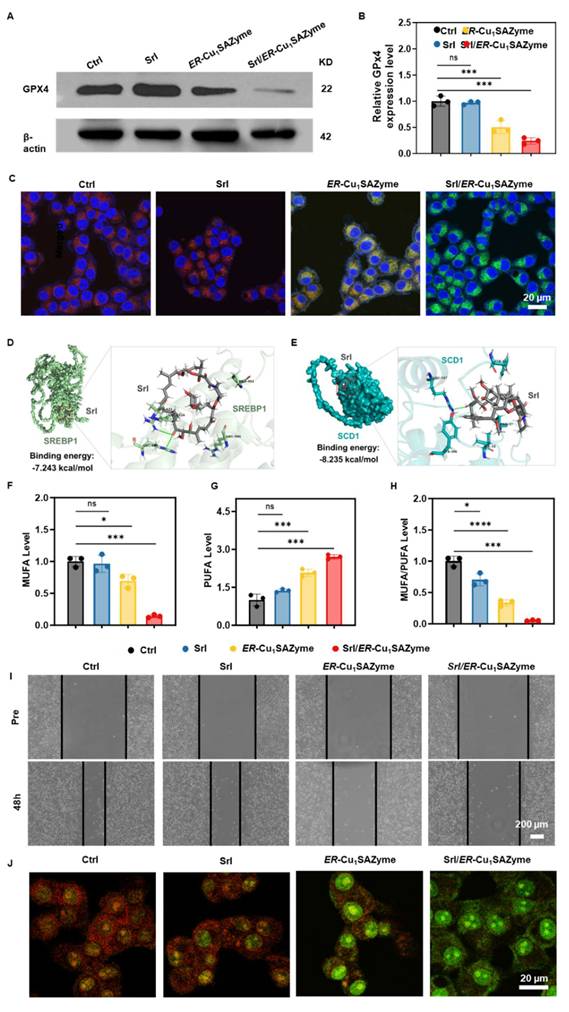
Potential antitumor mechanism of ER-Cu1SAZyme. (A) Western blot analysis of GPX4 expression in Cal-27 cells treated with Srl/ER-Cu1SAZyme. (B) Quantification of GPX4 protein levels normalized to untreated controls, with β-actin as a loading control. Band intensities were analyzed using ImageJ software and presented as fold change relative to control. (C) Lipid peroxidation levels measured by BODIPY-C11 staining in Cal-27 cells after Srl/ER-Cu1SAZyme treatment. Red fluorescence indicates non-oxidized lipids, while green fluorescence represents oxidized lipids. (D-E) Molecular docking schematic: Structural formulas of Srl, SREBP1 (D), and SCD1 (E), with close-up views of the interaction sites. (F-H) Intracellular levels of MUFA (F), PUFA (G), and the MUFA/PUFA ratio (H) in Cal-27 cells following Srl/ER-Cu1SAZyme treatment. (I) Representative images of Cal-27 cell migration at 48 hours after Srl/ER-Cu1SAZyme treatment. (J) CLSM images of Cal-27 cells stained with AO after Srl/ER-Cu1SAZyme treatment. Red fluorescence indicates intact lysosomes, while green fluorescence reflects damaged lysosomes. Data are presented as mean ± s.d. All measurements (n) are biologically independent.
To further confirm the catalytic LPO activity of ER-Cu1SAZyme, a model liposome system was established. As shown in Figure S23, untreated liposomes and those treated with ER-Cu1SAZyme or H2O2 alone emitted only red fluorescence around 590 nm, corresponding to the reduced form of the liposome. In contrast, the ER-Cu1SAZyme plus H2O2 group displayed a marked increase in green fluorescence, indicating that ER-Cu1SAZyme effectively catalyzes LPO in the liposome model. This supports its role in promoting membrane lipid oxidation during ferroptosis.
To elucidate the underlying mechanism, the regulation of lipid metabolism by Srl/ER-Cu1SAZyme was assessed, as the cell membrane, a primary target of LPO, plays a pivotal role in determining sensitivity to LPO [13]. SREBP1, a transcription factor involved in lipid metabolism, is regulated by mTOR and drives the expression of SCD1 [45-47]. SCD1, in turn, promotes the synthesis of MUFAs, which incorporate into the cell membrane, replacing PUFAs [48]. This shift reduces LPO and enhances resistance to ferroptosis [49]. To evaluate the effect of Srl on modulating membrane composition, molecular docking studies were conducted between Srl and key lipid metabolic proteins. As previously reported, Srl exhibits high affinity for mTOR [50], with a binding energy of -8.442 kcal/mol, indicating potent inhibition (Figure S24 and Tables S3-S4). Additionally, Srl showed strong binding affinity for SREBP1 and SCD1, with binding energies of -7.243 and -8.235 kcal/mol, respectively (Figures 5D, E and S25-26). The interacting residues between Srl and SREBP1 or SCD1 are listed in Tables S5-S6. These interactions suggest that Srl may interfere with MUFA content in the cell membrane. To further explore this, ELISA was used to measure MUFA levels in cells. The results demonstrated significant inhibition of MUFA synthesis in the Srl/ER-Cu1SAZyme group, highlighting the nanomedicine's effective therapeutic action (Figure 5F). In contrast, free Srl exhibited limited efficacy, likely due to its hydrophobic nature, which hinders bioavailability at low concentrations in physiological environments. Conversely, the Srl/ER-Cu1SAZyme nanomedicine enhanced cellular uptake, improving its therapeutic effectiveness. Furthermore, an unexpected increase in PUFA levels was observed after treatment with Srl/ER-Cu1SAZyme (Figure 5G). This could result from the reduction in MUFA and increased oxidative stress, which likely activates a compensatory mechanism to elevate PUFA levels, thereby helping cells mitigate the enhanced oxidative stress [51, 52]. This observation highlights the remarkable plasticity and survival capacity of cancer cells.
High-performance liquid chromatography-tandem mass spectrometry (HPLC-MS/MS) was employed to further analyze the lipid composition of the cell membrane. Five major membrane-associated lipid classes were examined: phosphatidylcholine (PC), phosphatidylethanolamine (PE), phosphatidylserine (PS), cholesteryl ester (CE), and sphingomyelin (SM), all of which are essential for membrane integrity and regulate ferroptosis sensitivity [53-57]. For each lipid class, the relative abundance of MUFA- and PUFA-containing species was quantified, and the MUFA/PUFA ratio was calculated. As shown in Figure S27, treatment with Srl/ER-Cu1SAZyme led to a reduction in MUFA/PUFA ratios across multiple lipid classes. Similar findings were confirmed by the ELISA assay (Figure 5H), indicating that Srl/ER-Cu1SAZyme modulates lipid metabolism, promotes LPO, and enhances ferroptosis.
Given the multifaceted nature of tumor cell death, the observed lipid-centered metabolic changes, along with significant LPO and multiple ferroptosis-associated molecular markers, strongly suggest that ferroptosis plays a critical role in the cellular response induced by Srl/ER-Cu1SAZyme. Apoptosis may also contribute, albeit to a lesser extent, as indicated by the partial rescue effect observed with the apoptosis inhibitor Z-VAD-FMK (Figure S28). In contrast, ferroptosis inhibition using Ferrostatin-1 provided a much stronger protective effect, further emphasizing ferroptosis as the dominant mechanism underlying the observed cytotoxicity.
Furthermore, scratch assays revealed that Srl/ER-Cu1SAZyme effectively inhibits cancer cell migration, suggesting that the compensatory increase in PUFA is insufficient to induce metastasis, thus enhancing therapeutic efficacy (Figures 5I and S29). Significant lysosomal damage was also observed. As shown in Figure 5J, Srl alone had minimal impact on lysosomal integrity, but under the strong oxidative stress induced by ER-Cu1SAZyme, a notable increase in lysosomal internal pH was observed, as evidenced by the diminished red fluorescence, indicating disruption of the lysosomal membrane. In the Srl/ER-Cu1SAZyme group, Srl's inhibition of mTOR-mediated lysosomal regeneration, combined with the potent oxidative damage from ER-Cu1SAZyme, exacerbated lysosomal dysfunction, as reflected by the complete loss of red fluorescence. Lysosomal damage impairs the degradation of cellular debris and damaged organelles, leading to the accumulation of toxic intermediates [58, 59]. Therefore, lysosomal destruction disrupts the tumor cells' ability to repair and maintain cellular homeostasis, reducing drug resistance and increasing sensitivity to ferroptosis.
The ferroptosis sensitization mechanism of the Srl/ER-Cu1SAZyme
After confirming that Srl/ER-Cu1SAZyme effectively induces ferroptosis in Cal-27 cells, the intrinsic mechanisms by which Srl/ER-Cu1SAZyme modulates lipid metabolism to enhance ferroptosis susceptibility were further investigated. To better simulate clinical conditions, 30 clinical samples from patients with OSCC were collected (Figure 6A and Table S7). Immunohistochemistry revealed marked overexpression of SCD1 in OSCC tissues from various anatomical sites, including the buccal mucosa, floor of mouth, gingiva, and tongue (Figures 6B and S30-31), alongside moderate increases in GPX4 (Figures S32-33). These results suggest that OSCC cells may resist ferroptosis by activating the MUFA synthesis pathway and enhancing antioxidant defense mechanisms, thus maintaining cell survival and proliferation. Specifically, the overexpression of SCD1 drives the conversion of saturated fatty acids to MUFAs, which reduces LPO mediated by PUFAs, while elevated GPX4 levels further inhibit ferroptosis by scavenging LPO. Moreover, significant overexpression of mTOR, an upstream regulator of SCD1, was observed (Figures 6C and S34-35), likely promoting SCD1 transcription via the activation of SREBP1 and thereby reinforcing a metabolic phenotype that resists ferroptosis.
Abnormalities in SCD1 and its upstream regulators in clinical patient tissue samples. (A) Schematic of clinical cancer tissue collection and analysis. (B) Representative image of SCD1 expression in OSCC tissues from 30 paired clinical samples. (C) Representative image of mTOR expression in OSCC tissues from 30 paired clinical samples. (D) Volcano plot comparing gene expression between Tumor and Control groups, with blue indicating downregulated DEGs and red indicating upregulated DEGs. (E-G) Expression of PIK3CA (E), AKT3 (F), and mTOR (G) across individual cancer stages in HNSCC. (H) DEG enrichment in KEGG pathways. Data are presented as mean ± s.d. P < 0.001, ***, P < 0.01, **, P < 0.05, *.
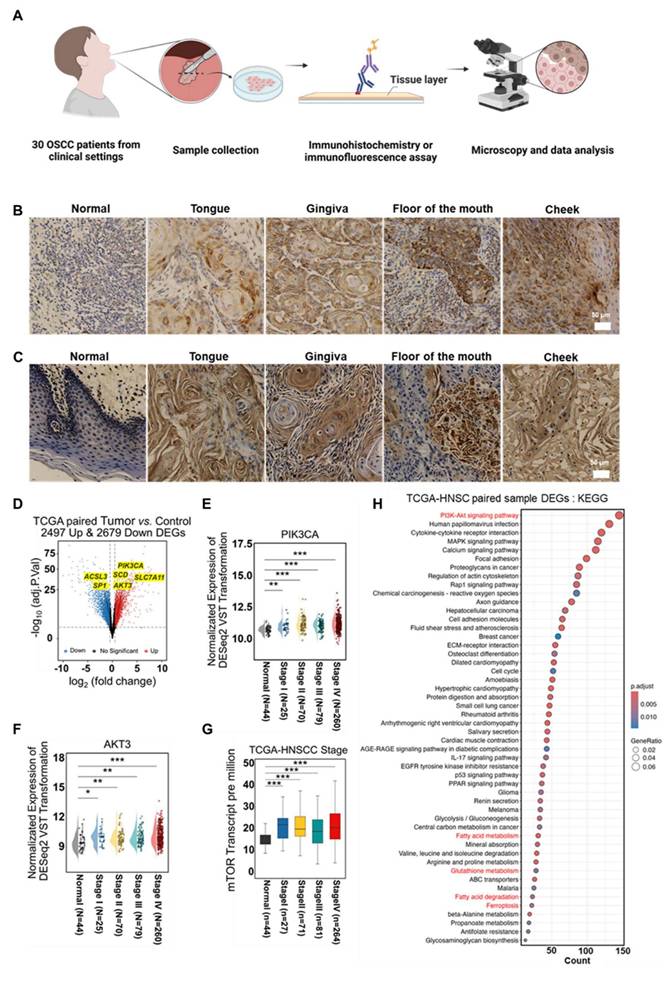
To further validate this hypothesis and enhance the clinical relevance of the findings, a comprehensive analysis was conducted using data from the TCGA (The Cancer Genome Atlas) and CPTAC (Clinical Proteomic Tumor Analysis Consortium) databases. Gene expression differences between normal and tumor tissues were compared systematically. Since OSCC represents the sixth most common cancer globally, accounting for over 90% of head and neck malignancies [60], data from head and neck squamous cell carcinoma (HNSCC) were also included to increase sample size and ensure statistically significant results. Comparative transcriptomics of paired tumor and normal tissues from TCGA revealed 2,497 upregulated genes and 2,679 downregulated genes in tumor tissues (Figure 6D). Key alterations included lipid metabolism-related genes (ACSL3, SCD1), redox homeostasis modulators (SLC7A11), and oncogenic drivers (PIK3CA, AKT3, SP1), suggesting significant rewiring of lipid metabolism, activation of pro-invasive signaling, and disruption of oxidative stress balance. Further analysis revealed a significant upregulation of SCD1 upstream regulators, including PIK3CA, AKT3, and mTOR, across all stages of HNSCC (Figures 6E-G).
Notably, mTOR expression exhibited a progressive increase with age (Figure S36) and differed by sex (Figure S37), with particularly high levels observed in lymph node metastases (Figure S38). Validation using TCGA and CPTAC datasets confirmed significantly higher mTOR expression in primary tumors compared to normal tissues (Figures S39-40). These results suggest that sustained activation of SCD1 upstream genes may contribute to tumor progression by promoting cell proliferation and survival through lipid metabolic reprogramming. Subsequently, differentially expressed genes (DEGs) underwent Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis (Figure 6H), revealing enrichment in pathways related to PI3K-AKT signaling, fatty acid metabolism, GSH metabolism, fatty acid degradation, and ferroptosis. These results provide strong theoretical support for targeting lipid metabolism remodeling as a potential strategy to enhance ferroptosis and improve therapeutic outcomes.
The identification of mutations and altered expression levels of MUFA-related genes in clinical tissue samples of patients with OSCC further substantiates the potential of lipid metabolism modulation to improve therapeutic outcomes in OSCC. Previous cellular data have demonstrated that Srl/ER-Cu1SAZyme effectively reduces the MUFA/PUFA ratio and activates ferroptosis in Cal-27 cells. To investigate the underlying mechanisms and assess the clinical relevance, gene and protein expression levels in Cal-27 cells treated with Srl/ER-Cu1SAZyme were analyzed to identify the molecular factors driving this therapeutic effect.
As shown in Figure 7A, the heatmap reveals significant differences in gene expression between the Srl/ER-Cu1SAZyme-treated and control groups. Principal component analysis (PCA) in Figure 7B confirms clear separation between the gene expression profiles of Srl/ER-Cu1SAZyme-treated and control cells, indicating that treatment significantly alters cellular gene expression. The heatmap (Figure S41) and volcano plot (Figure 7C) show 731 upregulated and 1123 downregulated DEGs in the Srl/ER-Cu1SAZyme-treated group compared to controls. Notably, downregulation of key genes such as FASN, SCD, and FADS2 was observed, all of which are involved in the SREBP1/SCD1-mediated synthesis of MUFAs. This downregulation leads to reduced MUFA levels. Conversely, ACSL4, which facilitates the incorporation of PUFAs into membrane phospholipids, was upregulated, suggesting an increased incorporation of PUFAs into the cell membrane. Combined with reduced MUFA synthesis, this shift lowers the MUFA/PUFA ratio in the membrane, increasing its susceptibility to LPO. These gene alterations impact several biological processes, including metabolism, human diseases, genetic information processing, and cellular functions (Figure 7D).
Gene and protein regulation of Cal-27 cells by ER-Cu1SAZyme. (A) Heat map showing intracellular transcriptomic regulation in Srl/ER-Cu1SAZyme-treated Cal-27 cells. Blue indicates downregulated DEGs, red indicates upregulated DEGs. (B) PCA of transcriptomics sequencing comparing the Control and Srl/ER-Cu1SAZyme groups. (C) Volcano plot comparing gene expression between Control and Srl/ER-Cu1SAZyme groups. Blue indicates downregulated DEGs, red indicates upregulated DEGs. (D) Pathway enrichment analysis of Srl/ER-Cu1SAZyme-treated Cal-27 cells. (E) Top 25 pathways enriched after Srl/ER-Cu1SAZyme treatment. (F-I) qPCR analysis of gene expression in Srl/ER-Cu1SAZyme-treated Cal-27 cells. (J) Western blot analysis of mTOR, p-mTOR, m-SREBP1, SCD1, and FASN expression in Srl/ER-Cu1SAZyme-treated Cal-27 cells. Data are presented as mean ± s.d., and all measurements (n) are biologically independent.
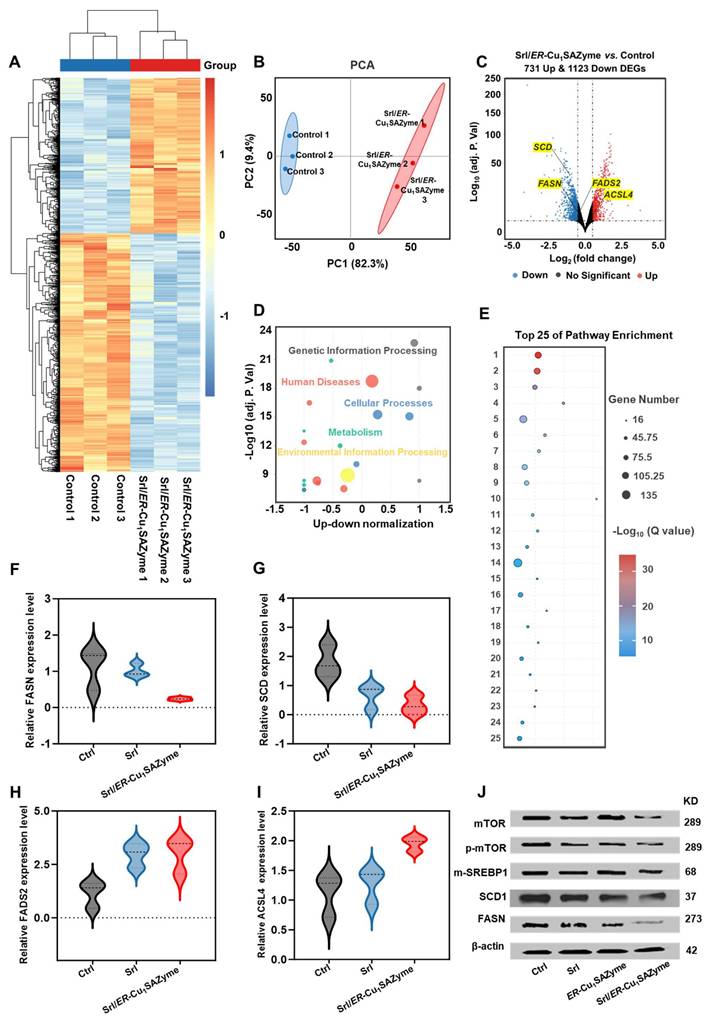
Pathway enrichment analysis further highlighted the involvement of processes related to cancer, cell cycle regulation, DNA repair, oxidative stress, and ferroptosis (Figure 7E), suggesting that Srl/ER-Cu1SAZyme-induced lipid metabolic reprogramming may influence these critical pathways. These findings were corroborated by qPCR analysis (Figures 7F-I, Table S8), which provided additional validation of gene expression changes, supporting the hypothesis that Srl/ER-Cu1SAZyme sensitizes cells to oxidative stress through lipid metabolism alterations. Western blot analysis (Figures 7J, S42, S43) further confirmed these changes at the protein level. Compared to the control group, the Srl/ER-Cu1SAZyme-treated group exhibited significantly reduced protein expression of SCD1 (Figure S44) and FASN (Figure S45), consistent with observed gene expression changes. Additionally, the expression of upstream regulators of SCD1, including mTOR (Figure S46) and SREBP1 (Figure S47), was notably decreased, providing further evidence that Srl/ER-Cu1SAZyme enhances ferroptosis sensitivity by modulating lipid metabolic pathways.
In conclusion, combined Srl/ER-Cu1SAZyme treatment enhances tumor cell sensitivity to ferroptosis by reprogramming lipid metabolism, particularly by modulating the overactive MUFA synthesis and reshaping the MUFA/PUFA ratio in the cell membrane. This effect is achieved through the suppression of SCD1-mediated MUFA synthesis via upstream regulators, such as SREBP1 and mTOR, while simultaneously promoting the incorporation of PUFAs into the membrane by upregulating ACSL4. This reprogramming of the cell membrane lipid composition increases ferroptosis sensitivity, enhancing LPO induced by the catalytic action of Srl/ER-Cu1SAZyme-generated •OH, thereby robustly inducing ferroptosis.
In vivo antitumor efficacy of the Srl/ER-Cu1SAZyme
Following the elucidation of the cancer cell-killing mechanism, the in vivo antitumor efficacy of Srl/ER-Cu1SAZyme was evaluated using a Balb/c nude mouse model bearing Cal-27 tumors. To establish a foundation for assessing the antitumor activity, the biodistribution of the nanotherapeutic agent was first investigated. As shown in Figure 8A, fluorescently labeled ER-Cu1SAZyme began to accumulate in tumor tissues 8 hours post-injection, peaking around 24 hours. Notably, significant amounts of the nanomedicine persisted in the tumor tissue even after 48 hours, indicating prolonged retention that could support sustained therapeutic action. To further confirm the in vivo distribution, ex vivo biodistribution analysis was conducted 48 hours after administration (Figure S48). The fluorescence signal was primarily detected in tumor tissues, with only low-level accumulation in the liver and negligible signals in other major organs, including the heart, spleen, lungs, and kidneys. This distribution pattern aligns with the anticipated clearance behavior of systemically administered nanomaterials and highlights the favorable tumor selectivity of Srl/ER-Cu1SAZyme [61, 62].
Efficient therapeutic efficacy against Cal-27 tumors by ER-Cu1SAZyme. (A) Real-time biodistribution images of fluorescently labeled Srl/ER-Cu1SAZyme before and after injection. (B) Illustration of treatment schedule. This Figure was created on Biorender.com. (C) Tumor growth curves of individual mice in each group (n = 5 biologically independent mice per group). (D-H) Histological microscopic images of dissected tumors stained with H&E (D), Ki67 (E), GPX4 (F), SCD1 (G), and TUNEL (H). Data are presented as mean ± s.d., and all measurements (n) are biologically independent.
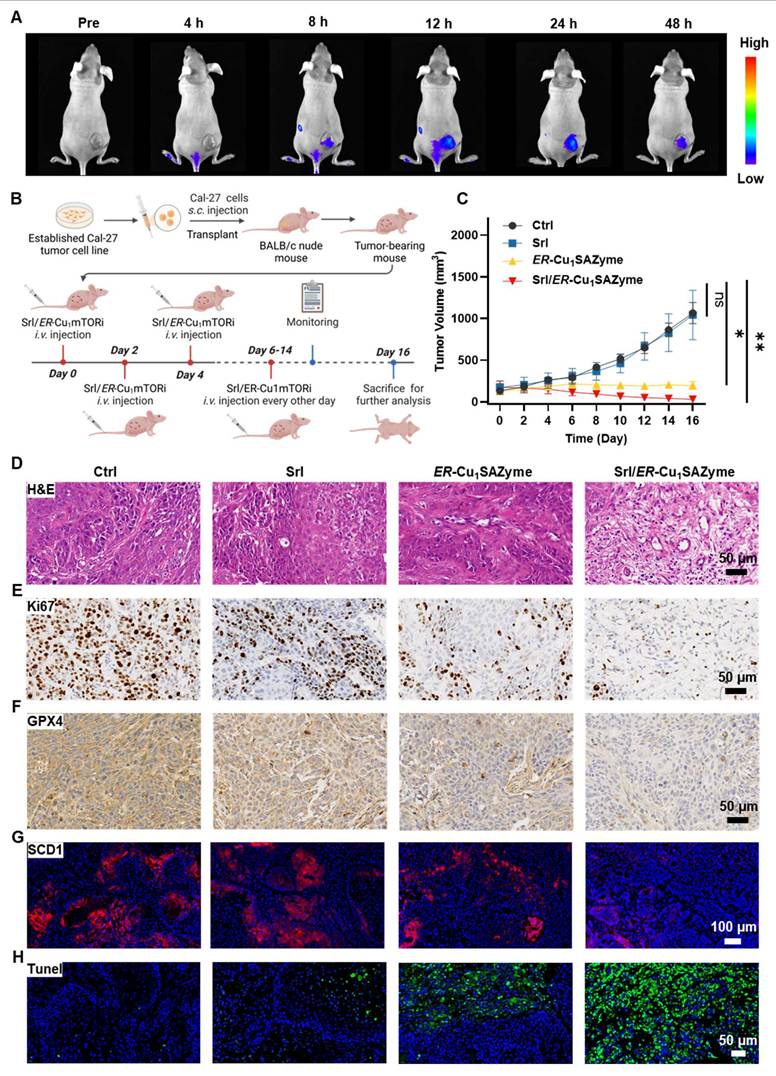
After confirming the accumulation of Srl/ER-Cu1SAZyme in tumors, its antitumor activity was rigorously assessed. Mice were randomly divided into four groups: Saline, Srl, ER-Cu1SAZyme, and Srl/ER-Cu1SAZyme. Treatments were administered via tail vein injections every other day for a total of eight doses to evaluate efficacy (Figure 8B). Tumor growth curves revealed that Srl alone had a negligible effect on tumor progression, while ER-Cu1SAZyme inhibited growth through catalytic damage, though it did not fully halt tumor development. In stark contrast, the combination treatment of Srl/ER-Cu1SAZyme resulted in significant tumor growth suppression, with tumors in this group being approximately 33 times smaller than those in the Saline group and 3.7 times smaller than their initial size (Figures 8C and S49-51). These results suggest that Srl enhances the catalytic therapeutic effect of ER-Cu1SAZyme, highlighting the combination treatment as a promising antitumor strategy.
The enhanced antitumor efficacy of Srl/ER-Cu1SAZyme was further validated through histological analysis. Tumors treated with Srl/ER-Cu1SAZyme exhibited more extensive necrosis and cavitation compared to those treated with ER-Cu1SAZyme alone, consistent with the tumor growth data (Figure 8D). The Ki67 proliferation marker, indicative of tumor cell proliferation, was significantly reduced in the Srl/ER-Cu1SAZyme-treated group, signifying widespread tumor cell death and suppressed proliferation (Figure 8E). To assess the role of ferroptosis, GPX4 expression levels were analyzed (Figure 8F). Both ER-Cu1SAZyme and Srl/ER-Cu1SAZyme treatments downregulated SCD1 expression, with the latter exhibiting a more pronounced effect, further supporting the cellular-level findings. This suggests that the inhibition of SCD1 disrupts MUFA synthesis, thereby sensitizing cells to ferroptosis induced by ROS damage mediated by ER-Cu1SAZyme, demonstrating the therapeutic efficacy in vivo. Notably, Srl/ER-Cu1SAZyme was more effective at reducing GPX4 levels compared to ER-Cu1SAZyme, further confirming the superior therapeutic potential of this ferroptosis-based approach for treating OSCC (Figures 8G and S52).
Additionally, apoptosis also contributed to the therapeutic effect, as confirmed by TUNEL staining. A significant increase in green fluorescent cells was observed in the ER-Cu1SAZyme-treated group, with an even higher number in the Srl/ER-Cu1SAZyme group (Figure 8H). Similar results were observed with Caspase-3 staining, where a significant number of positive cells were detected in the Srl/ER-Cu1SAZyme-treated tumors (Figure S53). These results suggest that the disruption of tumor cell metabolism reduces their adaptability and promotes apoptosis [63]. These results indicate that the Srl/ER-Cu1SAZyme strategy induces a complex and multifaceted tumor cell death process, making it a promising and potent approach for cancer treatment.
The biosafety of the Srl/ER-Cu1SAZyme
The biosafety of Srl/ER-Cu1SAZyme is a key factor for its potential application in nanodrug therapies, necessitating a thorough evaluation of its biocompatibility and safety profile. To assess potential kidney damage, human proximal tubule epithelial (HK-2) cells were co-incubated with Srl/ER-Cu1SAZyme. Results indicated that the viability of HK-2 cells remained largely unaffected, even at high concentrations of 256 μg/ml over a 48-hour exposure period, demonstrating the excellent biosafety of Srl/ER-Cu1SAZyme for normal cells (Figure 9A). To further assess hemocompatibility, hemolysis assays were conducted, as red blood cells are among the first to encounter nanodrugs in circulation (Figure 9B). The results showed no hemolysis at concentrations up to 256 μg/ml, indicating minimal acute hepatotoxicity associated with this nanotherapeutic agent.
In vivo safety of ER-Cu1SAZyme. (A) Cell viability of HK-2 cells treated with Srl/ER-Cu1SAZyme for 24 and 48 hours. (B) Hemolysis test of Srl/ER-Cu1SAZyme at different concentrations. (C-E) Serum biochemistry parameters, including Cr (C), GOT (D), and GPT (E) levels, measured on the 16th day post-administration. (F) Body weight changes in mice after treatment. (G) Histological microscopic images of tissues stained with H&E. Data are presented as mean ± s.d., and all measurements (n) are biologically independent.
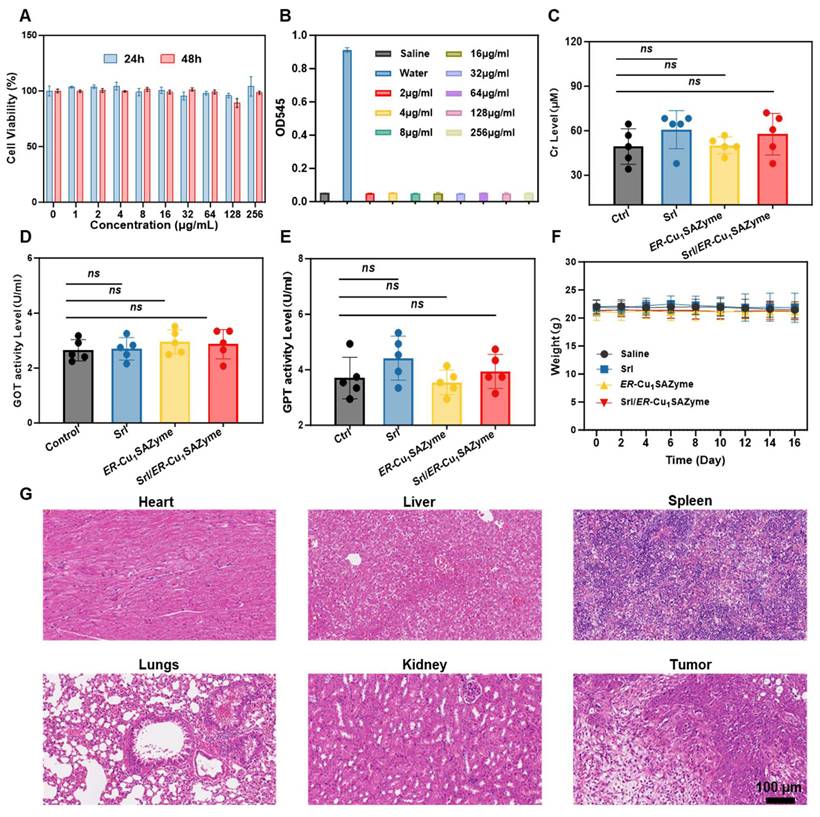
For in vivo hepatorenal safety evaluation, serum samples were collected at the end of treatment, and levels of creatinine, along with liver function markers such as glutamic-pyruvic transaminase (GPT) and glutamic oxaloacetic transaminase (GOT), were measured (Figures 9C-E). All parameters remained within the normal range, suggesting minimal liver and kidney toxicity associated with Srl/ER-Cu1SAZyme administration. Moreover, the steady weight gain observed in treated mice further supported the well-tolerated nature of the treatment (Figure 9F).
Histological analysis of various organs (heart, liver, spleen, lung, and kidney) was performed via H&E staining post-treatment. No significant tissue damage was detected, supporting the favorable in vivo safety profile and efficient clearance behavior of the treatment (Figure 9G). These results strongly suggest that Srl/ER-Cu1SAZyme exhibits excellent biocompatibility and biosafety in vivo.
Conclusions
In this study, a novel edge-rich Cu-N3 single-atom nanozyme (ER-Cu1SAZyme) was developed, characterized by its unique hollow, porous structure and catalytic centers enriched at the edges. This distinct architecture enhances catalytic activity, enabling efficient ROS generation that disrupts cellular redox homeostasis. In combination with the FDA-approved drug Srl, ER-Cu1SAZyme synergistically modulates lipid metabolism in tumor cells by inhibiting SCD1-mediated MUFA synthesis and promoting ACSL4-mediated PUFA activation. This lipid remodeling alters the tumor cell membrane composition, enhancing susceptibility to LPO and ferroptosis. In tumor-bearing murine models, treatment with Srl/ER-Cu1SAZyme resulted in a significant reduction in tumor volume, demonstrating its potent therapeutic efficacy with minimal toxicity. These results highlight the potential of targeting lipid metabolic reprogramming combined with enhanced ROS generation as an effective strategy to overcome tumor resistance and improve ferroptosis-based therapies. This study offers new insights and a strong data foundation for the design and application of novel single-atom nanozymes, paving the way for their future clinical translation in biomedical applications.
Abbreviations
ACSL4: acyl-CoA synthetase long-chain family member 4; BET: Brunauer-Emmett-Teller; CPTAC: Clinical Proteomic Tumor Analysis Consortium; DEGs: differentially expressed genes; DFT: density functional theory; DOS: density of states; ER-Cu1SAZyme: edge-rich Cu-N3 single-atom nanozyme; ESR: electron spin resonance; EXAFS: extended X-ray absorption fine structure; GSH: glutathione; HAADF-STEM: high-angle annular dark-field scanning transmission electron microscopy; HNSCC: head and neck squamous cell carcinoma; H2O2: hydrogen peroxide; ICP-MS: inductively coupled plasma mass spectrometry; KEGG: Kyoto Encyclopedia of Genes and Genomes; LPO: lipid peroxidation; MDA: malondialdehyde; MMP: mitochondrial membrane potential; MUFAs: monounsaturated fatty acids; OSCC: oral squamous cell carcinoma; PCA: principal component analysis; PUFAs: polyunsaturated fatty acids; ROS: reactive oxygen species; SAZymes: single-atom nanozymes; SCD1: stearoyl-CoA desaturase 1; TEM: transmission electron microscopy; TCGA: The Cancer Genome Atlas; TME: tumor microenvironment; XANES: X-ray absorption near-edge structure; XAS: X-ray absorption spectroscopy; XPS: X-ray photoelectron spectroscopy.
Supplementary Material
Supplementary methods, figures and tables.
Acknowledgements
This research is supported by funding from Research Personnel Cultivation Programme of Zhongda Hospital Southeast University (CZXM-GSP-RC148), Key scientific research projects of Anhui Provincial Department of Education (2023AH051741), Major scientific research projects of Anhui Provincial Health Commission (AHWJ2023A10146), National Natural Science Foundation of China (82272028, 81971621). All animal procedures were performed in accordance with the Articles of Association of Experimental Animal Welfare and Ethics Committee of Wannan Medical College, and approved by the Animal Welfare and Ethics Committee of Wannan Medical College (accreditation number: WNMC-AWE-2023473). Clinical sample collection and related procedures were approved by the Ethics Committee of the First Affiliated Hospital of Wannan Medical College (2025KY-25). Biorender.com was used to create Figures. ChatGPT (version 5.2) was used for language polishing.
Data availability
The corresponding author can provide the data supporting this study's findings upon a reasonable request.
Author contributions
X.X., J.Y. and Y.Z. contributed to the most experiments and experimental design, the experimental data analysis and prepared the manuscript. X.C., L.C., M.D. and J.D. were involved in parts of the experimental studies and supported the data analysis. Z.C, L.L. and J.C. supervised the research, analyzed the data and contributed to funding. All authors read and commented on the paper and Figures.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Ding J, Liu L, Zhang J, Liu Y, Xu H, Shen Z. et al. Unraveling Dynamic Structural Evolution of Single Atom Catalyst Via in Situ Surface-Enhanced Infrared Absorption Spectroscopy. J Am Chem Soc. 2025;147:9601-9
2. Qu Q, Mao Y, Ji S, Liao J, Dong J, Wang L. et al. Engineering the Lewis Acidity of Fe Single-Atom Sites Via Atomic-Level Tuning of Spatial Coordination Configuration for Enhanced Oxygen Reduction. J Am Chem Soc. 2025;147:6914-24
3. Tan H, Zhou P, Liu M, Gu Y, Chen W, Guo H. et al. Al-N3 Bridge Site Enabling Interlayer Charge Transfer Boosts the Direct Photosynthesis of Hydrogen Peroxide from Water and Air. J Am Chem Soc. 2024;146:31950-60
4. Wu J, Wang X, Wang Q, Lou Z, Li S, Zhu Y. et al. Nanomaterials with Enzyme-Like Characteristics (Nanozymes): Next-Generation Artificial Enzymes (II). Chem Soc Rev. 2019;48:1004-76
5. Li W, Liu S, Ding H, Zhao R, Zang P, Li S. et al. Three-Step Depletion Strategy of Glutathione: Tunable Metal-Organic-Framework-Engineered Nanozymes for Driving Oxidative/Nitrative Stress to Maximize Ferroptosis Therapy. Nano Lett. 2024;24:2071-80
6. Cheng J, Li L, Jin D, Dai Y, Zhu Y, Zou J. et al. Boosting Ferroptosis Therapy with Iridium Single-Atom Nanocatalyst in Ultralow Metal Content. Adv Mater. 2023;35:e2210037
7. Xue P, Zhuang H, Shao S, Bai T, Zeng X, Yan S. Engineering Biodegradable Hollow Silica/Iron Composite Nanozymes for Breast Tumor Treatment through Activation of the "Ferroptosis Storm". ACS Nano. 2024;18:25795-812
8. Cao C, Yang N, Su Y, Zhang Z, Wang C, Song X. et al. Starvation, Ferroptosis, and Prodrug Therapy Synergistically Enabled by a Cytochrome C Oxidase Like Nanozyme. Adv Mater. 2022;34:e2203236
9. Zhu Y, Wang W, Gong P, Zhao Y, Pan Y, Zou J. et al. Enhancing Catalytic Activity of a Nickel Single Atom Enzyme by Polynary Heteroatom Doping for Ferroptosis-Based Tumor Therapy. ACS Nano. 2023;17:3064-76
10. Jiang X, Stockwell BR, Conrad M. Ferroptosis: Mechanisms, Biology and Role in Disease. Nat Rev Mol Cell Biol. 2021;22:266-82
11. Friedmann Angeli JP, Krysko DV, Conrad M. Ferroptosis at the Crossroads of Cancer-Acquired Drug Resistance and Immune Evasion. Nat Rev Cancer. 2019;19:405-14
12. Zhao L, Zhou X, Xie F, Zhang L, Yan H, Huang J. et al. Ferroptosis in Cancer and Cancer Immunotherapy. Cancer communications (London, England). 2022;42:88-116
13. Zhang C, Liu X, Jin S, Chen Y, Guo R. Ferroptosis in Cancer Therapy: A Novel Approach to Reversing Drug Resistance. Mol Cancer. 2022;21:47
14. Wang Y, Wu X, Ren Z, Li Y, Zou W, Chen J. et al. Overcoming Cancer Chemotherapy Resistance by the Induction of Ferroptosis. Drug resistance updates: reviews and commentaries in antimicrobial and anticancer chemotherapy. 2023;66:100916
15. Zhou L, Wang Q, Yin P, Xing W, Wu Z, Chen S. et al. Serum Metabolomics Reveals the Deregulation of Fatty Acids Metabolism in Hepatocellular Carcinoma and Chronic Liver Diseases. Anal Bioanal Chem. 2012;403:203-13
16. Muir K, Hazim A, He Y, Peyressatre M, Kim DY, Song X. et al. Proteomic and Lipidomic Signatures of Lipid Metabolism in NASH-associated Hepatocellular Carcinoma. Cancer research. 2013;73:4722-31
17. Kawashima M, Tokiwa M, Nishimura T, Kawata Y, Sugimoto M, Kataoka TR. et al. High-Resolution Imaging Mass Spectrometry Combined with Transcriptomic Analysis Identified a Link between Fatty Acid Composition of Phosphatidylinositols and the Immune Checkpoint Pathway at the Primary Tumour Site of Breast Cancer. Br J Cancer. 2020;122:245-57
18. Van den Bossche V, Vignau J, Vigneron E, Rizzi I, Zaryouh H, Wouters A. et al. Pparα-Mediated Lipid Metabolism Reprogramming Supports Anti-EGFR Therapy Resistance in Head and Neck Squamous Cell Carcinoma. Nat Commun. 2025;16:1237
19. Wang X, Shu C, Wang G, Han P, Zheng L, Xu L. et al. Recent Progress of Noble Metal-Based Nanozymes: Structural Engineering and Biomedical Applications. Nanoscale. 2025;17:10557-80
20. Qiao B, Wang A, Yang X, Allard LF, Jiang Z, Cui Y. et al. Single-Atom Catalysis of CO Oxidation Using Pt1/Feox. Nat Chem. 2011;3:634-41
21. Cai S, Zhang W, Yang R. Emerging Single-Atom Nanozymes for Catalytic Biomedical Uses. Nano Res. 2023;16:13056-76
22. Nan F, Wei Q, Kou T, Zhao Y, Shen L, Li D. et al. Accurate Regulation of Carbon Skeletons Bonded to Fe-N4 Single Atom Nanozymes for Efficient Generation of Reactive Oxygen Species. Sci China:Chem. 2025;68:1067-77
23. Xue Q, Kang R, Klionsky DJ, Tang D, Liu J, Chen X. Copper Metabolism in Cell Death and Autophagy. Autophagy. 2023;19:2175-95
24. Budi CS, Deka JR, Hsu WC, Saikia D, Chen KT, Kao HM. et al. Bimetallic Co/Zn Zeolitic Imidazolate Framework ZIF-67 Supported Cu Nanoparticles: An Excellent Catalyst for Reduction of Synthetic Dyes and Nitroarenes. J Hazard Mater. 2021;407:124392
25. Li L, Chen G, Zheng H, Meng W, Jia S, Zhao L. et al. Room-Temperature Oxygen Vacancy Migration Induced Reversible Phase Transformation during the Anelastic Deformation in CuO. Nat Commun. 2021;12:3863
26. He C, Zhang Y, Zhang Y, Zhao L, Yuan LP, Zhang J. et al. Molecular Evidence for Metallic Cobalt Boosting Co2 Electroreduction on Pyridinic Nitrogen. Angew Chem. 2020;59:4914-9
27. Zhang J, Qu L, Shi G, Liu J, Chen J, Dai L. N, P-Codoped Carbon Networks as Efficient Metal-Free Bifunctional Catalysts for Oxygen Reduction and Hydrogen Evolution Reactions. Angew Chem Int Ed. 2016;128:2270-4
28. Zhang X, Wei H, Li S, Ren B, Jiang J, Qu G. et al. Manipulating Coordination Environment for a High-Voltage Aqueous Copper-Chlorine Battery. Nat Commun. 2023;14:6738
29. Zhang J, Liu J, Ran J, Lin X, Wang H, Qiu X. Oxidative Ammonolysis Modified Lignin-Derived Nitrogen-Doped Carbon-Supported Co/Fe Composites as Bifunctional Electrocatalyst for Zn-Air Batteries. Chin Chem Lett. 2024: 110403.
30. Li F, Sun H, Ren J, Zhang B, Hu X, Fang C. et al. A Nuclease-Mimetic Platinum Nanozyme Induces Concurrent Dna Platination and Oxidative Cleavage to Overcome Cancer Drug Resistance. Nat Commun. 2022;13:7361
31. Yan H, Wang Q, Wang J, Shang W, Xiong Z, Zhao L. et al. Planted Graphene Quantum Dots for Targeted, Enhanced Tumor Imaging and Long-Term Visualization of Local Pharmacokinetics. Adv Mater. 2023;35:2210809
32. Cheng R, Jiang L, Gao H, Liu Z, Mäkilä E, Wang S. et al. A pH-Responsive Cluster Metal-Organic Framework Nanoparticle for Enhanced Tumor Accumulation and Antitumor Effect. Adv Mater. 2022;34:2203915
33. Zhang W, Wang M, Liu B, Chen H, Tan J, Meng Q. et al. Glutathione Induced in Situ Synthesis of Cu Single-Atom Nanozymes with Anaerobic Glycolysis Metabolism Interference for Boosting Cuproptosis. Angew Chem Int Ed. 2024;63:e202402397
34. Zhou J, Xu D, Tian G, He Q, Zhang X, Liao J. et al. Coordination-Driven Self-Assembly Strategy-Activated Cu Single-Atom Nanozymes for Catalytic Tumor-Specific Therapy. J Am Chem Soc. 2023;145:4279-93
35. Wu Q, Zheng G, Li L, Wang L. Enhancing Peroxidase-Like Activity and Photothermal Property of Copper Single-Atom Nanozyme Via a Cascade Competition Strategy. Adv Funct Mater. 2025;35:2422588
36. Ye J, Li C, Xu J, Liu S, Qu J, Wang Q. et al. Engineered Nanozymes with Asymmetric Mn-O-Ce Sites for Intratumorally Leveraged Multimode Therapy. Adv Mater. 2025: 2419673.
37. Hu S, Lu H, Xie W, Wang D, Shan Z, Xing X. et al. TDO2+ Myofibroblasts Mediate Immune Suppression in Malignant Transformation of Squamous Cell Carcinoma. J Clin Invest. 2022 132
38. Zainal NS, Lee BKB, Wong ZW, Chin IS, Yee PS, Gan CP. et al. Effects of Palbociclib on Oral Squamous Cell Carcinoma and the Role of Pik3ca in Conferring Resistance. Cancer Biol Med. 2019;16:264-75
39. Gong W, Xu J, Wang Y, Min Q, Chen X, Zhang W. et al. Nuclear Genome-Derived Circular RNA circPUM1 Localizes in Mitochondria and Regulates Oxidative Phosphorylation in Esophageal Squamous Cell Carcinoma. Signal Transduction Targeted Ther. 2022;7:40
40. Gao J, Wang Z, Guo Q, Tang H, Wang Z, Yang C. et al. Mitochondrion-Targeted Supramolecular "Nano-Boat" Simultaneously Inhibiting Dual Energy Metabolism for Tumor Selective and Synergistic Chemo-Radiotherapy. Theranostics. 2022;12:1286-302
41. Stockwell BR. Ferroptosis Turns 10: Emerging Mechanisms, Physiological Functions, and Therapeutic Applications. Cell. 2022;185:2401-21
42. Yang X, Chen Y, Guo J, Li J, Zhang P, Yang H. et al. Polydopamine Nanoparticles Targeting Ferroptosis Mitigate Intervertebral Disc Degeneration Via Reactive Oxygen Species Depletion, Iron Ions Chelation, and GPX4 Ubiquitination Suppression. Adv Sci. 2023;10:e2207216
43. Wang R, Nie W, Yan X, Luo K, Zhang Q, Wang T. et al. Biomimetic Nanomotors for Deep Ischemia Penetration and Ferroptosis Inhibition in Neuroprotective Therapy of Ischemic Stroke. Adv Mater. 2025;37:2409176
44. Hu R, Chen X, Li Z, Zhao G, Ding L, Chen L. et al. Liquid Nanoparticles for Nanocatalytic Cancer Therapy. Adv Mater. 2023;35:2306469
45. Sinclair C, Bommakanti G, Gardinassi L, Loebbermann J, Johnson MJ, Hakimpour P. et al. mTOR Regulates Metabolic Adaptation of Apcs in the Lung and Controls the Outcome of Allergic Inflammation. Science. 2017;357:1014-21
46. Sinnott-Armstrong N, Sousa IS, Laber S, Rendina-Ruedy E, Nitter Dankel SE, Ferreira T. et al. A Regulatory Variant at 3q21.1 Confers an Increased Pleiotropic Risk for Hyperglycemia and Altered Bone Mineral Density. Cell Metab. 2021;33:615-28.e13
47. Chen J, Wu Z, Ding W, Xiao C, Zhang Y, Gao S. et al. SREBP1 siRNA Enhance the Docetaxel Effect Based on a Bone-Cancer Dual-Targeting Biomimetic Nanosystem against Bone Metastatic Castration-Resistant Prostate Cancer. Theranostics. 2020;10:1619-32
48. Yi J, Zhu J, Wu J, Thompson CB, Jiang X. Oncogenic Activation of PI3K-AKT-mTOR Signaling Suppresses Ferroptosis Via Srebp-Mediated Lipogenesis. Proc Natl Acad Sci, USA. 2020;117:31189-97
49. Wohlhieter CA, Richards AL, Uddin F, Hulton CH, Quintanal-Villalonga À, Martin A. et al. Concurrent Mutations in Stk11 and Keap1 Promote Ferroptosis Protection and Scd1 Dependence in Lung Cancer. Cell Rep. 2020;33:108444
50. Battaglioni S, Benjamin D, Wälchli M, Maier T, Hall MN. mTOR Substrate Phosphorylation in Growth Control. Cell. 2022;185:1814-36
51. Scharfman HE. Neuroscience. Metabolic Control of Epilepsy. Science. 2015;347:1312-3
52. Wang X, Zhang J, Zheng K, Du Q, Wang G, Huang J. et al. Discovering Metabolic Vulnerability Using Spatially Resolved Metabolomics for Antitumor Small Molecule-Drug Conjugates Development as a Precise Cancer Therapy Strategy. J Pharm Anal. 2023;13:776-87
53. Haider A, Wei Y-C, Lim K, Barbosa AD, Liu C-H, Weber U. et al. PCYT1A Regulates Phosphatidylcholine Homeostasis from the Inner Nuclear Membrane in Response to Membrane Stored Curvature Elastic Stress. Dev Cell. 2018;45:481-95 e8
54. Primrose MT, Claypool SM. Phosphatidylethanolamine. Trends Endocrinol Metab. 2024;35:929-30
55. Calianese DC, Birge RB. Biology of Phosphatidylserine (Ps): Basic Physiology and Implications in Immunology, Infectious Disease, and Cancer. Cell Commun Signaling. 2020;18:41
56. Dergunov AD, Baserova VB. Different Pathways of Cellular Cholesterol Efflux. Cell Biochem Biophys. 2022;80:471-81
57. Chakraborty M, Jiang X-C. Sphingomyelin and Its Role in Cellular Signaling. Lipid-mediated Protein Signaling. 2013:1-14
58. Sun X, Liu Y, Huang Z, Xu W, Hu W, Yi L. et al. SARS-CoV-2 Non-Structural Protein 6 Triggers NLRP3-Dependent Pyroptosis by Targeting ATP6AP1. Cell Death Differ. 2022;29:1240-54
59. Wang Y, Wang M, Liu Y, Tao H, Banerjee S, Srinivasan S. et al. Integrated Regulation of Stress Responses, Autophagy and Survival by Altered Intracellular Iron Stores. Redox Biol. 2022;55:102407
60. Dourado MR, Korvala J, Åström P, De Oliveira CE, Cervigne NK, Mofatto LS. et al. Extracellular Vesicles Derived from Cancer-Associated Fibroblasts Induce the Migration and Invasion of Oral Squamous Cell Carcinoma. J Extracell Vesicles. 2019;8:1578525
61. Li X, Liu H, Gao W, Yang Q, Li X, Zhou X. et al. Octadecyl Gallate and Lipid-Modified MnSe2 Nanoparticles Enhance Radiosensitivity in Esophageal Squamous Cell Carcinoma and Promote Radioprotection in Normal Tissues. Adv Mater. 2024;36:2311291
62. Chen J, Zhu Z, Pan Q, Bai Y, Yu M, Zhou Y. Targeted Therapy of Oral Squamous Cell Carcinoma with Cancer Cell Membrane Coated Co-Fc Nanoparticles Via Autophagy Inhibition. Adv Funct Mater. 2023;33:2300235
63. Xu F, Li X, Huang X, Pan J, Wang Y, Zhou S. Development of a pH-Responsive Polymersome Inducing Endoplasmic Reticulum Stress and Autophagy Blockade. Sci Adv. 2020;6:eabb8725
Author contact
Corresponding authors: Zhiyi Chen (zhiyi_chenedu.cn); Li Li (lilimpg.de); Junjie Cheng (jjchengedu.cn; jjchengedu.cn).
 Global reach, higher impact
Global reach, higher impact