Impact Factor
Theranostics 2019; 9(15):4508-4524. doi:10.7150/thno.34314 This issue Cite
Research Paper
Development of a Biocompatible Copolymer Nanocomplex to Deliver VEGF siRNA for Triple Negative Breast Cancer
Division of Pharmacology and Pharmaceutical Sciences, School of Pharmacy, University of Missouri-Kansas City, 2464 Charlotte Street, Kansas City, MO 64108, USA
Received 2019-2-21; Accepted 2019-5-20; Published 2019-6-9
Abstract
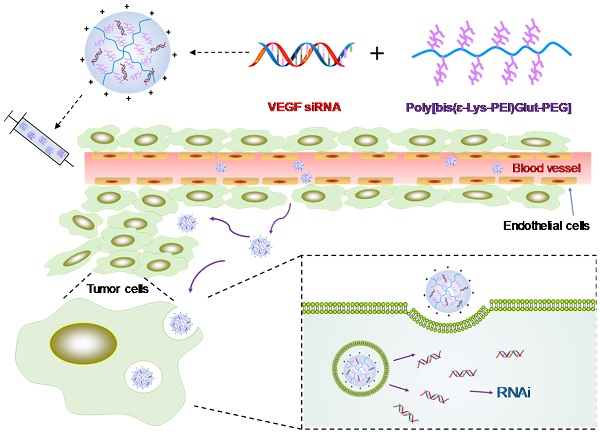
Triple negative breast cancer (TNBC) is the most difficult breast cancer subtype to treat. TNBC patients have significantly higher expression of vascular endothelial growth factor (VEGF) in tumors compared to non-TNBC patients. VEGF not only exerts its pro-angiogenic effects on endothelial cells but also acts as a survival and autocrine growth factor for VEGF receptor (VEGFR) expressing cancer cells. Silencing the expression of VEGF is therefore a potential therapy for TNBC.
Methods: A novel biocompatible linear copolymer poly[bis(ε-Lys-PEI)Glut-PEG] (PLEGP) was developed to deliver VEGF siRNA for TNBC therapy. The copolymer is composed of lysine and glutaric acid, a natural metabolite of amino acids in the body. Low-molecular weight polyethyleneimine (PEI) was grafted to the copolymer to efficiently condense siRNA into nanocomplex without inducing cytotoxicity. Various in vitro studies were performed to evaluate the stability, cellular uptake, tumor penetration, and biological activities of the VEGF siRNA nanocomplex. The anti-tumor activities of the nanocomplex was also evaluated in an orthotopic TNBC mouse model.
Results: PEIs with different molecular weights were evaluated, and the copolymer PLEGP1800 was able to easily form a stable nanocomplex with siRNAs and protect them from serum degradation. The siRNA/PLEGP1800 nanocomplex exhibited negligible cytotoxicity but showed high cellular uptake, high transfection efficiency, and high tumor penetration. In vitro activity studies showed that the siRNA nanocomplex significantly inhibited migration and invasion of TNBC cells. Moreover, the VEGF siRNA nanocomplex efficiently inhibited tumor growth in an orthotopic TNBC mouse model and down-regulated VEGF expression in the tumor.
Conclusion: PLEGP1800 is a safe and efficient copolymer to deliver siRNAs for TNBC therapy. It could potentially be applied to other cancers by changing the cargo and incorporating tumor-specific ligands.
Keywords: copolymer, glutaric acid, low-molecular weight PEI, triple negative breast cancer, siRNA, VEGF
Introduction
Triple negative breast cancer (TNBC) is an aggressive type of breast cancer that lacks the expression of human epidermal growth factor receptor 2, progesterone receptor, and estrogen receptor. Although TNBC only accounts for 12-17% of all breast cancers, it represents the most difficult breast cancer subtype to treat because of its aggressiveness, high proliferation, poor prognosis, and low survival rates [1, 2]. Current treatments for TNBC are limited to chemotherapy, including taxanes and anthracyclines, instead of receptor-targeted therapies, such as trastuzumab, pertuzumab, and tamoxifen [3-7]. The vascular endothelial growth factor (VEGF) family comprises VEGF-A, VEGF-B, VEGF-C, VEGF-D, and placenta growth factor (PGF). VEGF plays important roles in angiogenesis and is correlated with high metastasis of breast cancer. In particular, TNBC patients have significantly higher VEGF expressions in tumors compared to non-TNBC patients [8-11]. However, anti-VEGF therapies using antibodies, such as bevacizumab and ramucirumab, failed to improve the survival rate of TNBC patients in a few phase III clinical trials [12-14]. The disappointing result is possibly due to the fact that VEGF not only exerts its pro-angiogenic effects on endothelial cells but also acts as a survival and autocrine growth factor for VEGF receptor (VEGFR) expressing cancer cells [15, 16]. Moreover, VEGFR-1 is highly expressed in TNBC cells, and it is predominantly expressed internally in the nuclear envelope [17]. As a result, anti-VEGF antibodies used in those clinical trials only blocked VEGF's interaction with the VEGFR on the plasma membrane of tumor cells and nearby endothelial cells, but not the VEGFR inside tumor cells. We, therefore, hypothesize that silencing the expression of VEGF in TNBC cells will not only reverse VEGF's pro-angiogenic effect on endothelial cells but also block its pro-tumor activity on tumor cells.
Small interfering RNAs (siRNAs) have the capability to specifically down-regulate target genes with high potency [18]. In our previous study, we identified a VEGF siRNA and reported its inhibitory effect on the proliferation and migration of HER2+ breast cancer cells [19].However, poor stability, negative charge and large molecular weight (~14 kDa) of siRNAs limit their delivery to target cells in animals to exert biological activities [20]. Due to their safety and ease of production, non-viral vectors, such as polymers, micelles, liposomes, and nanoparticles, have attracted a great deal of attention to overcome these obstacles of siRNA delivery [21-24]. In August 2018, the lipid nanoparticle-formulated siRNA (Patisiran) treating hereditary transthyretin-mediated amyloidosis was approved by the Food and Drug Administration (FDA) as the first siRNA drug in the world.
Polyethyleneimine (PEI) is one of the promising polymers for siRNA delivery. PEI efficiently neutralizes the negative charge of siRNAs, forms stable nanocomplexes with siRNAs through electrostatic interactions and releases siRNAs from the endosome to the cytosol via the “proton sponge effect” [25, 26]. However, the transfection efficiency and cytotoxicity of siRNA/PEI complexes are highly correlated with the molecular weight of PEI [27]. High-molecular-weight (HMW) PEI has high transfection efficiency but with considerable cytotoxicity. On the contrary, low-molecular-weight (LMW) PEI has low cytotoxicity but also exhibits low gene transfection efficiency because of its low condensation capacity [27, 28]. The low polymerization degree of LMW-PEI is correlated with its small size and limits its ability to form strong electrostatic associations with nucleic acids in a nanoscale structure [27]. One strategy is grafting LMW PEIs to a polymer to increase their condensation capability with nucleic acids [29, 30].
Polyethylene glycol (PEG) has been widely used to improve the pharmacokinetics and stability of nanoscale drug delivery systems, including siRNA nanocomplexes, through the “stealth effect” [31]. We recently developed a PEG-glutaric acid-lysine based biodegradable copolymer to prepare a polymer-drug conjugate of rapamycin [32]. In the present study, LMW branched PEIs with different molecular weights (600 Da, 1800 Da, and 10k Da) were conjugated to the copolymer to form a positively charged and biodegradable nanocarrier to deliver the VEGF siRNA to TNBC. The polymer grafted with PEI1800 efficiently delivered the VEGF siRNA into TNBC cells with a high transfection efficiency and low cytotoxicity. In vitro activity studies showed that the siRNA nanocomplex significantly inhibits migration and invasion of TNBC cells. Moreover, the nanocomplex exhibited efficient tumor penetration in a 3D tumor spheroid model, suggesting a good penetration capability of the nanocomplex in tumor microenvironment in vivo. The VEGF siRNA nanocomplex efficiently inhibit tumor growth in vivo and successfully down-regulate VEGF expression in the tumor.
Materials
Poly(ethyleneimine) (MW 600, 1800, 10k), BD Matrigel, Annexin V-FITC apoptosis kit, GelRedTM and chemical regents were purchased from Fisher Scientific (Pittsburg, PA). Lipofectamine® 2000, VEGF siRNA, and Cy5-labeled siRNA were obtained from Invitrogen (Carlsbad, CA). VEGF ELISA kit was ordered from R&D system (Minneapolis, MN). The CellTiter-Glo luminescent cell viability assay kit was purchased from Promega (Madison, WI).
Cell Culture
HeLa, MDA-MB-231 and MCF-10A cells were purchased from American Type Culture Collection. HeLa and MDA-MB-231 cells were cultured in DMEM with 10% fetal bovine serum (FBS), 100 µg/mL streptomycin and 100 units/mL penicillin. MCF-10A cells were cultured in Mammary Epithelial Cell Growth Medium (Lonza, MD) with 100 ng/ml cholera toxin (Fisher, PA). Human Umbilical Vein Endothelial Cells (HUVEC) were purchased from Fisher Scientific (Pittsburg, PA) and cultured in Endothelial Cell Growth Medium containing supplement reagents (R&D Systems Inc., Minneapolis, MN). The cells were incubated in a humidified atmosphere at 37°C with 5% CO2.
Synthesis of the poly[bis(Lys-PEI)Glut-PEG] Copolymer (PLEGP)
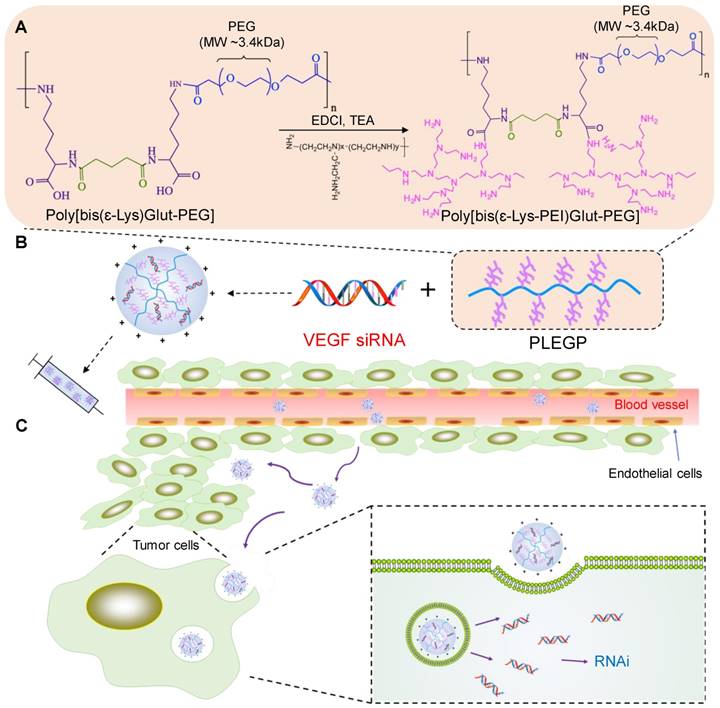
The synthetic reaction scheme is presented in Figure 1A. Poly[bis(ε-Lys)Glut-PEG] was synthesized as we described before [32], and 50 mg of the polymer was dissolved in dimethylformamide (DMF) with 75 mg of 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide (EDC) and 75 µl of Triethylamine (TEA). PEI600, PEI1800 and PEI10k were added to the solution at different polymer:PEI molar ratios (1:1, 1:10, and 1:20). The mixture was stirred at room temperature for 24 h with N2 protection, and the polymers PLEGP600, PLEGP1800 and PLEGP10k were precipitated by adding cold diethyl ether and dialyzed for 24 h (MWCO: 14kDa). Structures of the polymers were confirmed by 1H-NMR (Figure S1).
Schematics of the siRNA/PLEGP nanocomplex. (A) Synthetic reaction scheme of PLEGP. (B) Fabrication of the siRNA/PLEGP nanocomplex. (C) Delivery of the nanocomplex to tumor cells. The copolymer (PLEGP600, PLEGP1800, PLEGP10k) were synthesized by conjugating LMW branched PEIs to poly[bis(ɛ-Lys)Glut-PEG]. PLEGP efficiently condenses VEGF siRNA into a stable nanocomplex structure and protect the siRNA from serum degradation. The siRNA/PLEGP nanocomplex accumulated in tumor tissues via the enhanced permeability and retention (EPR) effect. The nanocomplex enters the cells via endocytosis and release the siRNA into the cytoplasm via the “proton sponge effect.”
Fabrication and Characterization of the siRNA/PLEGP Nanocomplexes
PLEGPs were mixed with 20 μM VEGF siRNA (sequence: 5'-GCUACUGCCAUCCAAUCGAtt-3') in different siRNA:PLEGP w/w ratios (1:1, 1:5, 1:10, and 1:20 ) at room temperature for 30 min, and then diluted to 100 nM with PBS and Opti-MEM for characterization and transfection, respectively. The siRNA/PLEGP ratio was optimized using a gel retardation assay with a 2% agarose gel as previously reported [31]. Particle size and zeta potential of the siRNA/PLEGP nanocomplexes were determined using a Malvern Zetasizer Nano-ZS (Malvern Instruments, MA) in 100 mM HEPES buffer (pH 7.4). Morphology of the nanocomplexes was studied using a CM12 Transmission Electron Microscope (TEM) (Philips, Germany).
2,4,6-Trinitrobenzene Sulfonic Acid (TNBS) Assay
Substitution efficiency of the primary amines by PEI in PLEGPs was determined with a TNBS assay. Briefly, PLEGP600, PLEGP1800 and PLEGP10k synthesized at a polymer/PEI molar ratio 1:10 were added to 96-well plates at a concentration of 1mg/mL and incubated with TNBS buffer at 37°C for 2 h. After adding 10% sodium dodecyl sulfonate buffer and 1N hydrochloric acid, absorbance of the samples was measured at 335 nm using a Spectramax 190 microplate spectrophotometer (Molecular Devices, CA). A series of concentrations of PEI600, PEI1800 and PEI10k were added as described above to generate standard curves.
Serum Stability Assay
VEGF siRNAs were incubated with PLEGPs at a ratio of 1:10 (w/w) to form siRNA/PLEGP nanocomplexes and then incubated with 50% rat serum at 37°C for various time intervals. siRNAs in the nanocomplexes were then released by incubation with 40 μM heparin, electrophoresed in 10% native polyacrylamide gel electrophoresis (PAGE) gel, and visualized with GelRedTM staining.
In Vitro Cytotoxicity Study
Cytotoxicity of the siRNA/PLEGP nanocomplexes was evaluated using the 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) assay. Briefly, 1×104 HeLa, MDA-MB-231 and MCF-10A cells were seeded in 96-well plates and incubated at 37°C overnight. PLEGP600, PLEGP1800 and PLEGP10k were mixed with scrambled siRNAs to form nanocomplexes, diluted in DMEM medium containing 10% FBS with a final siRNA concentration of 100 nM, and incubated with the cells for 24 and 48 h. MTT assays were then performed to measure the cytotoxicity of the nanocomplexes as described before [31].
Cellular Uptake Study
HeLa and MDA-MB-231 cells (1×105) were seeded in 24-well plates and incubated at 37°C overnight. 5' ends of the VEGF siRNA antisense strands were labeled with Cy5. The cells were transfected with Cy5-siRNA/PLEGP nanocomplexes at an siRNA concentration of 100 nM in Opti-MEM for 6 h. The cells were then washed with 40 μM heparin to remove non-specifically bound nanocomplexes and then subjected to fluorescence analysis with a FACS II flow cytometer (BD instrument, NJ) as described before [31]. Cellular uptake of nanocomplex was also evaluated using confocal microscope. Co-localization of Cy5-siRNA and lysosome were analyzed as the Pearson's correlation coefficient using ImageJ as previously reported [33].
Three-Dimensional (3D) Spheroid Penetration Study
3D spheroids were prepared using Spheroid Formation ECM as per the company's protocol (Amsbio, Cambridge, MA). Three thousand MDA-MB-231 cells were suspended in 50 µL Spheroid Formation ECM, added into a Corning™ 96-well Ultra-low attachment microplate (Pittsburgh, PA), centrifuged at 200 g for 3 min at 4°C, and incubated at 37°C for 120 h to form spheroids. The spheroids were incubated with the Cy5-siRNA/PLEGP nanocomplexes diluted in Opti-MEM medium at 37°C for 6 h, washed with PBS, and fixed in 10% paraformaldehyde. Penetration of the Cy5-siRNA inside the spheroids were analyzed using confocal microscopy (Leica TCS SP5, Germany).
Transfection Study of the Luciferase siRNA/PLEGP Nanocomplex
Approximately 5×103 HeLa cells were seeded into each well of a black well, clear bottom plate and incubated at 37°C overnight. Sixty nanograms of Luciferase-pcDNA3 plasmid (Addgene, MA) and 30 ng pMCS-Green Renilla Luc plasmid (ThermoFisher, CA) were mixed with 0.2 μL of Lipofectamine® 2000 (ThermoFisher, CA) in 100 μL of Opti-MEM and incubated at 37°C for 6 h for the first step transfection. Firefly luciferase siRNAs (5'-CUUACGCUGAGUACUUCGAtt-3') or scrambled siRNAs were mixed with PLEGP to form nanocomplexes as mentioned above, diluted in Opti-MEM, and incubated with the cells at 37°C for 6 h in the second-step transfection. Next, the medium was replaced with DMEM containing 10% FBS and incubated for another 42 h. Silencing activity of the luciferase siRNA/PLEGP nanocomplex was quantitated using a Dual-Luciferase® Reporter Assay kit (Promega, WI). For each siRNA, the ratios of Firefly to Renilla luciferase activity were normalized to the cells treated with scrambled siRNA and presented as the percentage mean ± SD.
Silencing Activity of the VEGF siRNA/PLEGP Nanocomplexes
Silencing activity of the VEGF siRNA/PLEGP nanocomplexes at the protein level was measured using an ELISA kit. Approximately 1×105 MDA-MB-231 cells were seeded in each well of a 24-well plate and incubated at 37°C overnight. The VEGF siRNA/PLEGP nanocomplexes were prepared as described above at a 100 nM final concentration, transfected into cells, and incubated for 24 h. The medium was replaced with DMEM complete medium and incubated for 24 h at 37°C. Lipofectamine® 2000 was used as a control. After transfection, the media were collected, and the protein levels of VEGF were quantified using a DueSet Human VEGF ELISA kit.
Migration and Invasion Assays
Migration and invasion assays were performed as we described before [19]. For the invasion assay, 50 μg of Matrigel were coated on the top of a transwell chamber. MDA-MB-231 cells were treated with the VEGF siRNA/PLEGP1800 nanocomplex as described above. Forty-eight hours after transfection, the cells were detached, re-suspended in DMEM supplemented with 0.5% FBS, and transferred to transwell chambers at a density of 5×104 cells/well. After 6 h for the migration assay and 24 h for the invasion assay, cells in the lower chamber were fixed with 10% paraformaldehyde, stained with 0.05% crystal violet, and counted under a microscope. Quantitative analysis was performed by counting the cells on the bottom of the membrane in six random microscope fields from three independent samples. The number of migrated and invaded cells were normalized to the cells treated with scrambled siRNA and presented as the percentage mean ± SD.
3D Tumor Spheroid Invasion Study
3D spheroids of MDA-MB-231 cells were prepared as described above, and the spheroid invasion assay was performed as per the company's protocol (Amsbio, Cambridge, MA). Seventy-two hours after the formation of spheroids, 50 µL of invasion matrix (Amsbio, Cambridge, MA) was added to the wells, and the plate was centrifuged at 300 g for 4 min at 4°C, followed by incubation at 37°C for 60 min to induce gel formation. The spheroids were then incubated with the VEGF siRNA/PLEGP1800 nanocomplex diluted in Opti-MEM at 37°C for 24 h, followed by replacing the medium with DMEM supplemented with 10% FBS and incubation for up to 3 days. The spheroids in each well were photographed every 24 h at 40x magnification, and invasion of the 3D spheroids was analyzed using ImageJ. The spheroid areas of each group were normalized to the spheroid area of the group at day 1 and presented as the percentage mean ± SD.
Cells Proliferation Assay
Five thousand MDA-MB-231 cells were plated each well into a black well, clear bottom 96-well plate and incubated at 37°C overnight. After washing with PBS, the cells were incubated with the VEGF siRNA/PLEGP1800 nanocomplex diluted in Opti-MEM at 37°C for 48 or 72 h. One hundred microliters of CellTiter-Glo buffer (Promega, WI) was then added into the plate, and luminescent intensity was measured using a Spectramax M5e spectrophotometer (San Jose, CA).
For HUVEC proliferation assay, 5×103 HUVEC cells were plated each well into a 96-well plate and incubated with Endothelial Cell Growth Medium at 37°C overnight. After washing with PBS, the cells were cultured in DMEM medium containing 10% (v/v) culture medium derived from siRNA/PLEGP1800-treated MDA-MB-231 cells for 24 h. Cell proliferation was evaluated by determining the luminescent intensity as described above.
Endothelial Cell Tube Formation Assay
A 24-well plate was coated with 500 µl of 1:1 (v/v) diluted matrigel in Endothelial Cell Growth Medium for 60 min at 37°C [34]. HUVEC cells were suspended with culture medium derived from siRNA/PLEGP1800-treated MDA-MB-231 cells and added into the matrigel-coated plate at a density of 5×104 cells/well. Tube formation was evaluated after 24 h incubation and counted under a microscope at 40x magnification.
Biodistribution, in Vivo Pharmacokinetics and Anti-tumor Activity Study
The animal protocol was approved by the Institutional Animal Care and Use Committee (IACUC) of the University of Missouri-Kansas City. Approximately 5×105 MDA-MB-231 cells were implanted into the right mammary gland of female nude mice to generate an orthotopic model of TNBC as described [35]. For biodistribution study, six mice were randomly divided into two groups and intravenously injected with Cy5-labeled siRNA or Cy5-siRNA/PLEGP1800 nanocomplex at a siRNA dose of 0.1 mg/kg [31]. After 24 h, the mice were euthanized, and major organs (liver, tumor, lung, spleen, kidney, muscle of the foreleg, and heart) were harvested for fluorescence imaging using a Bruker MS FX PRO imaging system (Billerica, CA). For the anti-tumor activity study, tumor-bearing mice were randomly divided into three groups and intravenously injected with VEGF siRNA/PLEGP1800, scrambled siRNA/PLEGP1800 and saline. The injections were performed every 4 days for 4 times with a dose of 1.5 mg siRNA/kg [36]. Tumor volumes were calculated using the formula V= longest diameter  shortest diameter2/2.
shortest diameter2/2.
VEGF protein expressions in tumor tissues were measured using a VEGF ELISA kit, as previously reported [37]. Briefly, tumor tissues were homogenized in RIPA buffer and centrifuged at 13,000 g for 10 min to remove the precipitate. After determining the total protein concentration using a BCA protein assay kit (ThermoFisher, CA), the concentrations of VEGF protein in tumor tissues were determined using the VEGF ELISA kit and presented as VEGF (%) compared with the saline treatment group. Western blot was also performed to confirm the silencing activity of the VEGF siRNA using a VEGF antibody (ThermoFisher, CA).
Alanine aminotransferase (ALT) and aspartate aminotransferase (AST) assays (Sigma-Aldrich, MO) were performed to evaluate hepatotoxicity of the siRNA/PLEGP1800 nanocomplex. Immunohistochemical analysis was performed to determine the microvessel density in tumor tissues using a mouse anti-human CD31 antibody (ThermoFisher, CA). Hematoxylin and Eosin (H&E) staining was also performed on tumor, liver, kidney and lung tissue sections to evaluate the in vivo cytotoxicity as previously reported [38].
For pharmacokinetic study, six BALA/c mice were randomly divided into two groups and intravenously injected with siRNA or siRNA/PLEGP1800 nanocomplex at a siRNA dose of 1.5 mg/kg, which contains 10% Cy5-labeled siRNA. At various time points (5 min, 10 min, 30 min, 1 h, 2 h, 4 h, 6 h, 24 h and 48 h) after injection, 10 µL blood was collected from tail vein and dissolved in 200 µL lysis buffer as describe before[39]. The Cy5 fluorescence intensity were measured using a Bruker MS FX PRO imaging system, and the area under curve (AUC) was calculated.
Statistical analysis
Data are presented as the mean ± standard deviation (SD). Statistical analysis was performed using a two-way analysis of variance (ANOVA) with Tukey's post hoc test). P<0.05 was considered statically significant.
Results
Fabrication and Characterization of siRNA/PLEGP Nanocomplexes
The 1H NMR spectra of the PLEGP polymers were presented in Figure S1, and fabrication of the siRNA/PLEGP nanocomplexes is illustrated in Figure 1. For each of the LMW-PEI, PLEGP was synthesized at three different polymer/PEI molar ratios (1:1, 1:10, and 1:20). PLEGPs were then condensed with siRNAs at different PLEGP/siRNA weight ratios of 1:1, 5:1, 10:1, and 20:1. As shown in Figure 2A, condensation between siRNAs and PLEGPs increases with the PLEGP/siRNA ratio as well as the molecular weight of PEI. For example, when PLEGPs were synthesized at the polymer/PEI molar ratio of 1:1, PLEGP600, PLEGP1800, and PLEGP10k efficiently condensed the siRNA at a PLEGP/siRNA weight ratio of 10:1, 5:1, and 1:1, respectively. PEI with high molecular weight has high charge density and therefore can efficiently condense siRNAs. For the same PEI, the condensation capability of PLEGP is correlated with the polymer/PEI ratio during synthesis. More PEI molecules were grafted to the polymer backbone at high polymer/PEI ratios, thus increasing its charge density and subsequently enhancing its condensation capability with siRNAs. Zeta potential results in Figure 2B were consistent with the gel retardation results in Figure 2A. Zeta potentials of PLEGP600, PLEGP1800 and PLEGP10k were performed in Figure S2.
Characterizations of the siRNA/PLEGP600, siRNA/PLEGP1800 and siRNA/PLEGP10k nanocomplexes. (A) Gel retardation assay. (B) Zeta potential. (C) Particle size and TEM. The scale bar represents 500 nm.
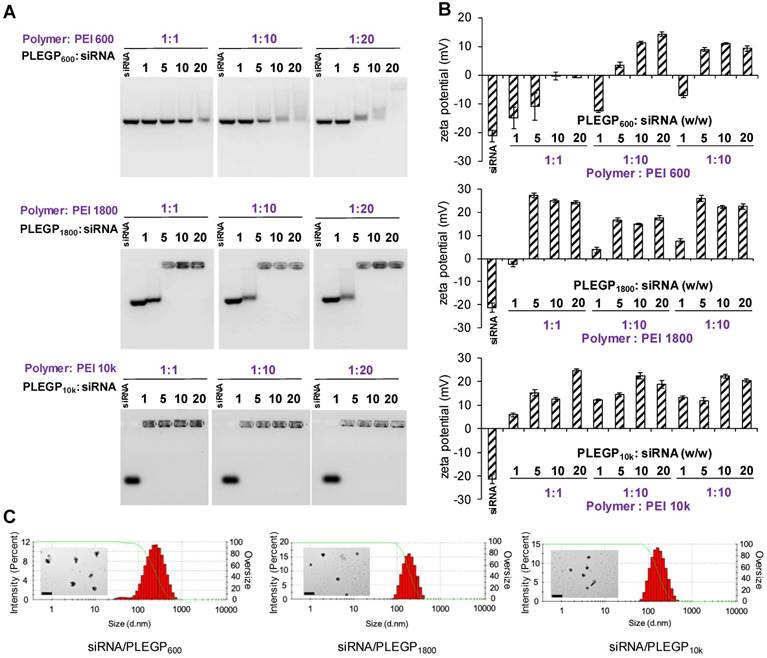
Cytotoxicity of PEI is the major hurdle in its therapeutic applications. Therefore, we aim to optimize the polymer/PEI ratio to condense siRNAs into nanocomplexes using the least amount of PEI, thus reducing the potential cytotoxicity associated with PEI. As shown in Figure 2A and 2B, the polymer/PEI molar ratio at 1:10 and 1:20 showed similar condensation capability, we decided to use the 1:10 ratio for future studies to minimize the amount of PEI in the siRNA nanocomplex. Based on gel retardation assay and zeta potential analysis, we selected PLEGPs synthesized with the polymer/PEI molar ratio of 1:10 for the following studies. A TNBS assay was performed to determine the grafting efficiency of PEI on the polymer. As shown in Table 1, the percentage of the amine groups that were grafted with PEI600, PEI1800, and PEI10k was 57.7%, 49.3%, and 16.2%, respectively. The grafting efficiency is inversely proportional with the molecular weight of PEI, suggesting that steric hindrance is the determining factor in the grafting efficiency of PEI. Molecular weight of Poly[bis(ε-Lys)Glut-PEG] was determined using gel permeation chromatography as we reported before [32], and molecular weight of PLEGP was calculated based on the grafting efficiency of PEI to the polymer.
Substitution Efficiency of the Primary Amine by PEI in PLEGP
| Polymer/PEI molar ratio | PEI (nmole)/ Polymer (mg) | Substitution Efficiency of Primary Amine | Molecular Weight | |
|---|---|---|---|---|
| PLEGP600 | 1:10 | 236.8 | 57.7% | 31471 |
| PLEGP1800 | 1:10 | 165.8 | 49.3% | 38487 |
| PLEGP10k | 1:10 | 43.6 | 16.2% | 47844 |
We selected the PLEGP/siRNA weight ratio of 10:1 to ensure that all three PLEGPs can efficiently form stable siRNA/PLEGP nanocomplexes. Particle size of the siRNA/PLEGP600, siRNA/PLEGP1800, and siRNA/PLEGP10k nanocomplexes was 267, 208 and 183 nm, respectively. Polydispersity index (PDI) of the siRNA/PLEGP600, siRNA/PLEGP1800, and siRNA/PLEGP10k nanocomplexes was 0.263, 0.185 and 0.181, respectively, suggesting a narrow size distribution of the nanocomplexes. Morphology of the nanocomplexes was studied with TEM (Figure 2C), and the result is consistent with particle size analysis.
Serum Stability
Poor stability of siRNAs in the serum is one of the major hurdles in delivering siRNAs to target tissues in vivo. We thus sought to evaluate the stability of the siRNA/PLEGP nanocomplexes in 50% rat serum. As shown in Figure 3A, all the three nanocomplexes protected the siRNA from serum degradation for at least 24 h. The siRNA/PLEGP600 nanocomplex exhibited some degree of degradation after 36 h incubation in the serum. By contrast, both siRNA/PLEGP1800 and siRNA/PLEGP10k nanocomplexes protected the siRNA in the serum for up to 36 h.
Serum stability and cytotoxicity of the siRNA/PLEGP nanocomplexes. (A) Serum stability of siRNA/PLEGP600, siRNA/PLEGP1800 and siRNA/PLEGP10k nanocomplexes in 50% rat serum for 0, 1, 3, 6, 12, 24, and 36 h. (B) Viability of HeLa cells treated with the siRNA/PLEGP nanocomplexes for 24 and 48 h. (C) Viability of MDA-MB-231 cells treated with the siRNA/PLEGP nanocomplexes for 24 and 48 h. (D) Viability of MCF-10A cells treated with the siRNA/PLEGP nanocomplexes for 24 and 48 h. Lipofectamine® 2000 was used as a control. All results are presented as the mean ± SD (n=3). (* P <0.05)
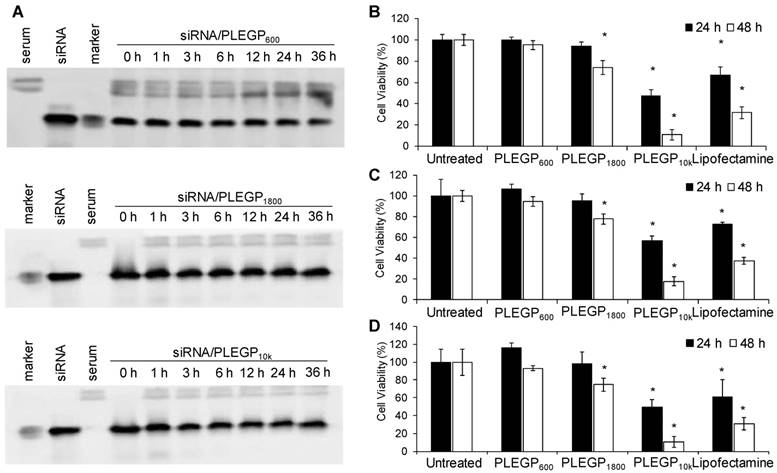
Cytotoxicity Study
Cytotoxicity of HMW-PEIs is the major limitation for their therapeutic applications. We, therefore, used LMW-PEIs in this study to efficiently deliver siRNAs without inducing cytotoxicity. As shown in Figure 3B, PLEGP600 did not induce any cytotoxicity in HeLa cells after transfection for 24 and 48 hr. PLEGP1800 showed somewhat toxicity in HeLa cells only after transfection for 48 h. By contrast, PLEGP10k and Lipofectamine® 2000 exhibited high toxicity at 24 and 48 h post-transfection. While the toxicity of Lipofectamine® 2000 is well documented, the toxicity of PLEGP10k is mainly due to the relatively high molecular weight of the grafted PEI. Similar results were also observed in MDA-MB-231 cells in Figure 3C and MCF-10A cells in Figure 3D.
Cellular Uptake Study
We next evaluated cellular uptake of the siRNA/PLEGP nanocomplexes in HeLa (Figure 4A and C) and MDA-MB-231 (Figure 4B and D) cells using flow cytometry and confocal microscopy. Cellular uptake of the siRNA/PLEGP nanocomplexes is strongly correlated with the molecular weight of PEI in the polymer. PLEGP10k exhibited much higher cellular uptake than Lipofectamine® 2000 in HeLa and MDA-MB-231 cells. By contrast, PLEGP600 showed the lowest uptake in the cells, suggesting that PLEGP600 may not be a good candidate for siRNA delivery regardless of its safety.
Cellular uptake of the siRNA/PLEGP nanocomplexes in HeLa and MDA-MB-231 cells. siRNA was labeled with Cy5 for fluorescence analysis using flow cytometry (A, B) and confocal microscopy (C, D). Fluorescence intensity of HeLa cells (A) and MDA-MB-231 cells (B) that take up the siRNA/PLEGP nanocomplexes. Representative confocal images of HeLa cells (C) and MDA-MB-231 cells (D) treated with the siRNA nanocomplexes. The scale bar represents 20 µm. Co-localization of Cy5-siRNA and Lysotracker shown in HeLa cells (E) and MDA-MB-231 cells (F) was quantitated using ImageJ. All results are presented as the mean ± SD (n=3). (* P <0.05; ** P <0.01).
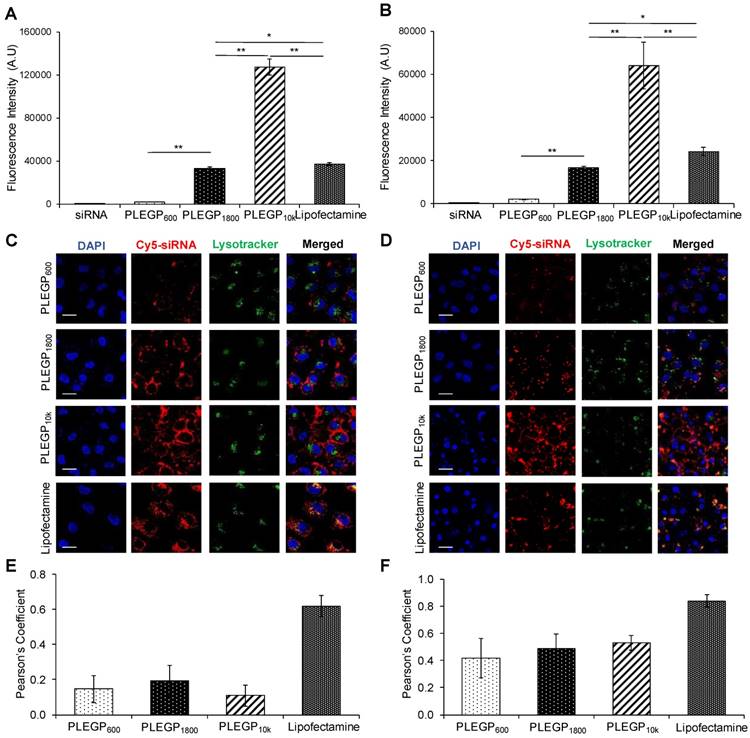
Subsequently, we used confocal microscopy to study intracellular distribution of the siRNA/PLEGP nanocomplexes in HeLa (Figure 4C) and MDA-MB-231 (Figure 4D) cells. The results are consistent with the data from flow cytometry analysis. The siRNA/PLEGP1800 nanocomplex showed similar cellular uptake as Lipofectamine® 2000, and the siRNA/PLEGP10k nanocomplex exhibited the highest cellular uptake in both HeLa and MDA-MB-231 cells. Moreover, in contrast to the Lipofectamine® 2000 group, which displayed substantial overlap between Cy5-labeled siRNAs and LysoTracker in HeLa (Figure 4E) and MDA-MB-231 (Figure 4F), cells treated with the siRNA/PLEGP nanocomplexes exhibited minimal entrapment in lysosomes, indicating that PLEGP can efficiently release siRNAs into the cytoplasm through the distinctive endosome escape capability of PEI.
3D Spheroid Penetration Study
Tumor penetration is another challenge for cancer therapeutics including siRNAs. We, therefore, investigated the penetration capability of the siRNA/PLEGP nanocomplexes in a 3D spheroid model. MDA-MB-231 spheroids with a diameter of approximately 500 µm were generated to mimic the tumor microenvironment [40, 41]. After 6 h incubation with MDA-MB-231 spheroids, the Cy5-siRNA/PLEGP10k nanocomplex exhibited the deepest penetration and the highest fluorescence intensity in the center of the spheroids (Figure 5A). The Cy5-siRNA/PLEGP600 nanocomplex was mainly observed in the peripheral areas of the spheroids, with much weaker fluorescence intensity, indicating its limited penetration capability. As shown in Figure 5A, the Cy5-siRNA/PLEGP1800 nanocomplex also penetrated into the core of the spheroids, and the fluorescence intensity was approximately 63% of the Cy5-siRNA/PLEGP10k nanocomplex (Figure 5B).
3D spheroid penetration study and silencing activity of the siRNA/PLEGP nanocomplexes. (A) Representative z-stacked confocal images of the spheroids with a z-step of 50 µm. The scale bar represents 50 µm. (B) Mean fluorescence intensity of the z-stacked confocal images vs. the distance from the periphery of the spheroids. (C) Silencing activity of the luciferase siRNA/PLEGP nanocomplex in HeLa cells. Dual luciferase reporter assay was used to detect silencing activity. For each siRNA, the ratios of Firefly to Renilla luciferase activity were normalized to the cells treated with scrambled siRNA and presented as the percentage mean ± SD (n=3). (D) Silencing activity of the VEGF siRNA/PLEGP nanocomplexes at the protein level in MDA-MB-231 cells. Silencing activity of each siRNA was normalized to scrambled siRNA and presented as the percentage mean ± SD (n=3).
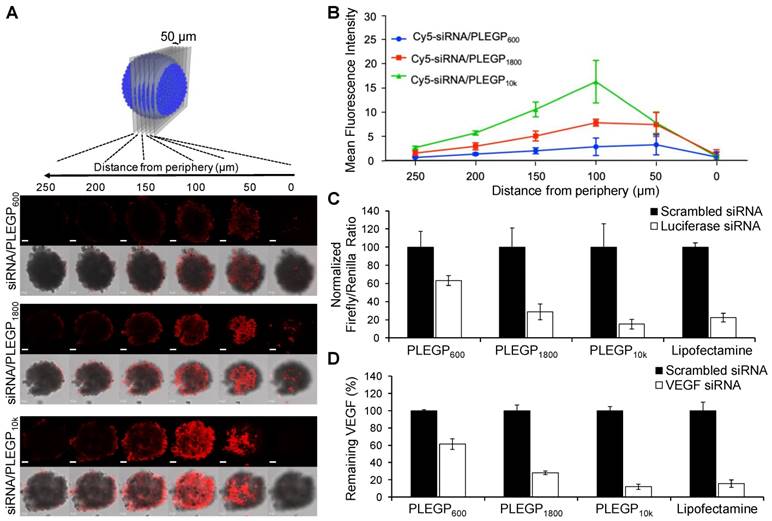
Transfection Efficiency of the Luciferase siRNA/PLEGP Nanocomplex in HeLa Cells
Transfection efficiency of the siRNA/PLEGP nanocomplexes was first evaluated in HeLa cell line, which has been widely used in transfection studies because of its high transfection efficiency [42]. First, luciferase and renilla plasmids were co-transfected into HeLa cells, followed by transfecting the cells with the luciferase siRNA/PLEGP nanocomplexes. As shown in Figure 5C, 85% of the expression of the luciferase plasmid was silenced by the luciferase siRNA/PLEGP10k nanocomplex, followed by 71% for PLEGP1800 and only 36% for PLEGP600. The positive control Lipofectamine® 2000 displayed a 77% silencing efficiency in HeLa cells.
Silencing Activity of VEGF siRNA/PLEGP Nanocomplexes in TNBC cells
We next examined the silencing activity of VEGF siRNA/PLEGP nanocomplexes in MDA-MB-231 cells. The results illustrated in Figure 5D are consistent with the cellular uptake results in MDA-MB-231 cells (Figure 4) and the transfection study using HeLa cells (Figure 5C). The VEGF siRNA/PLEGP10k nanocomplex exhibited 88% silencing activity of VEGF, which is the highest among all the three siRNA/PLEGP nanocomplexes. The VEGF siRNA/PLEGP1800 nanocomplex displayed 72% silencing activity, a lower value than the VEGF siRNA/PLEGP10k nanocomplex and VEGF siRNA/Lipofectamine (84%), but the value is much higher than the VEGF siRNA/PLEGP600 nanocomplex, which exhibits approximately 39% silencing activity.
Among the three PLEGP polymers, PLEGP10k showed the highest cellular uptake, highest tumor penetration, highest silencing activity, but also the highest cytotoxicity. By contrast, PLEGP1800 exhibited reasonably high cellular uptake, tumor penetration, and silencing activity without inducing significant cytotoxicity. We therefore selected PLEGP1800 as a safe and highly efficient polymer for the following in vitro and in vivo activity studies of the VEGF siRNA.
Migration and Invasion Assays
It was reported that autocrine VEGF stimulates the migration and invasion of breast cancer cells [43]. Thus, we sought to examine whether the VEGF siRNA/PLEGP1800 nanocomplex inhibits the migration and invasion of MDA-MB-231 cells. Compared with scrambled siRNA/ PLEGP1800 nanocomplex, the VEGF siRNA/PLEGP1800 nanocomplex inhibited approximately 64% of the migration (Figure 6A and B) and 67% of the invasion (Figure 6C and D) of MDA-MB-231 cells.
The VEGF siRNA/PLEGP1800 nanocomplex inhibits the migration and invasion of MDA-MB-231 cells. (A, B) Representative pictures and quantitative analysis of migrated MDA-MB-231 cells after transfection with the VEGF siRNA/PLEGP1800 nanocomplex for 6 h. Quantitative analysis was performed by counting the number of cells on the bottom of the membrane in six random microscope fields from three independent samples. (C, D) Representative pictures and quantitative analysis of invasion of MDA-MB-231 cells treated with the VEGF siRNA/PLEGP1800 nanocomplex for 24 h. Quantitative analysis was performed by counting the number of cells on the bottom of the membrane in six random microscope fields from three independent samples. The number of migrated and invaded cells were normalized to the group. treated with scrambled siRNA and presented as the percentage mean ± SD (n = 3). (** P < 0.01 (E) Representative images of 3D spheroid invasion of MDA-MB-231 cells (n = 4). (F) Quantification of spheroid invasion based on the changes of the total area of the spheroids. The spheroid areas of each group were normalized to the spheroid area of the group at day 1 and presented as the percentage mean ± SD (n = 4). (** P < 0.01)
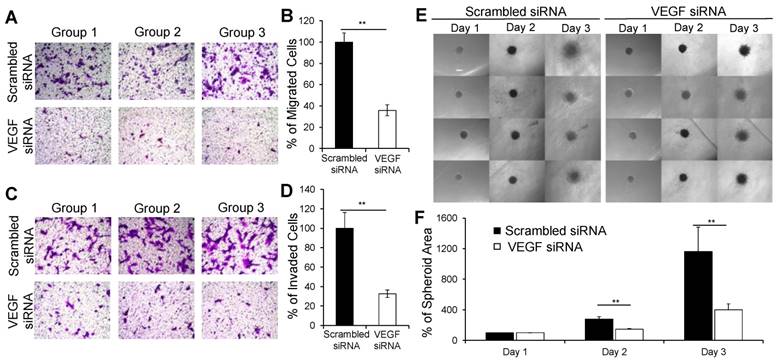
We also investigated the inhibitory effect of the VEGF siRNA/PLEGP1800 nanocomplex on the invasion of MDA-MB-231 cells in a 3D spheroid model, which mimics the tumor microenvironment (Figure 6E). The spheroids-treated with the scrambled siRNA spread with a typical “starburst” invasion pattern [44]. By contrast, the spheroids-treated with the VEGF siRNA displayed a dramatically retracted invasion with a 39% inhibition of invasion after 24 h and 58% inhibition after 48 h (Figure 6F). Based on these results, the VEGF siRNA/PLEGP1800 nanocomplex safely and efficiently delivered the siRNA into TNBC cells and exerted the anti-metastatic activity in the cells.
Cells Proliferation Study
We previously reported that VEGF siRNA significantly inhibited the proliferation of HER2+ breast cancer cells [19]. Herein, we treated MDA-MB-231 cells with the VEGF siRNA/PLEGP1800 nanocomplex for 48 h and 72 h and performed a cell proliferation study using the CellTiter-Glo luminescent cell viability assay (Figure 7A). The VEGF siRNA/PLEGP1800 nanocomplex-treated group exhibited a lower proliferation rate (73.1% for 48 h and 67.0% for 72 h) than the scrambled siRNA/PLEGP1800-treated group.
The VEGF siRNA/PLEGP1800 nanocomplex inhibits the proliferation of MDA-MB-231 and HUVEC cells. (A) Proliferation of MDA-MB-231 cells treated with the VEGF siRNA/PLEGP1800 nanocomplex for 48 h and 72 h. (B, C) Proliferation of HUVECs. HUVECs were cultured with serum-free DMEM medium, complete DMEM containing 10% FBS, and the culture medium derived from MDA-MB-231cells transfected with VEGF siRNA/PLEGP1800 or scrambled siRNA/PLEGP1800 for 24 h (B) and 48 h (C). (D) Tube formation of HUVECs in serum-free DMEM medium, complete DMEM containing 10% FBS, and the culture medium derived from MDA-MB-231cells transfected with the VEGF siRNA/PLEGP1800 or scrambled siRNA/PLEGP1800 for 24 h. All results are presented as the mean ± SD (n = 3). (* P < 0.05; ** P < 0.01).
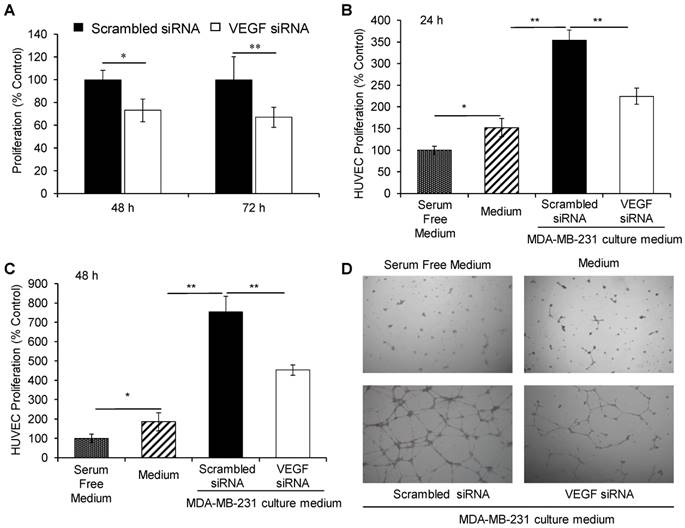
We then determined the proliferation of HUVECs in the presence of cell culture medium derived from VEGF siRNA/PLEGP1800 nanocomplex-treated MDA-MB-231 cells. As illustrated in Figure 7B & C, the cell proliferation rate of HUVEC was stimulated by incubation with MDA-MB-231 cells' culture medium, showing a 2.5-fold increase relative to control cells at 24 h and a 6.5-fold increase at 48 h. This result demonstrated that MDA-MB-231 cells secreted a significant amount of VEGF during culturing. Meanwhile, the VEGF siRNA/PLEGP1800 nanocomplex-treated group had a much lower HUVEC cell proliferation rate (63.5% for 24 h and 60.0% for 48 h) than the scrambled siRNA/PLEGP1800 nanocomplex-treated group, which indicates that silencing VEGF expression in TNBC cells can inhibit vascular endothelial cell proliferation in the tumor microenvironment.
In Vitro HUVEC Tube Formation
VEGF plays an important role in endothelial tube formation during tumor growth by positively regulating angiogenesis [17]. After demonstrating the inhibitory effect of the VEGF siRNA/PLEGP1800 nanocomplex on the proliferation of vascular endothelial cells HUVECs, we next sought to study the role of VEGF siRNA/PLEGP1800 nanocomplex in HUVEC tube formation. Similar to the proliferation study, HUVECs were seeded on a matrigel pre-coated 24-well plate and cultured for 24 h in the medium derived from MDA-MB-231 cells that were treated with the siRNA/PLEGP1800 nanocomplex. Tube formation of HUVECs was promoted by the medium derived from the scrambled siRNA-treated MDA-MB-231 cells, suggesting that MDA-MB-231 cells secret significant amount of VEGF into the medium. Accordingly, the stimulation was dramatically hampered in HUVECs treated with the medium derived from the VEGF siRNA-treated MDA-MB-231 cells (Figure 7D), suggesting that silencing VEGF expression in TNBC cells using the VEGF siRNA/PLEGP1800 nanocomplex can diminish the function of vascular endothelial cells.
In Vivo Biodistribution, Pharmacokinetic and Anti-Tumor Activity Study
Nanoscale particles can accumulate in primary and metastatic tumors by the enhanced permeability and retention (EPR) effect [45]. Cy5-labeled VEGF siRNAs were encapsulated in the siRNA/PLEGP nanocomplex for the biodistribution study. The mean fluorescence intensity of each organ was measured and presented as a ratio compared to the mean fluorescence intensity in the heart (Figure 8A & B). After systematic administration, free siRNAs are mainly degraded in the serum and eliminated from the kidney due to their small molecular weight. Thus, 24 h after administration via tail vein, the free Cy5-siRNA only showed negligible accumulation in tumors and other major organs. By contrast, the Cy5-siRNA/PLEGP1800 nanocomplex stayed in the circulation for a prolonged time and accumulated more in the tumor by the EPR effect. Interestingly, the Cy5-siRNA/PLEGP1800 nanocomplex also showed a high accumulation in the kidney, which may be caused by the positive charge on the surface of the glomerular basement membrane [46].
Biodistribution and anti-tumor activity of the VEGF siRNA/PLEGP1800 nanocomplex in an orthotopic TNBC mouse model. (A) Free Cy5-labeled siRNA or the Cy5-siRNA/PLEGP1800 nanocomplex were injected at an siRNA dose of 0.1 mg/kg via tail vein. Fluorescence images of the liver, tumor, lung, spleen, kidney, heart, and muscle from three mice were taken at 24 h post-injection. (B) Mean fluorescence intensity of each organ was normalized to the mean fluorescence intensity of the heart. The results are presented as the mean ± SD (n = 3). Relative tumor volume (C) and tumor weight (D) of the mice orthotopically implanted MDA-MB-231 cells. (E) Expression of VEGF in tumor tissues. In the end of the activity study, sera were collected from the mice to measure ALT (F) and AST (G) levels. The results of relative tumor volume are presented as the mean ± SE (n = 6). All other results are presented as the mean ± SD (n=6). (* P < 0.05, ** P < 0.01). (H) CD31staining of tumor tissues and H&E staining of the tumor, liver, kidney and lung. The scale bar represents 100 µm.
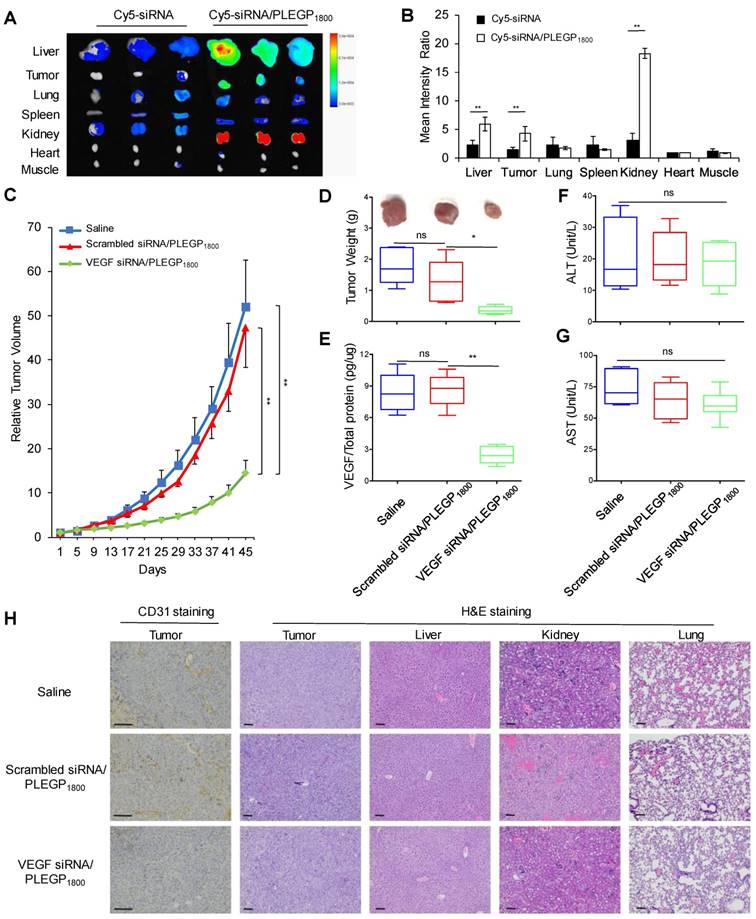
Pharmacokinetic profiles of free siRNA and the siRNA nanocomplex were evaluated in mice (Figure S3). The concentration of free siRNA in the blood decreased to 24.4% after 10 min, and the concentration was not detectable after 2 h. In contrast, the PLEGP1800 nanocomplex prolonged the circulation of encapsulated siRNA to more than 24 h. The AUCs for free siRNA and siRNA/PLEGP1800 nanocomplex were 543.7 min•µg/ml and 4609.2 min•µg/ml, respectively.
Anti-tumor activity of the VEGF siRNA/PLEGP1800 nanocomplex was evaluated in an orthotopic nude mouse model implanted with MDA-MB-231 cells. As illustrated in Figure 8C and 8D, the VEGF siRNA/PLEGP1800 nanocomplex significantly inhibited tumor growth, while the scrambled siRNA/PLEGP1800 nanocomplex did not show any antitumor effect against tumor growth. VEGF expression in tumors was also determined using an ELISA kit (Figure 8E) and western blot (Figure S4). The VEGF siRNA/PLEGP1800 nanocomplex-treated group showed significantly lower VEGF expression, demonstrating that the VEGF siRNA efficiently silenced the expression of VEGF in the tumor. The lower level of VEGF expression in tumors was consistent with the reduced CD31 expression in tumor tissues (Figure 8H, S5), suggesting that inhibition of VEGF leads to impaired angiogenesis signals from TNBC cells to vascular endothelial cells in the tumor microenvironment. Meanwhile, no significant hepatotoxicity (ALT & AST) was observed in the mice treated with the siRNA/PLEGP1800 nanocomplex (Figure 8F, G). This is in agreement with H&E staining of the major organs, including the liver, kidney and lung, in which no distinct morphology changes were observed.
Discussion
PEIs with high molecular weights (10 to 30 kDa) have been widely used as efficient transfection agents for nucleic acids, but high cytotoxicity largely limits their therapeutic applications [29]. The toxicity of HMW PEIs is mainly due to membrane destabilization caused by binding to plasma membrane proteoglycans and mitochondria-mediated apoptosis [47]. LMW PEIs with molecular weights less than 2kDa are nontoxic in vivo even at high concentrations. However, they cannot efficiently condense and deliver siRNAs to target cells. One solution is to use biodegradable polymers to modify LMW PEIs to increase their condensation capability with siRNAs and subsequently enhance the siRNA delivery efficiency [29, 30, 48]. Compared with linear PEIs, branched PEIs have a higher amine density and are increasingly being used to synthesize PEI-grafted polymers with a longer half-life and lower toxicity [49].
In the present study, a biodegradable multiblock copolymer was synthesized and grafted to LMW PEIs for siRNA delivery. The novelty of the polymer lies in the fact that the monomer backbone is composed of lysine, a natural amino acid, and glutaric acid, a natural metabolite of lysine and tryptophan in the body. Therefore, the polymer is fully biocompatible and can be completely degraded in the body. In addition, grafting of LMW PEIs to the polymer increases the charge density of PEIs, leading to the use of less amount of PEI to condense siRNAs. A small PEG (3.4 kDa) is also incorporated in the monomer to increase the space within the monomer so that PEI can be efficiently grafted. As shown in Table 1, steric hindrance is the determining factor in the grafting efficiency of the copolymer. Another function of PEG is to form hydrophilic shell (Figure 1) on the surface of the siRNA nanocomplex to reduce its nonspecific binding with serum proteins and prolong its circulation time in the body.
As illustrated in Figure 1, we synthesized three PLEGPs grafted with branched PEIs with different molecular weights (600, 1800, and 10K Da). Regardless of the PEI's molecular weight, PLEGPs efficiently condensed siRNAs into nanoscale particles with a narrow size distribution (Figure 2), suggesting that grafting of LMW PEIs to the polymer dramatically enhance PEIs' condensation capability. Meanwhile, condensation capability of PLEGPs is correlated with the grafting efficiency and the molecular weight of the PEIs (Figure 2), suggesting that PEI in the copolymer is the essential part for siRNA condensation. Similarly, PLEGPs' cellular uptake, cytotoxicity, and tumor penetration are strongly correlated with the molecular weight of PEI (Figures 3, 4, and 5). While PEIs with high molecular weight exhibited higher cellular uptake and tumor penetration, their cytotoxicity was also higher. PLEGP1800 exhibits similar cellular uptake as Lipofectamine® 2000 but with a much lower cytotoxicity. We subsequently selected PLEGP1800 as the candidate for the activity studies because of its low cytotoxicity as well as a reasonably high cellular uptake and tumor penetration.
Unlike the monolayer two-dimensional cell culture system, the 3D spheroid model mimics the complex cell-cell and cell-ECM complexity interactions in the tumor microenvironment [50, 51]. Despite accumulation in the tumor microenvironment by the EPR effect, polymeric nanoparticles have poor penetration into the tumors, which limit their therapeutic potential in cancer therapy. One solution is to modify the nanoparticles with PEG to improve the penetration efficiency by blocking nonspecific interaction with proteins and tumor cells in the tumor microenvironment [52, 53]. In our study (Figure 5), MDA-MB-231 spheroids with a diameter of approximately 500 µm were prepared to mimic the in vivo tumor microenvironment and determine the penetration ability of the VEGF siRNA/PLEGP nanocomplex [40, 41]. All the three PLEGP nanocomplexes demonstrated efficient tumor penetration in a 3D spheroid model. At the same time, the penetration efficiency of PLEGP1800 and PLEGP10k were much higher than that of PLEGP600. This could be due to the relatively smaller particle size (Figure 2) and higher transfection efficiency (Figure 4) of the PLEGP1800 and PLEGP10k nanocomplexes. This is in accordance with another study showing the similar correlation between tumor penetration and cellular transfection efficiency [54].
VEGF is highly expressed in the tumors of TNBC patients. The expression of VEGF in TNBC cells is 3 times higher than ER/PR+ subtypes and 1.5 times higher than HER2+ subtypes. VEGF regulates the proliferation and survival of TNBC cells through an autocrine signaling pathway [8-11]. For example, Bachelder and colleagues found that the survival of three different TNBC cells is dependent on the VEGF autocrine signaling pathway, and silencing VEGF with an antisense oligodeoxynucleotide induces apoptosis of the cells [55]. Similarly, silencing VEGF with siRNA also inhibits the proliferation and induces apoptosis of various breast cancer cells in vitro and in vivo [56-59]. In accordance with these findings, the VEGF siRNA/PLEGP1800 nanocomplex significantly inhibited the proliferation of TNBC MDA-MB-231 cells (Figure 7). Moreover, the nanocomplex suppressed tumor growth in an orthotopic TNBC mouse model (Figure 8).
Metastasis is responsible for more than 90% of cancer-related mortality and is also the main cause of treatment failure of TNBC patients [60, 61]. Compared to other types of breast cancer, TNBC has similar local recurrence rate but dramatically higher rate of distant recurrence, indicating a high metastasis rate of TNBC [62]. VEGF not only induces angiogenesis in the tumor microenvironment but also promotes the invasion of TNBC cells by regulating chemokine receptor type 4 (CXCR4) signaling pathway in an autocrine manner [43]. Downregulation of VEGF in the TNBC microenvironment therefore inhibits tumor cell invasion [43]. In the present study, the siRNA/PLEGP1800 nanocomplex significantly silences VEGF expression and subsequently inhibits the migration and invasion of TNBC cells (Figure 6).
Another protumor activity of VEGF is to promote angiogenesis endothelial cells in the tumor microenvironment [17]. Blocking the interaction between tumor-secreted VEGF and VEGFR2 on the surface of endothelial cells utilizing anti-VEGF or anti-VEGFR2 antibodies has shown inhibitory effect on the proliferation, migration, survival and adhesion of endothelial cells [63-66]. After transfection with the VEGF siRNA/PLEGP1800 nanocomplex, VEGF secreted by MDA-MB-231 cells was reduced by 72% compared to the control group (Figure 5D). Compared to the cell culture medium derived from MDA-MB-231 cells, the medium derived from the nanocomplex-treated MDA-MB-231 showed less stimulation of the proliferation and tube formation of HUVECs (Figure 7B, 7C, and 7D). Accordingly, lower levels of VEGF and CD31 were observed in tumor tissues from the mice treated with the VEGF siRNA/PLEGP1800 nanocomplex (Figure 8F), demonstrating that inhibition of VEGF expression by the VEGF siRNA/PLEGP1800 nanocomplex leads to impaired angiogenesis in the tumor microenvironment in vivo.
In summary, we have developed a LMW PEI-grafted biocompatible copolymer to deliver the VEGF siRNA for TNBC therapy. The copolymer is biodegradable and composed of lysine and glutaric acid, which is a natural metabolite of amino acids in the body. The copolymer PLEGP1800 can easily form stable nanocomplexes with siRNAs and protect them from serum degradation. The siRNA/PLEGP1800 nanocomplex exhibited negligible cytotoxicity but showed high cellular uptake, high transfection efficiency, and high tumor penetration. The nanocomplex also demonstrated promising anti-tumor activities in vitro and in vivo in an orthotopic TNBC mouse model. To our knowledge, this is the first report showing antitumor activity of VEGF siRNA in an orthotopic TNBC mouse model after systematic administration. Taken together, our evidences suggest that PLEGP1800 is a safe and efficient polymer to deliver siRNAs for TNBC therapy. It could potentially be applied to other cancers by changing the siRNA sequence and incorporating tumor-specific ligands. It can also be used as a platform for cancer theranostics after incorporating an imaging agent.
Supplementary Material
Supplementary figures.
Acknowledgements
This work is supported by the American Cancer Society-Lee national Denim Day Research Scholar Grant (RSG-15-132-01-CDD). The work is partially supported by the awards (1R01GM121798 and 1R01CA23109901) from the National Institutes of Health.
Competing Interests
The authors have declared that no competing interest exists.
References
1. Morris GJ, Naidu S, Topham AK, Guiles F, Xu Y, McCue P. et al. Differences in breast carcinoma characteristics in newly diagnosed African-American and Caucasian patients: a single-institution compilation compared with the National Cancer Institute's Surveillance, Epidemiology, and End Results database. Cancer. 2007;110:876-84
2. Wang C, Zheng X, Shen C, Shi Y. MicroRNA-203 suppresses cell proliferation and migration by targeting BIRC5 and LASP1 in human triple-negative breast cancer cells. J Exp Clin Cancer Res. 2012;31:58
3. Hudis CA, Gianni L. Triple-negative breast cancer: an unmet medical need. Oncologist. 2011;16(Suppl 1):1-11
4. Sanford M. Subcutaneous trastuzumab: a review of its use in HER2-positive breast cancer. Target Oncol. 2014;9:85-94
5. Amiri-Kordestani L, Wedam S, Zhang L, Tang S, Tilley A, Ibrahim A. et al. First FDA approval of neoadjuvant therapy for breast cancer: pertuzumab for the treatment of patients with HER2-positive breast cancer. Clin Cancer Res. 2014;20:5359-64
6. Turner NC, Ro J, Andre F, Loi S, Verma S, Iwata H. et al. Palbociclib in Hormone-Receptor-Positive Advanced Breast Cancer. N Engl J Med. 2015;373:209-19
7. Kalimutho M, Parsons K, Mittal D, Lopez JA, Srihari S, Khanna KK. Targeted Therapies for Triple-Negative Breast Cancer: Combating a Stubborn Disease. Trends Pharmacol Sci. 2015;36:822-46
8. Kristensen TB, Knutsson ML, Wehland M, Laursen BE, Grimm D, Warnke E. et al. Anti-vascular endothelial growth factor therapy in breast cancer. Int J Mol Sci. 2014;15:23024-41
9. Linderholm BK, Hellborg H, Johansson U, Elmberger G, Skoog L, Lehtio J. et al. Significantly higher levels of vascular endothelial growth factor (VEGF) and shorter survival times for patients with primary operable triple-negative breast cancer. Ann Oncol. 2009;20:1639-46
10. Mohammed RA, Ellis IO, Mahmmod AM, Hawkes EC, Green AR, Rakha EA. et al. Lymphatic and blood vessels in basal and triple-negative breast cancers: characteristics and prognostic significance. Mod Pathol. 2011;24:774-85
11. Dales JP, Garcia S, Bonnier P, Duffaud F, Carpentier S, Djemli A. et al. [Prognostic significance of VEGF receptors, VEGFR-1 (Flt-1) and VEGFR-2 (KDR/Flk-1) in breast carcinoma]. Ann Pathol. 2003;23:297-305
12. Sledge GW. Anti-vascular endothelial growth factor therapy in breast cancer: game over? J Clin Oncol. 2015;33:133-5
13. Cameron D, Brown J, Dent R, Jackisch C, Mackey J, Pivot X. et al. Adjuvant bevacizumab-containing therapy in triple-negative breast cancer (BEATRICE): primary results of a randomised, phase 3 trial. Lancet Oncol. 2013;14:933-42
14. Miles DW, Dieras V, Cortes J, Duenne AA, Yi J, O'Shaughnessy J. First-line bevacizumab in combination with chemotherapy for HER2-negative metastatic breast cancer: pooled and subgroup analyses of data from 2447 patients. Ann Oncol. 2013;24:2773-80
15. Fan F, Wey JS, McCarty MF, Belcheva A, Liu W, Bauer TW. et al. Expression and function of vascular endothelial growth factor receptor-1 on human colorectal cancer cells. Oncogene. 2005;24:2647-53
16. Vincent L, Jin DK, Karajannis MA, Shido K, Hooper AT, Rashbaum WK. et al. Fetal stromal-dependent paracrine and intracrine vascular endothelial growth factor-a/vascular endothelial growth factor receptor-1 signaling promotes proliferation and motility of human primary myeloma cells. Cancer Res. 2005;65:3185-92
17. Lee TH, Seng S, Sekine M, Hinton C, Fu Y, Avraham HK. et al. Vascular endothelial growth factor mediates intracrine survival in human breast carcinoma cells through internally expressed VEGFR1/FLT1. PLoS Med. 2007;4:e186
18. Leng Q, Woodle MC, Lu PY, Mixson AJ. Advances in Systemic siRNA Delivery. Drugs Future. 2009;34:721
19. Tai W, Qin B, Cheng K. Inhibition of breast cancer cell growth and invasiveness by dual silencing of HER-2 and VEGF. Mol Pharm. 2010;7:543-56
20. Wittrup A, Lieberman J. Knocking down disease: a progress report on siRNA therapeutics. Nat Rev Genet. 2015;16:543-52
21. Hu Y, Zhou D, Li C, Zhou H, Chen J, Zhang Z. et al. Gene delivery of PEI incorporating with functional block copolymer via non-covalent assembly strategy. Acta Biomater. 2013;9:5003-12
22. Shukla RS, Tai W, Mahato R, Jin W, Cheng K. Development of streptavidin-based nanocomplex for siRNA delivery. Mol Pharm. 2013;10:4534-45
23. Guan J, Sun J, Sun F, Lou B, Zhang D, Mashayekhi V. et al. Hypoxia-induced tumor cell resistance is overcome by synergistic GAPDH-siRNA and chemotherapy co-delivered by long-circulating and cationic-interior liposomes. Nanoscale. 2017
24. Qin B, Chen Z, Jin W, Cheng K. Development of cholesteryl peptide micelles for siRNA delivery. J Control Release. 2013;172:159-68
25. Nel AE, Madler L, Velegol D, Xia T, Hoek EM, Somasundaran P. et al. Understanding biophysicochemical interactions at the nano-bio interface. Nat Mater. 2009;8:543-57
26. Akinc A, Thomas M, Klibanov AM, Langer R. Exploring polyethylenimine-mediated DNA transfection and the proton sponge hypothesis. J Gene Med. 2005;7:657-63
27. Godbey WT, Wu KK, Mikos AG. Size matters: molecular weight affects the efficiency of poly(ethylenimine) as a gene delivery vehicle. J Biomed Mater Res. 1999;45:268-75
28. Zhong Z, Feijen J, Lok MC, Hennink WE, Christensen LV, Yockman JW. et al. Low molecular weight linear polyethylenimine-b-poly(ethylene glycol)-b-polyethylenimine triblock copolymers: synthesis, characterization, and in vitro gene transfer properties. Biomacromolecules. 2005;6:3440-8
29. Chiper M, Tounsi N, Kole R, Kichler A, Zuber G. Self-aggregating 1.8kDa polyethylenimines with dissolution switch at endosomal acidic pH are delivery carriers for plasmid DNA, mRNA, siRNA and exon-skipping oligonucleotides. J Control Release. 2017;246:60-70
30. Thomas M, Ge Q, Lu JJ, Chen J, Klibanov AM. Cross-linked small polyethylenimines: while still nontoxic, deliver DNA efficiently to mammalian cells in vitro and in vivo. Pharm Res. 2005;22:373-80
31. Zhao Z, Li Y, Jain A, Chen Z, Liu H, Jin W. et al. Development of a peptide-modified siRNA nanocomplex for hepatic stellate cells. Nanomedicine. 2018;14:51-61
32. Tai W, Chen Z, Barve A, Peng Z, Cheng K. A novel rapamycin-polymer conjugate based on a new poly(ethylene glycol) multiblock copolymer. Pharm Res. 2014;31:706-19
33. Lou B, Connor K, Sweeney K, Miller IS, O'Farrell A, Ruiz-Hernandez E. et al. RGD-decorated cholesterol stabilized polyplexes for targeted siRNA delivery to glioblastoma cells. Drug Deliv Transl Res. 2019;9:679-93
34. Ray A, Dhar S, Ray BK. Control of VEGF expression in triple-negative breast carcinoma cells by suppression of SAF-1 transcription factor activity. Mol Cancer Res. 2011;9:1030-41
35. Tavera-Mendoza LE, Brown M. A less invasive method for orthotopic injection of breast cancer cells into the mouse mammary gland. Lab Anim. 2017;51:85-8
36. Essex S, Navarro G, Sabhachandani P, Chordia A, Trivedi M, Movassaghian S. et al. Phospholipid-modified PEI-based nanocarriers for in vivo siRNA therapeutics against multidrug-resistant tumors. Gene Ther. 2015;22:257-66
37. Feng Q, Yu MZ, Wang JC, Hou WJ, Gao LY, Ma XF. et al. Synergistic inhibition of breast cancer by co-delivery of VEGF siRNA and paclitaxel via vapreotide-modified core-shell nanoparticles. Biomaterials. 2014;35:5028-38
38. Li Y, Wu Y, Huang L, Miao L, Zhou J, Satterlee AB. et al. Sigma receptor-mediated targeted delivery of anti-angiogenic multifunctional nanodrugs for combination tumor therapy. J Control Release. 2016;228:107-19
39. Chen Y, Gu H, Zhang DS, Li F, Liu T, Xia W. Highly effective inhibition of lung cancer growth and metastasis by systemic delivery of siRNA via multimodal mesoporous silica-based nanocarrier. Biomaterials. 2014;35:10058-69
40. Tredan O, Galmarini CM, Patel K, Tannock IF. Drug resistance and the solid tumor microenvironment. J Natl Cancer Inst. 2007;99:1441-54
41. Yao Q, Choi JH, Dai Z, Wang J, Kim D, Tang X. et al. Improving Tumor Specificity and Anticancer Activity of Dasatinib by Dual-Targeted Polymeric Micelles. ACS Appl Mater Interfaces. 2017;9:36642-54
42. Horibe T, Torisawa A, Akiyoshi R, Hatta-Ohashi Y, Suzuki H, Kawakami K. Transfection efficiency of normal and cancer cell lines and monitoring of promoter activity by single-cell bioluminescence imaging. Luminescence. 2014;29:96-100
43. Bachelder RE, Wendt MA, Mercurio AM. Vascular endothelial growth factor promotes breast carcinoma invasion in an autocrine manner by regulating the chemokine receptor CXCR4. Cancer Res. 2002;62:7203-6
44. Vinci M, Box C, Eccles SA. Three-dimensional (3D) tumor spheroid invasion assay. J Vis Exp. 2015 e52686
45. Maeda H. Toward a full understanding of the EPR effect in primary and metastatic tumors as well as issues related to its heterogeneity. Adv Drug Deliv Rev. 2015;91:3-6
46. Zuckerman JE, Choi CH, Han H, Davis ME. Polycation-siRNA nanoparticles can disassemble at the kidney glomerular basement membrane. Proc Natl Acad Sci U S A. 2012;109:3137-42
47. Moghimi SM, Symonds P, Murray JC, Hunter AC, Debska G, Szewczyk A. A two-stage poly(ethylenimine)-mediated cytotoxicity: implications for gene transfer/therapy. Mol Ther. 2005;11:990-5
48. Son S, Singha K, Kim WJ. Bioreducible BPEI-SS-PEG-cNGR polymer as a tumor targeted nonviral gene carrier. Biomaterials. 2010;31:6344-54
49. Jere D, Jiang HL, Arote R, Kim YK, Choi YJ, Cho MH. et al. Degradable polyethylenimines as DNA and small interfering RNA carriers. Expert Opin Drug Deliv. 2009;6:827-34
50. Zanoni M, Piccinini F, Arienti C, Zamagni A, Santi S, Polico R. et al. 3D tumor spheroid models for in vitro therapeutic screening: a systematic approach to enhance the biological relevance of data obtained. Sci Rep. 2016;6:19103
51. Desoize B, Jardillier J. Multicellular resistance: a paradigm for clinical resistance? Crit Rev Oncol Hematol. 2000;36:193-207
52. Al-Abd AM, Lee SH, Kim SH, Cha JH, Park TG, Lee SJ. et al. Penetration and efficacy of VEGF siRNA using polyelectrolyte complex micelles in a human solid tumor model in-vitro. J Control Release. 2009;137:130-5
53. Kim TH, Mount CW, Gombotz WR, Pun SH. The delivery of doxorubicin to 3-D multicellular spheroids and tumors in a murine xenograft model using tumor-penetrating triblock polymeric micelles. Biomaterials. 2010;31:7386-97
54. Bhise NS, Gray RS, Sunshine JC, Htet S, Ewald AJ, Green JJ. The relationship between terminal functionalization and molecular weight of a gene delivery polymer and transfection efficacy in mammary epithelial 2-D cultures and 3-D organotypic cultures. Biomaterials. 2010;31:8088-96
55. Bachelder RE, Crago A, Chung J, Wendt MA, Shaw LM, Robinson G. et al. Vascular endothelial growth factor is an autocrine survival factor for neuropilin-expressing breast carcinoma cells. Cancer Res. 2001;61:5736-40
56. Doan CC, Le LT, Hoang SN, Do SM, Le DV. Simultaneous silencing of VEGF and KSP by siRNA cocktail inhibits proliferation and induces apoptosis of hepatocellular carcinoma Hep3B cells. Biol Res. 2014;47:70
57. Wang L, Zhang WJ, Xiu B, Ding Y, Li P, Ye WD. et al. Nanocomposite-siRNA approach for down-regulation of VEGF and its receptor in myeloid leukemia cells. Int J Biol Macromol. 2014;63:49-55
58. Kim SH, Jeong JH, Lee SH, Kim SW, Park TG. Local and systemic delivery of VEGF siRNA using polyelectrolyte complex micelles for effective treatment of cancer. J Control Release. 2008;129:107-16
59. Chen Y, Wang X, Liu T, Zhang DS, Wang Y, Gu H. et al. Highly effective antiangiogenesis via magnetic mesoporous silica-based siRNA vehicle targeting the VEGF gene for orthotopic ovarian cancer therapy. Int J Nanomedicine. 2015;10:2579-94
60. Brabletz T, Lyden D, Steeg PS, Werb Z. Roadblocks to translational advances on metastasis research. Nat Med. 2013;19:1104-9
61. Sun B, Zhang S, Zhang D, Li Y, Zhao X, Luo Y. et al. Identification of metastasis-related proteins and their clinical relevance to triple-negative human breast cancer. Clin Cancer Res. 2008;14:7050-9
62. Dent R, Trudeau M, Pritchard KI, Hanna WM, Kahn HK, Sawka CA. et al. Triple-negative breast cancer: clinical features and patterns of recurrence. Clin Cancer Res. 2007;13:4429-34
63. Wu Y, Sun WL, Feng JF. Antiangiogenic therapy in the management of breast cancer. Asia Pac J Clin Oncol. 2013;9:110-6
64. Deissler HL, Deissler H, Lang GE. Actions of bevacizumab and ranibizumab on microvascular retinal endothelial cells: similarities and differences. Br J Ophthalmol. 2012;96:1023-8
65. Rusovici R, Patel CJ, Chalam KV. Bevacizumab inhibits proliferation of choroidal endothelial cells by regulation of the cell cycle. Clin Ophthalmol. 2013;7:321-7
66. Ferrara N, Gerber HP, LeCouter J. The biology of VEGF and its receptors. Nat Med. 2003;9:669-76
Author contact
Corresponding author: Kun Cheng, Ph.D., Professor, Division of Pharmacology and Pharmaceutical Sciences, School of Pharmacy, University of Missouri-Kansas City, 2464 Charlotte Street, Kansas City, MO 64108. Phone: (816) 235-2425; Fax: (816) 235-5779; E-mail: chengkunedu.
 Global reach, higher impact
Global reach, higher impact