13.3
Impact Factor
Theranostics 2021; 11(1):132-146. doi:10.7150/thno.47525 This issue Cite
Research Paper
Transcriptomic analysis identifies a tumor subtype mRNA classifier for invasive non-functioning pituitary neuroendocrine tumor diagnostics
1. Department of Neurosurgery, Pituitary Center, Peking Union Medical College Hospital (PUMCH), Chinese Academy of Medical Sciences (CAMS) & Peking Union Medical College (PUMC), Beijing 100730, PR China.
2. State Key Laboratory of Medical Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences (CAMS) & School of Basic Medicine, Peking Union Medical College (PUMC), Beijing 100005, PR China.
3. Department of Biochemistry, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences (CAMS) & School of Basic Medicine, Peking Union Medical College (PUMC), Beijing 100005, PR China.
4. Department of Pathology, Peking Union Medical College Hospital (PUMCH), Chinese Academy of Medical Sciences (CAMS) & Peking Union Medical College (PUMC), Beijing 100730, PR China.
5. Medical Epigenetic Research Center, Chinese Academy of Medical Sciences (CAMS), Beijing 100005, PR China.
#These authors contributed equally to this work.
Abstract
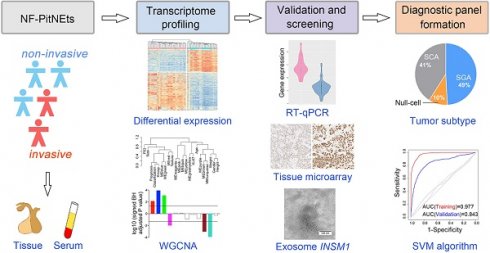
Rationale: The invasive behavior of non-functioning pituitary neuroendocrine tumors (NF-PitNEts) presents obstacles for complete surgical resection and is indicative of poor prognosis. Therefore, developing reliable diagnostic tools for identifying invasive PitNEts would be helpful in guiding surgical decisions and, in particular, the follow-up treatment.
Methods: We analyzed differential gene expression profiles between 39 non-invasive and 22 invasive NF-PitNEts by high-throughput sequencing, gene co-expression, and functional annotation. Twenty-one transcripts were further validated by Taqman-qPCR in another 143 NF-PitNEt samples. The histological expression and serum-exosomal mRNA of three candidate genes were examined by tissue microarray and droplet digital PCR.
Results: Non-invasive and invasive NF-PitNEts were clustered into distinct groups with a few outliers because of their gonadotroph, corticotroph, or null cell lineages. The gene signature with strong invasive potential was enriched in 'Pathways in cancers' and 'MAPK pathway', with significantly higher in situ INSM1 and HSPA2 protein expression in invasive NF-PitNEts. Further integration of the 20 qPCR-validated differentially expressed genes and pituitary cell lineages provided a gene-subtype panel that performed 80.00-90.24% diagnostic accuracy for the invasiveness of NF-PitNEts.
Conclusion: Our approach defined new characteristics in the core molecular network for patients at risk for invasive NF-PitNEt, representing a significant clinical advance in invasive PitNEt diagnostics.
Keywords: Pituitary neuroendocrine tumors, transcriptome, diagnostic panel, INSM1, HSPA2
 Global reach, higher impact
Global reach, higher impact