13.3
Impact Factor
Theranostics 2021; 11(1):426-444. doi:10.7150/thno.50281 This issue Cite
Research Paper
Oscillating lncRNA Platr4 regulates NLRP3 inflammasome to ameliorate nonalcoholic steatohepatitis in mice
1. College of Pharmacy, Jinan University, Guangzhou 510632, China.
2. Integrated Chinese and Western Medicine Postdoctoral research station, Jinan University, Guangzhou, 510632, China.
3. Institution of Laboratory Animal, Jinan University, 601 Huangpu Avenue West, Guangzhou, China.
#These authors contributed equally to this work.
Abstract
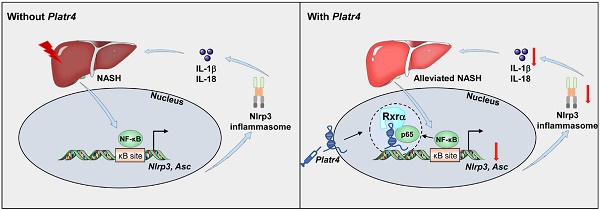
Background: Understanding the molecular events and mechanisms underlying development and progression of nonalcoholic steatohepatitis is essential in an attempt to formulating a specific treatment. Here, we uncover Platr4 as an oscillating and NF-κB driven lncRNA that is critical to the pathological conditions in experimental steatohepatitis
Methods: RNA-sequencing of liver samples was used to identify differentially expressed lncRNAs. RNA levels were analyzed by qPCR and FISH assays. Proteins were detected by immunoblotting and ELISA. Luciferase reporter, ChIP-sequencing and ChIP assays were used to investigate transcriptional gene regulation. Protein interactions were evaluated by Co-IP experiments. The protein-RNA interactions were studied using FISH, RNA pull-down and RNA immunoprecipitation analyses
Results: Cyclic expression of Platr4 is generated by the core clock component Rev-erbα via two RevRE elements (i.e., -1354/-1345 and -462/-453 bp). NF-κB transcriptionally drives Platr4 through direct binding to two κB sites (i.e., -1066/-1056 and -526/-516 bp), potentially accounting for up-regulation of Platr4 in experimental steatohepatitis. Intriguingly, Platr4 serves as a circadian repressor of Nlrp3 inflammasome pathway by inhibiting NF-κB-dependent transcription of the inflammasome components Nlrp3 and Asc. Loss of Platr4 down-regulates Nlrp3 inflammasome activity in the liver, blunts its diurnal rhythm, and sensitizes mice to experimental steatohepatitis, whereas overexpression of Platr4 ameliorates the pathological conditions in an Nlrp3-dependent manner. Mechanistically, Platr4 prevents binding of the NF-κB/Rxrα complex to the κB sites via a physical interaction, thereby inhibiting the transactivation of Nlrp3 and Asc by NF-κB.
Conclusions: Platr4 functions to inactivate Nlrp3 inflammasome via intercepting NF-κB signaling. This lncRNA might be an attractive target that can be modulated to ameliorate the pathological conditions of steatohepatitis.
Keywords: Platr4, NLRP3 inflammasome, NF-κB, RXR, Steatohepatitis
 Global reach, higher impact
Global reach, higher impact